🚨🚨🚨 Gear up. Revealing three theses from Wuhan Institute of Virology which provides new details on Mojiang sampling, 4991 genome, 7896-clade, unpublished CoVs, coronavirus reverse genetics and more! All theses were supervised by Shi Zhengli. Links at the end of the thread.
1/ Serological Cross-Reactivity Analysis of Coronavirus Based on Nucleocapsid Proteins (by Ning Wang, May 2014) 



Here are some key takeaways:
• It is the 1st time a WIV study mentions the miners outbreak: "Three miners died from pneumonia in Mojiang..we investigated the virus carried by bats in this cave..it is likely that the six miners were infected with the pathogen carried by bats."
• It is the 1st time a WIV study mentions the miners outbreak: "Three miners died from pneumonia in Mojiang..we investigated the virus carried by bats in this cave..it is likely that the six miners were infected with the pathogen carried by bats."
• This very obviously - and for the third time - flies in the face of WIV's claim that the miner’s illness were thought to be related to fungal infection.
https://twitter.com/MarionKoopmans/status/1369901874313330690?s=20
• What I find most fascinating: In this study, "fever patient sera were obtained from a hospital in Yunnan Province." 30 sera were obtained in total, and all marked MJ (almost certainly Mojiang).
Note: Shi's @Nature addendum mentions only 13 sera collected from 4 patients.
Note: Shi's @Nature addendum mentions only 13 sera collected from 4 patients.

• A total of 170 bat samples were collected from Mojiang in 2012, of which 109 were positive for coronavirus, with a positive rate of 64%!
Seems the mine was indeed a holy grail for virus hunters!
Seems the mine was indeed a holy grail for virus hunters!
• The author notes: "The full-length sequences of N and RdRp genes of several coronaviruses have been successfully amplified, but only a small number of full-length sequences of S genes have been amplified."
• On S gene sequences, the author notes: “there are still some urgent technical barriers to the amplification of the S gene sequences, especially the full-length sequences, and it is urgent to accomplish the relevant technical breakthroughs in future studies”
• In the acknowledgement section, the author thanks Linfa Wang for his "guidance" on the project. This confirms my long held suspicion that Linfa Wang knew about the miners outbreak.
"If three people died and it was controlled would we know it?"
"If three people died and it was controlled would we know it?"
https://twitter.com/TheSeeker268/status/1273074790933250049?s=20
• And finally the study reports, for the first time ever, Ra4991: "Overlap PCR was performed on the amplified bat coronavirus N gene, of which 4991, 3740(β)-N was amplified by laboratory associate" 



2/ Reverse Genetic System of Bat SARS-like Coronaviruses and Function of ORFX (by Lei-Ping Zeng, April 2017) 





A few illustrative quotes from the thesis:
• “In this study, we combined the advantages of two mainstream coronavirus reverse genetics methods, established a new efficient and cost-effective method”, and "a scheme to replace the S gene without traces”.
• “In this study, we combined the advantages of two mainstream coronavirus reverse genetics methods, established a new efficient and cost-effective method”, and "a scheme to replace the S gene without traces”.
• The reverse genetics system established in this study “can be used to rescue viruses whose sequences have only been identified, to study the function of virus-encoded proteins, to help assess their pathogenicity, and to explore their evolutionary and transmission mechanisms.”
• The author also points out: “It can also help to evaluate the effectiveness of existing antibodies and drugs against SARS against these bat-derived SL-CoV.”
• And: “determining in the lab whether recombination occurs after co-infection with different SL-CoV strains and what new strains may arise..is beneficial for studying the evolutionary pattern of SL-CoV and providing a basis for preventing possible future outbreaks..” 

• The author summarizes the outlook as: “At this point, our laboratory can establish a complete system from viral pathogen discovery, pathogen rescue and pathogen pathogenicity assessment to deepen and accelerate the epidemiological study of emerging viruses.”
• Zeng also mentions new methods that have emerged recently: “yeast recombination or in vitro recombination”, which “applied to the reverse genetics of coronaviruses could develop more efficient and economical method” for virology research. What happened to this line of study?
• “This study conducted large-scale detection and analysis of SARS-like coronavirus in bat samples collected from twenty regions in China from 2011 to 2016." A total of 9261 samples were collected (2815 from Yunnan), and 170 SARS-like coronavirus were detected (92 from Yunnan). 

