Protein folding is so important. In 2023, DeepMind won the $250,000 Lasker award for their solution to the problem. A lot of people have asked me to explain protein folding in simple, understandable terms.
Here is my attempt at explaining just the problem.
🧵OPEN THE THREAD🧵
Here is my attempt at explaining just the problem.
🧵OPEN THE THREAD🧵
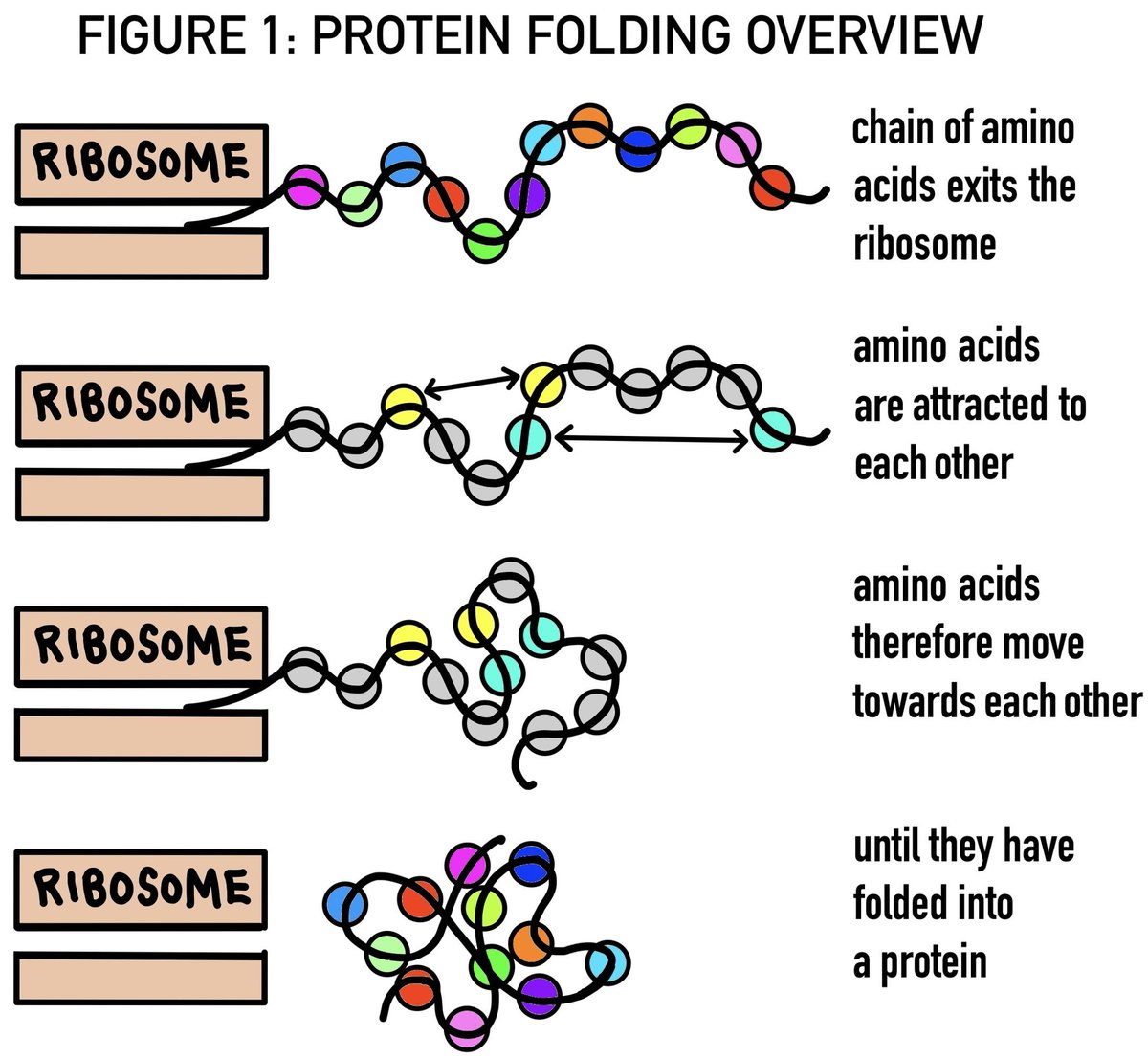
Understanding how a protein's amino acid sequence dictates its 3D shape—known as the "protein folding problem"—is a fundamental question in biology. Proteins are the workhorses of cells, and their functions depend on their shapes (structure).

