Thrilled to share our new preprint 🌎🌍🌏! We made a generative model of the zebrafish brain, and show that by co-activating in different combis, 200 neural assemblies can explain the correlations and connectivity of 40k+ neurons!
doi.org/10.1101/2021.1…
In short: 1/8
doi.org/10.1101/2021.1…
In short: 1/8
Of course first a big thank you to everyone on the (largely twitterless) and inspiring interdisciplinary team; Jérôme, Guillaume, Geoffrey, @MichaelKunst5, @skeptencephalon @BormuthVolker, Bernhard & Georges 😎2/8
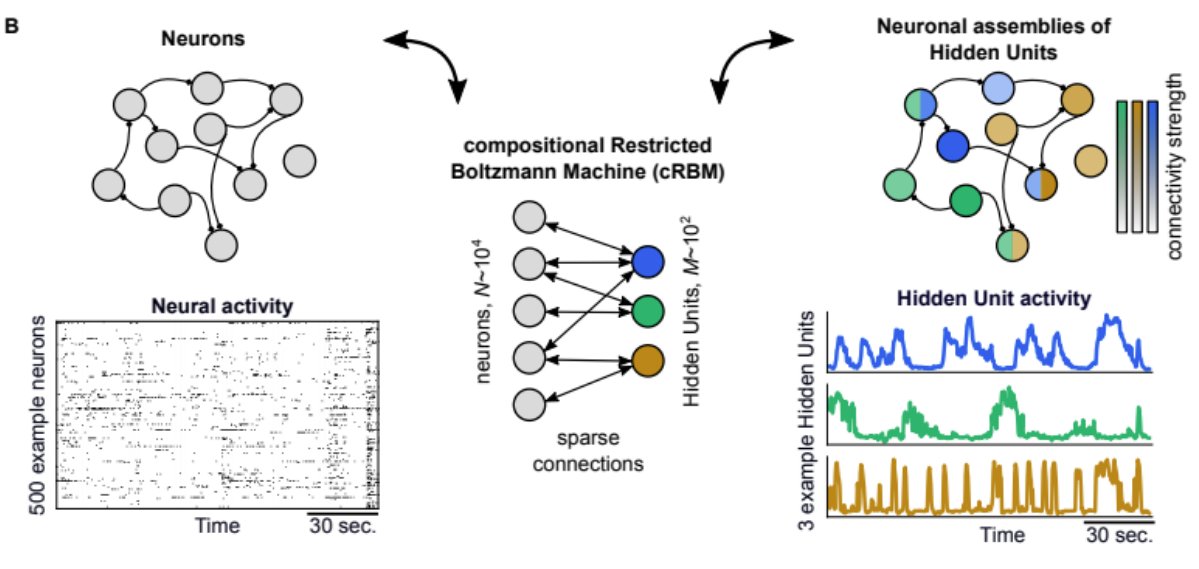
Neural assemblies are groups of neurons with correlated activity; neurons can be in multiple assemblies. More than just clusters, assemblies are thought to constrain spontaneous dynamics. To prove this, we need an assembly model that can generate neural data. Enter RBMs... 3/8