• "In this study, the full-length RdRp sequence (2757 bp) was amplified for phylogenetic analysis of the virus, and finally 60 full-length RdRp sequences of SARS-like coronaviruses were obtained from positive samples."
• This is important: “SL-CoVs without deletions in the RBM region are able to utilize ACE2 receptors.. whereas SL-COVs with deletions in this region are not able to utilize ACE2 receptors.. Currently, bat SL-CoVs without deletions in the RBM region are only found in Yunnan.”
• Based on the phylogenetic analysis of the coronaviruses, the author concludes: "bat SARS-like coronaviruses tend to cluster together more by geographic location than by host species, suggesting that space plays a greater barrier role."
• In this study they sequenced the whole genome of Ra4991, and three other SARS-like coronaviruses. (The table could hold important clues.) 
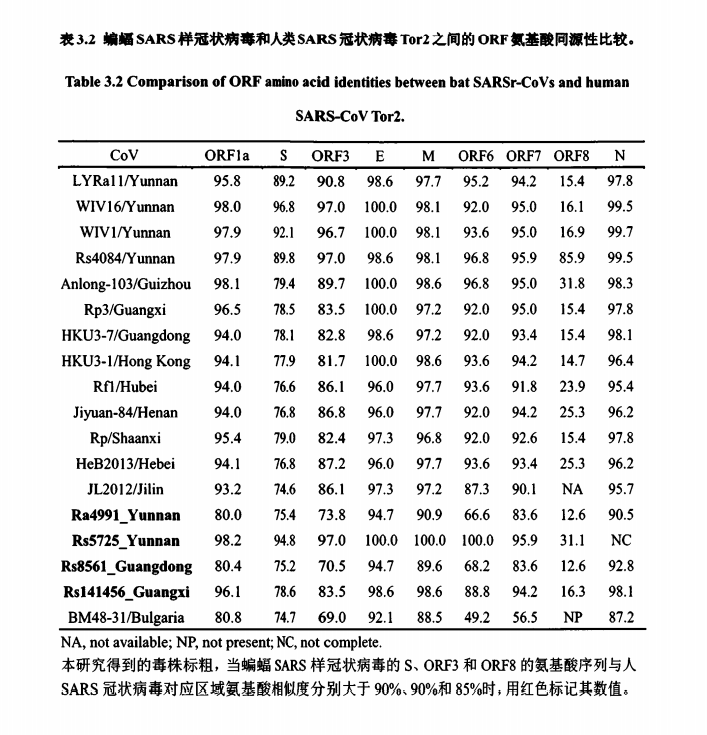
• And this is big: The study confirms, for the first time, that the 7896-clade from the mojiang mine were sequenced way before the covid outbreak. And more than just the RdRp fragment: i.e. the Spike of 7896; ORF8 of 7896, 7909 & 7952; and the RBD+NTD of 7905, 7909, 7924 & 7931. 





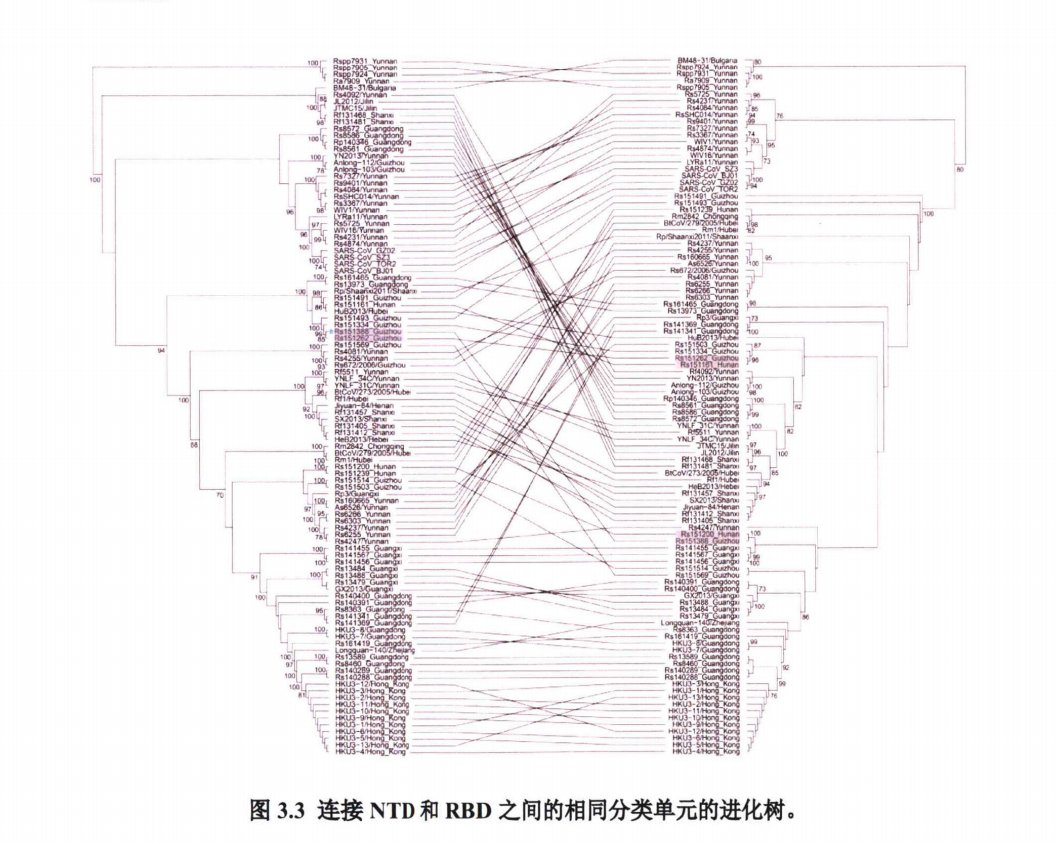

• So just to be clear: WIV had all these sequences at the time most closely related to SARS-CoV-2 and they chose not to disclose it in Jan 2020, and even after 15 months.
And that's a wrap! Here is the link for the theses and their translations:
drive.google.com/drive/folders/…
Copy-edited and translated by: @francsicodeasis (thank you so much!) and me, with generous help from Google, Adobe and DeepL.
drive.google.com/drive/folders/…
Copy-edited and translated by: @francsicodeasis (thank you so much!) and me, with generous help from Google, Adobe and DeepL.
• • •
Missing some Tweet in this thread? You can try to
force a refresh
















