I'm curious about the most tweeted about #rstats packages in the past ~week, let's explore this using R! 🧵
I'm going to use these packages:
*️⃣ tidyverse
*️⃣ rtweet
*️⃣ rvest
*️⃣ tidytext
you can install them with:
install.packages(c("tidyverse", "rtweet", "rvest", "tidytext"))
I'm going to use these packages:
*️⃣ tidyverse
*️⃣ rtweet
*️⃣ rvest
*️⃣ tidytext
you can install them with:
install.packages(c("tidyverse", "rtweet", "rvest", "tidytext"))
First, let's get a vector of all #rstats packages on CRAN!
library(rvest)
r_pkgs <- read_html('cloud.r-project.org/web/packages/a…') %>%
html_nodes('td:nth-child(1)') %>%
html_text()
library(rvest)
r_pkgs <- read_html('cloud.r-project.org/web/packages/a…') %>%
html_nodes('td:nth-child(1)') %>%
html_text()

Then let's pull in all tweets in the past week or so that use the #rstats hashtag
library(rtweet)
df <- search_tweets(q = "#rstats", n = 7000, include_rts = FALSE)
library(rtweet)
df <- search_tweets(q = "#rstats", n = 7000, include_rts = FALSE)

Alright! If you're following along live, we've got 4,948 tweets (you may get a different number if you run this in a few minutes! In fact, I expect you'd at least pick up one more tweet: this one! #rstats) 

🧹 Let's tidy this up! I'm going to build up step-by-step here.
First let's unnest the text into words.
library(tidyverse)
library(tidytext)
pkgs <- df %>%
unnest_tokens(word, text)
First let's unnest the text into words.
library(tidyverse)
library(tidytext)
pkgs <- df %>%
unnest_tokens(word, text)

Now let's remove any "stop words" this will be things like "not", "do" etc.
library(tidyverse)
library(tidytext)
pkgs <- df %>%
unnest_tokens(word, text) %>%
anti_join(stop_words)
library(tidyverse)
library(tidytext)
pkgs <- df %>%
unnest_tokens(word, text) %>%
anti_join(stop_words)

Now I'm going to keep only the "words" that are found in that character vector of packages we created before (I called it r_pkgs)
library(tidyverse)
library(tidytext)
pkgs <- df %>%
unnest_tokens(word, text) %>%
anti_join(stop_words) %>%
filter(word %in% r_pkgs)
library(tidyverse)
library(tidytext)
pkgs <- df %>%
unnest_tokens(word, text) %>%
anti_join(stop_words) %>%
filter(word %in% r_pkgs)

Let's count up the words and keep the top 5 to plot.
library(tidyverse)
library(tidytext)
pkgs <- df %>%
unnest_tokens(word, text) %>%
anti_join(stop_words) %>%
filter(word %in% r_pkgs) %>%
count(word) %>%
slice_max(n, n = 5)
library(tidyverse)
library(tidytext)
pkgs <- df %>%
unnest_tokens(word, text) %>%
anti_join(stop_words) %>%
filter(word %in% r_pkgs) %>%
count(word) %>%
slice_max(n, n = 5)