Full details in the preprint, the compositional RBM does exactly that. After training on zebrafish LSM data, we can sample new data with the same statistics as the empirical training data. And it defines assemblies via its low-dimensional bottleneck. 4/8 
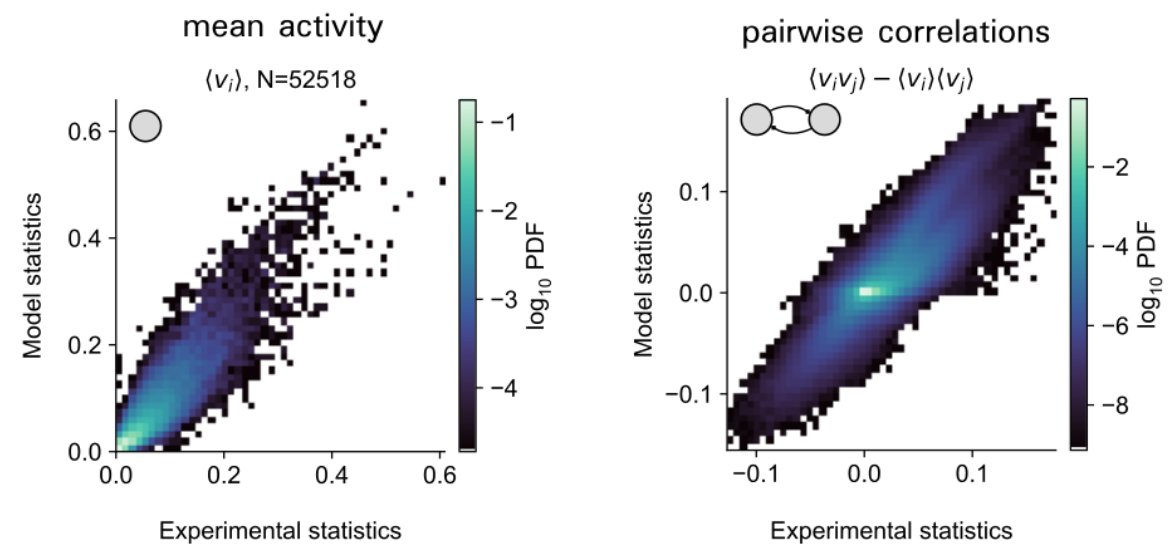
We can now verify that 200 assemblies are sufficient to generate spontaneous high-dim neural activity. We thus have 1) a model that can accurately generate neural data and 2) its underlying assemblies. 5/8 
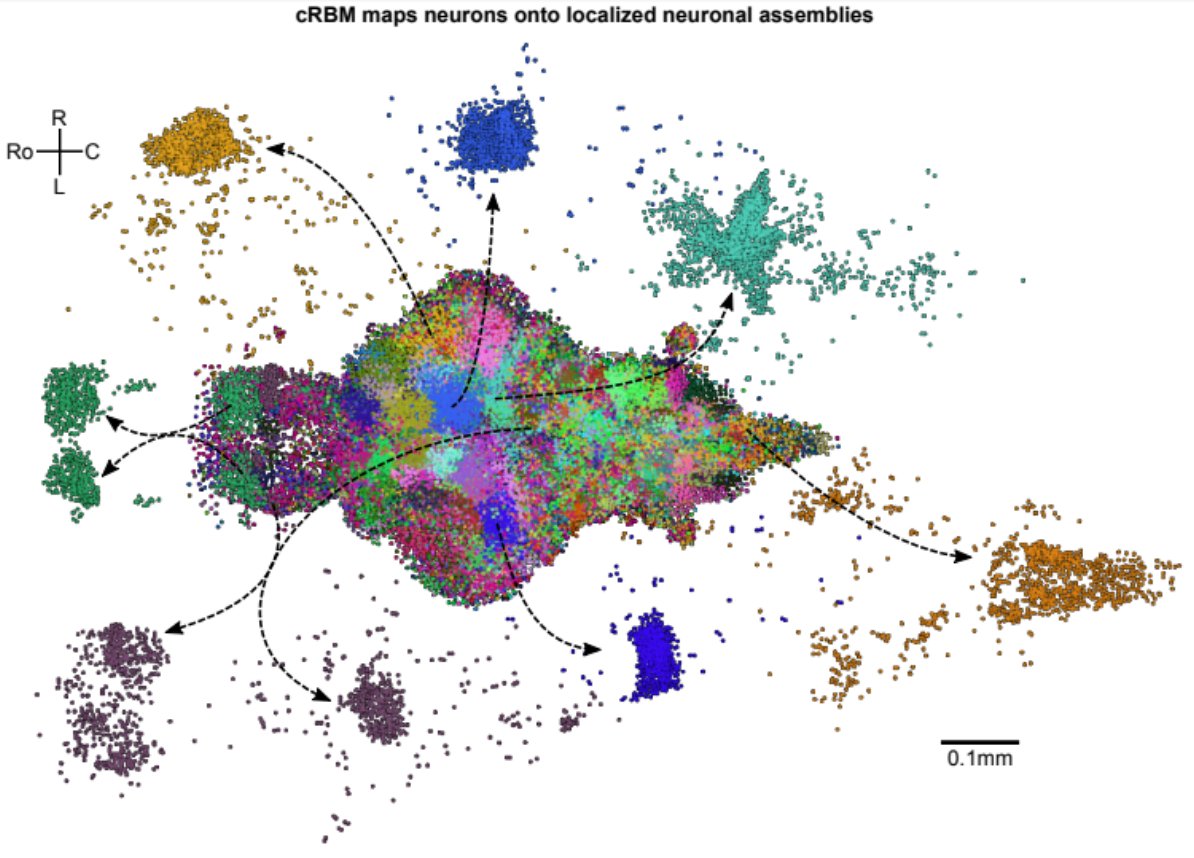
These assemblies show great variety in morphology; here are two example assemblies. Typically locally dense, but globally sparse. Assemblies can overlap, hence they are not independent and can be co-active in many different configurations. 6/8 