Problem: Predicting this 3D shape from just the amino acid sequence.
This is tricky because proteins can fold in an astronomical number of ways, but only a few are biologically relevant.
Knowing the shape helps us understand function and design better treatments.
This is tricky because proteins can fold in an astronomical number of ways, but only a few are biologically relevant.
Knowing the shape helps us understand function and design better treatments.
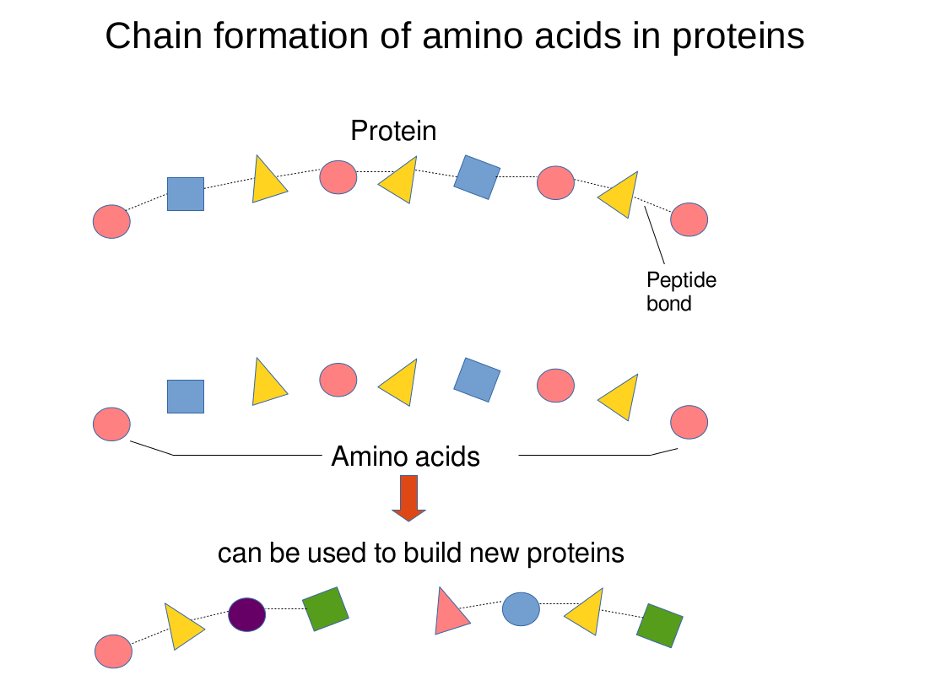
Before WWII, it was thought that protein properties were defined merely by their amino acid composition. However, post-1949, Frederick Sanger’s methods revealed that the sequence of these amino acids plays a crucial role. 
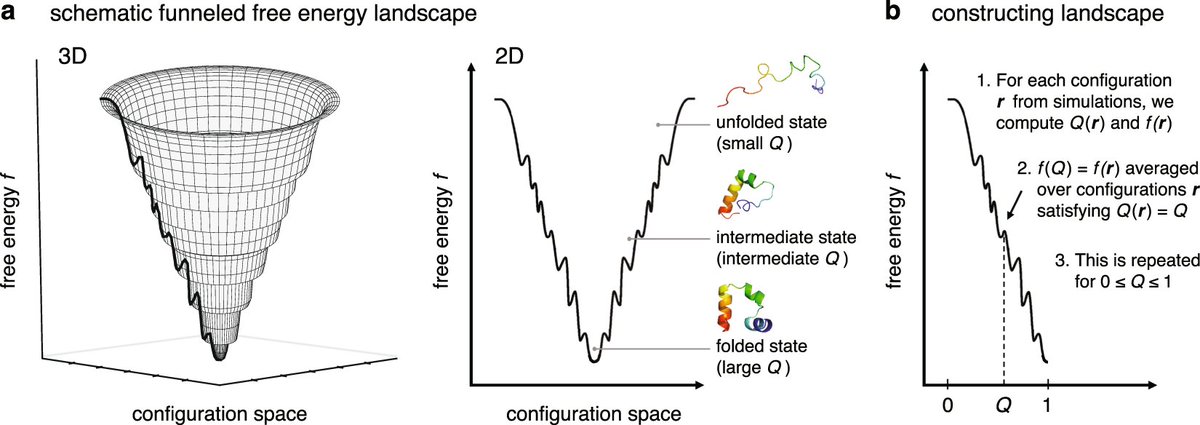
Cyrus Levinthal noted in the 1960s that it would take an astronomical amount of time for a protein to randomly try each possible fold before finding the correct structure. Yet, proteins fold correctly and quickly, usually within milliseconds. 

