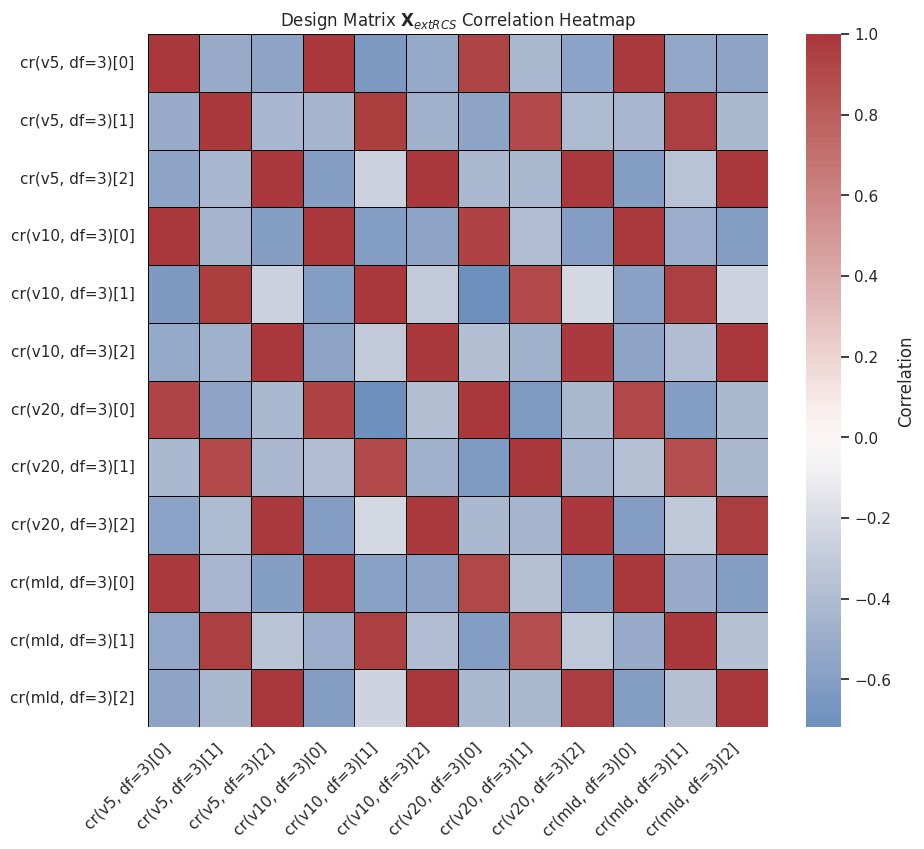
🧵 A new Nature paper on “spatial ecotypes” claims liquid biopsy can predict immunotherapy response. The tumor biology is cool. The statistics are a masterclass in how to do bad biomarker research. @f2harrell has a checklist for this. Let’s go. 1/
https://twitter.com/aaronnewmanlab/status/2052048106506813883
First, Harrell’s Biomarker Uncertainty Principle: “A molecular signature derived from high-dimensional data can be either parsimonious or predictive, but not both.” This paper tries to be both. It earns neither. 2/fharrell.com/post/badb
DICHOTOMANIA. They split 78 patients at the MEDIAN to draw KM curves. Harrell’s BBR shows median dichotomization reduces efficiency to 63.7% vs. continuous analysis — you need 158 patients to match what 100 give you. They burned 36% of their tiny cohort 3/
“Virtually all published cutpoints are analysis artifacts.” . The median isn’t biology. Add patients to one tail and the threshold moves. The tumor didn’t change. Your KM curves did. 4/hbiostat.org/bbr/info
Natura non facit saltus. Nature doesn’t make jumps. The median cutpoint assumes patients just above and just below are maximally different, while patients just above and far above are identical. This is not how SE levels work. Not how tumors work. Not how anything works. 5/
The “winning biomarker” problem. @f2harrell bootstrap example: the apparent winner of 213 markers had a 95% rank CI of [30–213]. Data could only rule out it being in the bottom 29. SE8 “wins” across 41 features here.
Nobody ran the bootstrap. Nobody showed those rank CIs. 6/
Nobody ran the bootstrap. Nobody showed those rank CIs. 6/
@f2harrell Harrell checklist item: “Pick a winning biomarker even though tiny changes in the sample result in a different winner.” SE8 beats SE7 overall. SE7 beats SE8 in melanoma alone. That’s not biology. That’s rank instability.! 7/fharrell.com/post/badb
@f2harrell Harrell checklist: “Validate using a sample too small or that should have been in training data.” Paired plasma-tumor validation: n=23. No CIs on the Spearman correlations shown. With n=23 those CIs span from “meaningless” to “strong.” That’s not a result. That’s a range. 8/
@f2harrell BBR chapter 18: dichotomizing a continuous outcome can require 5x the sample size for equivalent power. This paper dichotomizes the predictor (high/low SE) AND the outcome (responder/non-responder). Double dichotomania. 9/
@f2harrell Harrell: “Touting a new biomarker while ignoring basic clinical info that may be more predictive.” TCGA survival models here adjust for age and sex only. No stage. No performance status. SE7 beats TMB! Did it beat a properly specified clinical model? We will never know. 10/
@f2harrell Liquid EcoTyper trained on SIMULATED cfDNA — clean math mixtures of tumor DNA + healthy plasma. Harrell: “Live within the confines of the information content of the data.” A simulation’s info content is not real plasma’s. Real plasma has biology that hates your assumptions. 11/
@f2harrell The abstract: “implications for improved risk stratification and therapy personalization.” The data: n=78, median split, simulated training, no calibration curves, no PH check, no rank CIs, adjusts for age/sex only. One of these is a Nature paper. The other is the truth. 12/
@f2harrell Harrell’s bad biomarker checklist: don’t have a stat analysis plan; categorize continuous variables; overstate predictive utility; avoid checking absolute accuracy. This paper hits them all. 13/13 /end
• • •
Missing some Tweet in this thread? You can try to
force a refresh