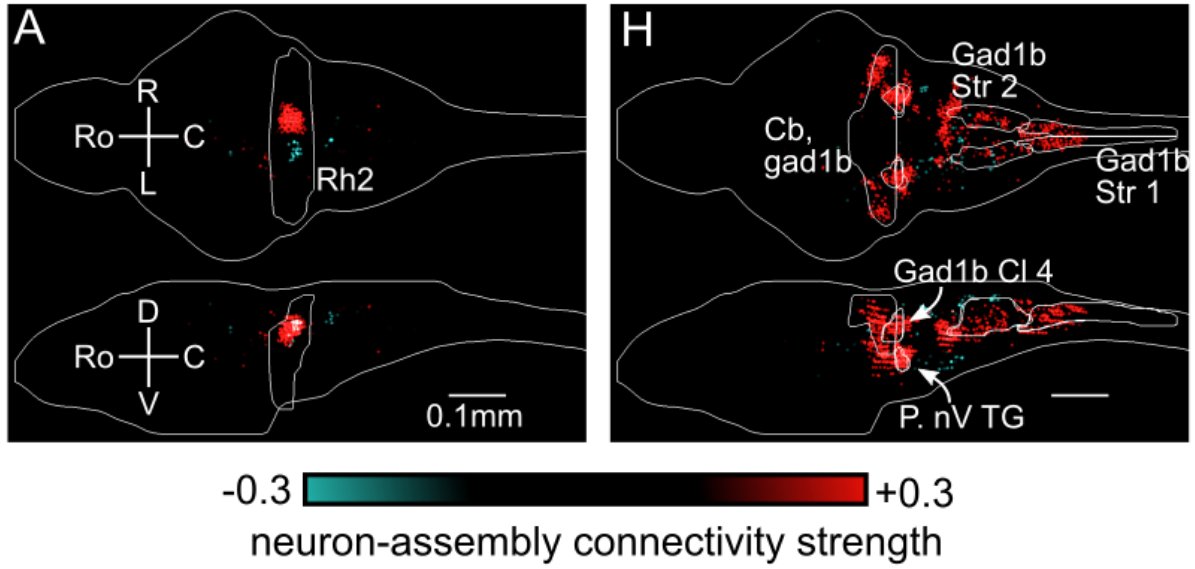
We mathematically derived the functional connectivity between all neurons, and by assessing which assemblies connect to which brain regions; a region-level connectivity map. This is conserved across individuals; and correlates to structural connectivity. 7/8 
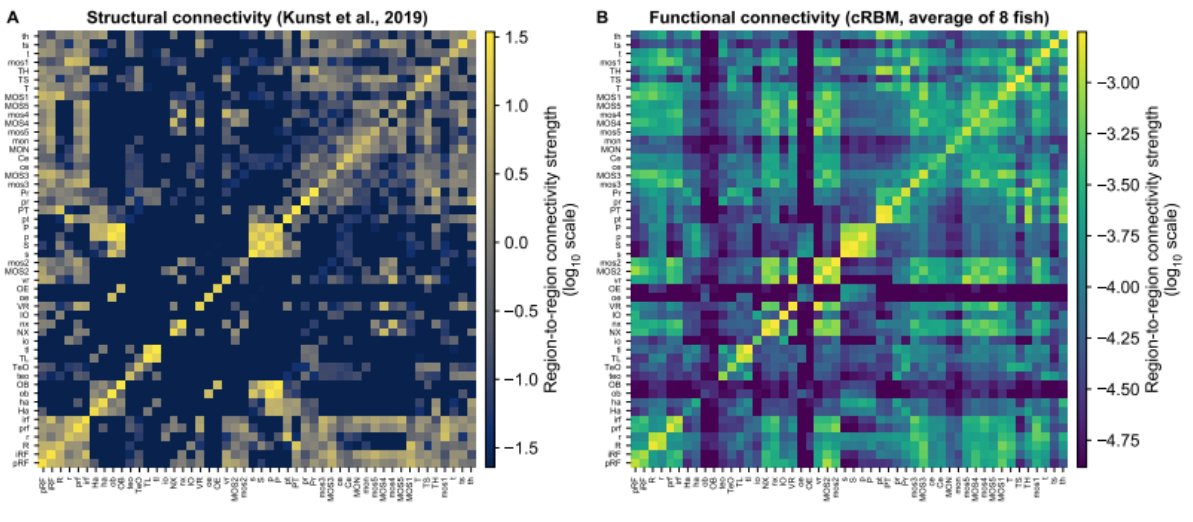
In summary; using a generative assembly-based model, we show that assemblies give rise to neural correlations and connectivity. Our cRBM was implemented in Python, and can be applied to any high-dim data. Code in preprint. Thanks for reading (my first tweet)! 8/8🥳
• • •
Missing some Tweet in this thread? You can try to
force a refresh