Ok, final tidying step, let's reorder so it plots nicely
library(tidyverse)
library(tidytext)
pkgs <- df %>%
unnest_tokens(word, text) %>%
anti_join(stop_words) %>%
filter(word %in% r_pkgs) %>%
count(word) %>%
slice_max(n, n = 5) %>%
mutate(word = reorder(word, n))
library(tidyverse)
library(tidytext)
pkgs <- df %>%
unnest_tokens(word, text) %>%
anti_join(stop_words) %>%
filter(word %in% r_pkgs) %>%
count(word) %>%
slice_max(n, n = 5) %>%
mutate(word = reorder(word, n))
And there we have it! Top 5 recently tweeted about #rstats packages:
🥇 tensorflow
🥈 tidyverse
🥉 ggplot2
🏅 shiny
🏅 gt
Thanks for following along!
🥇 tensorflow
🥈 tidyverse
🥉 ggplot2
🏅 shiny
🏅 gt
Thanks for following along!
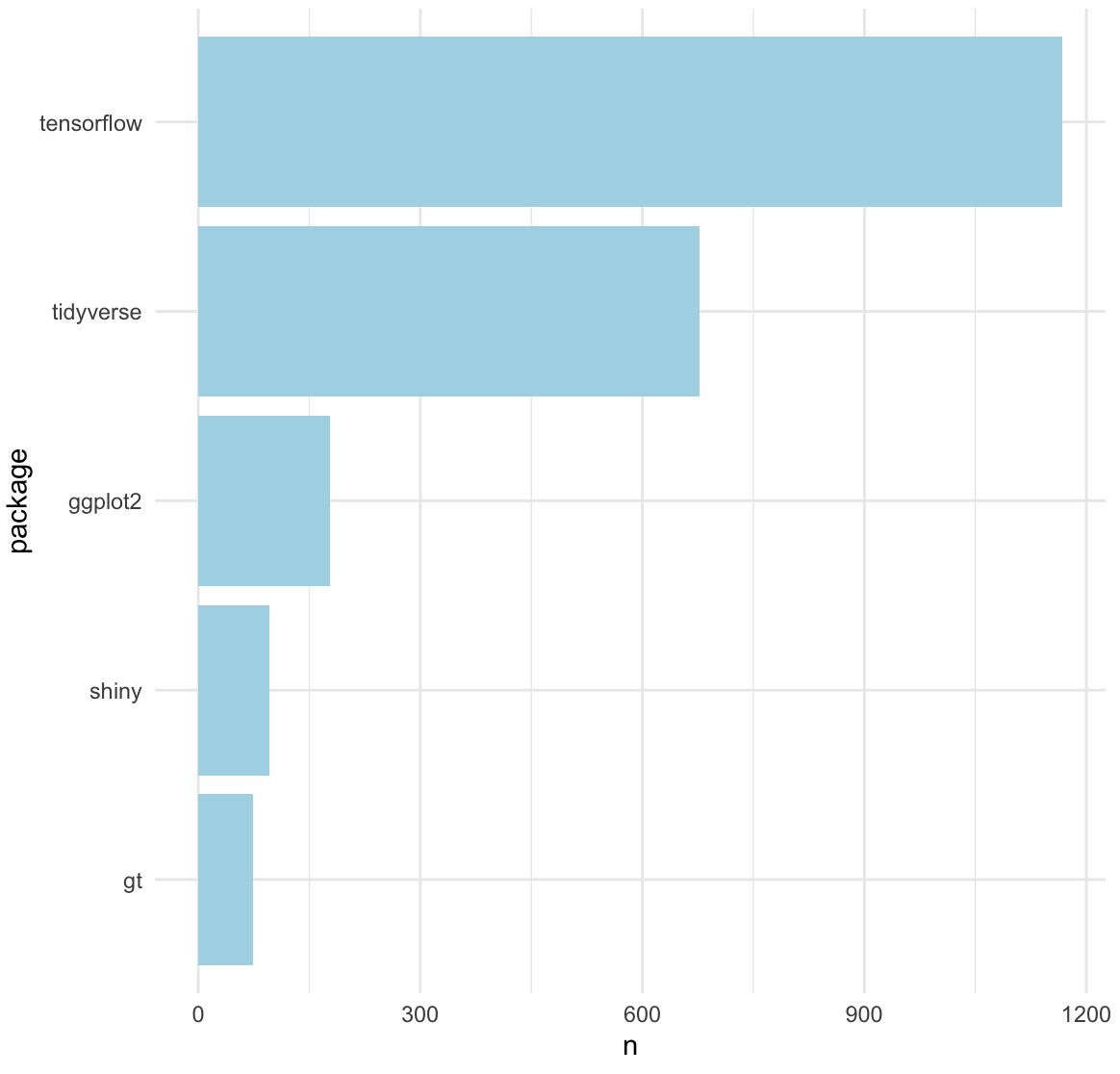
• • •
Missing some Tweet in this thread? You can try to
force a refresh









