
biorxiv.org/content/10.110… We developed a novel microfluidic isotachophoresis method (Ribo-ITP) to measure ribosome occupancy from ultra-low input samples including single cells. Here, we characterized transcriptome-wide ribosome occupancy during early mouse development. (1/6)
Our high coverage data enabled the first analysis of allele-specific ribosome occupancy in mouse preimplantation development. We discovered gene-specific differences in the timing of engagement of paternal RNAs with ribosomes. (2/6) 
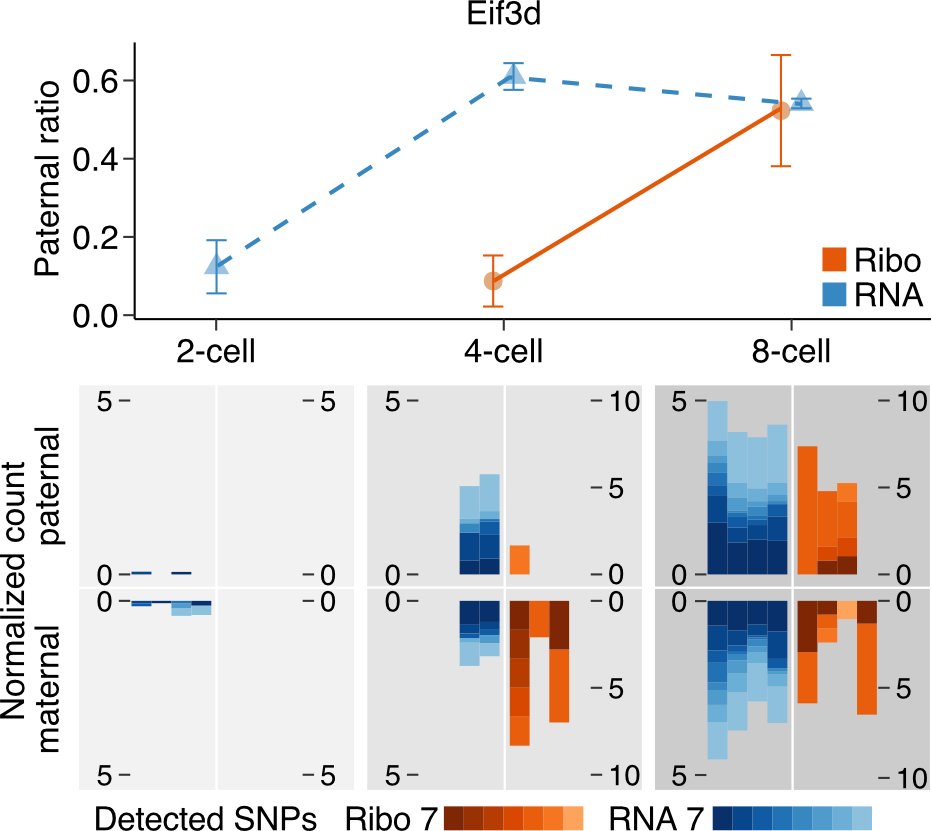
Moreover, we uncovered translation control as a key regulatory mechanism for the expression of proteins involved in the transition from meiosis to mitosis and N6-methyladenosine modification of RNAs. (3/6) 
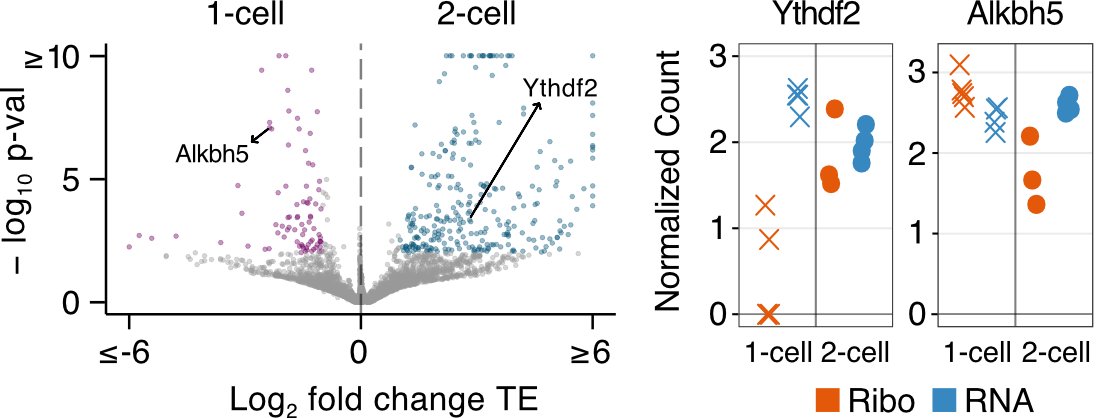
We assessed the contribution of ribosome occupancy to the proteome of mouse preimplantation embryos and identified temporal dynamics that eluded previous RNA expression analyses. (4/6) 

We are excited about the numerous applications that will be enabled by Ribo-ITP! Look out for a revised version of the manuscript which will be accompanied by step-by-step instructions and videos to help the research community utilize this exciting technology. (5/6)
We have already initiated several collaborative projects utilizing Ribo-ITP and look forward to many more productive collaborations! (6/6)
• • •
Missing some Tweet in this thread? You can try to
force a refresh