Protein folding is guided by an energy landscape shaped like a funnel. While there are many possible folded states, natural selection has optimized proteins to fold into a minimum-energy structure rapidly. This funnel guides the protein to its native state. 
A popular hypothesis, Anfinsen’s dogma, essentially states 'the amino acid sequence of a protein contained all of the information needed for the protein to reach the native conformation.'
This is the 'thermodynamic hypothesis of protein folding.' AlphaFold uses this dogma.
This is the 'thermodynamic hypothesis of protein folding.' AlphaFold uses this dogma.
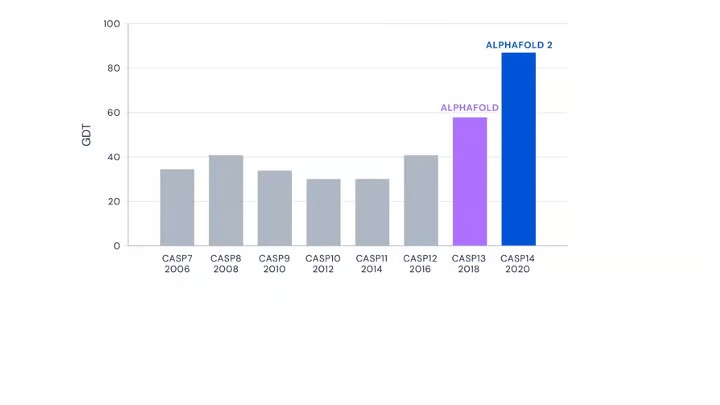
From the perspective of performance, AlphaFold2 (and this dogma) have cracked the likely structure of various proteins. However, it is well-accepted that this dogma may not hold true for all proteins.
The low-hanging fruit was picked. Some problems remain.

The low-hanging fruit was picked. Some problems remain.
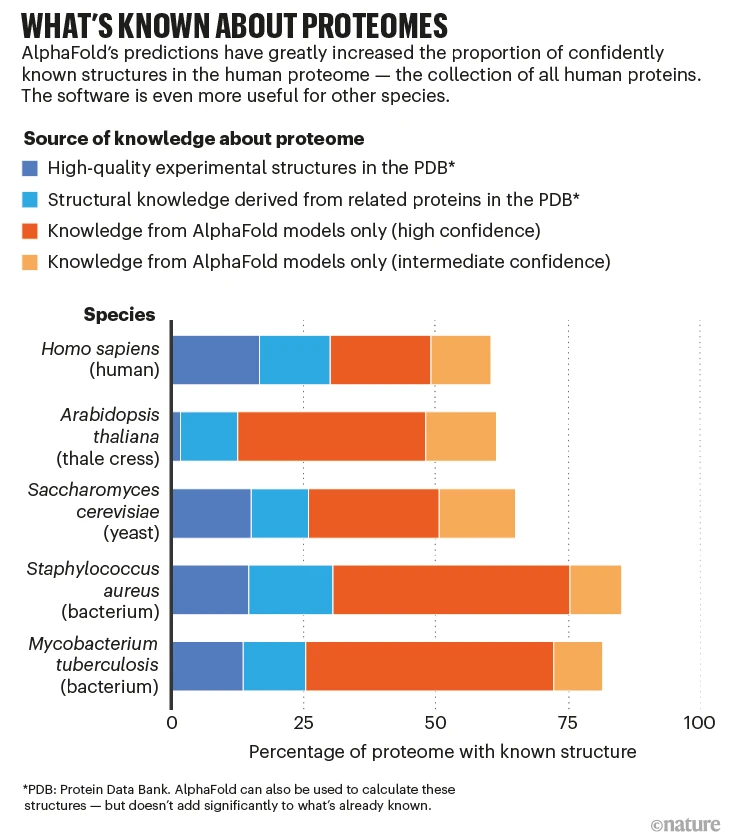

Folding doesn’t happen in empty space but in the bustling environment of a cell, where other molecules can influence the folding pathway. This cellular context adds another layer of complexity. As an example, let's consider molecular chaperones.
Not all proteins fold spontaneously; molecular chaperones assist in the folding of many proteins. These chaperones prevent misfolding and aggregation that can lead to complex diseases. 
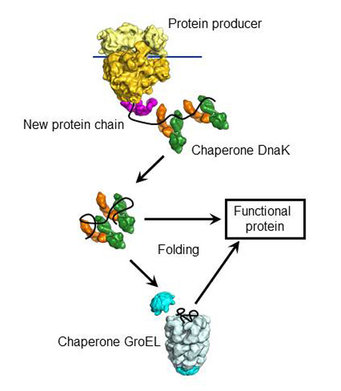
Solutions like AlphaFold model direct physicochemical interactions between amino acids to determine the most likely 3D structure of a protein but do not account for the cellular processes, like the action of chaperones, that can affect protein folding in vivo.
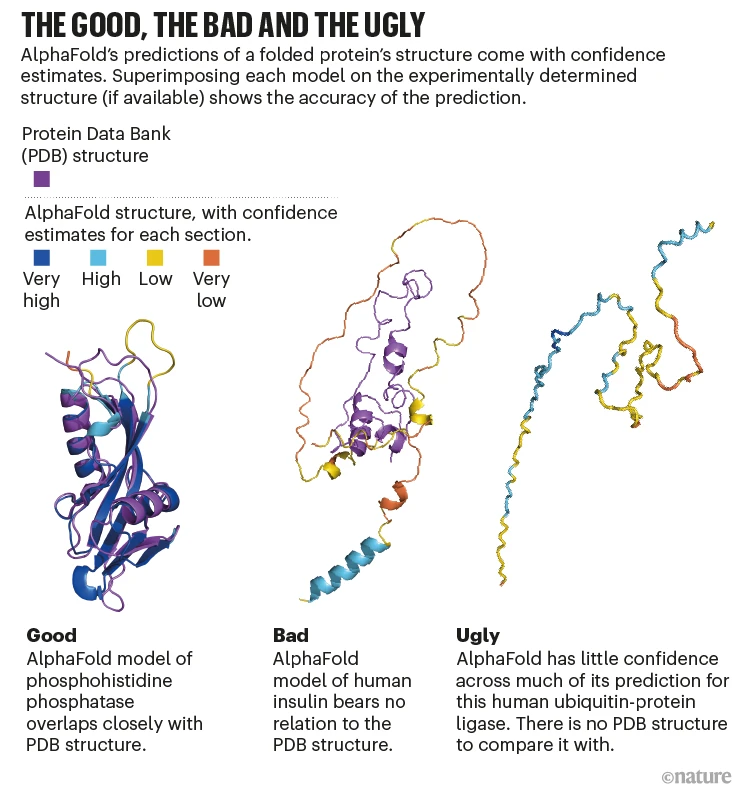
There is a lot to this problem (to be covered in other 🧵's), and the problem of protein-molecule, protein-protein, and protein-drug interactions, making the usage of AlphaFold2 in real-life scenarios difficult. The functional problem extends beyond a static training database.
The problem of predicting likely structures, assuming they are static and isolated, is solved. However, it is fair to say that the functional 'protein folding problem' is now solving protein complexes, based on interactions.
DeepMind's AlphaFold-Multimer, their protein complex solution, was not half as successful as their protein structure solution.
Protein complex prediction, in my opinion, bridges computational biochemistry and systems biology in unthought-of ways.
Protein complex prediction, in my opinion, bridges computational biochemistry and systems biology in unthought-of ways.
When it comes to solutions for drug discovery, understanding protein-drug interactions is a prerequisite. Essentially, here is an example of how solving a problem on paper is never enough in biology, and blackboxes might not necessarily work. This is also an example of technical constraints in data collection.
Observing protein folding in real time challenges even the most advanced scientific instruments, demanding ultra-fast and precise techniques to catch these fleeting processes; this, on top of in-vivo measurements within a cell being a grand, traditional challenge.
I will cover this topic in more detail over time, but tl;dr - protein structures? somewhat solved. protein complexes? we are so early.
Observing protein folding in real time challenges even the most advanced scientific instruments, demanding ultra-fast and precise techniques to catch these fleeting processes; this, on top of in-vivo measurements within a cell being a grand, traditional challenge.
I will cover this topic in more detail over time, but tl;dr - protein structures? somewhat solved. protein complexes? we are so early.
• • •
Missing some Tweet in this thread? You can try to
force a refresh




