🧵Another continuing thread branch
1/ 2021 PLOS Pathogens (Henan Univ + Western Univ of Health Sciences + Univ Kansas Med Ctr): hantaviruses use host factor **P58IPK** to counter PKR defenses. (1)
1/ 2021 PLOS Pathogens (Henan Univ + Western Univ of Health Sciences + Univ Kansas Med Ctr): hantaviruses use host factor **P58IPK** to counter PKR defenses. (1)
https://x.com/CaffeenHeadache/status/2052420977972355281
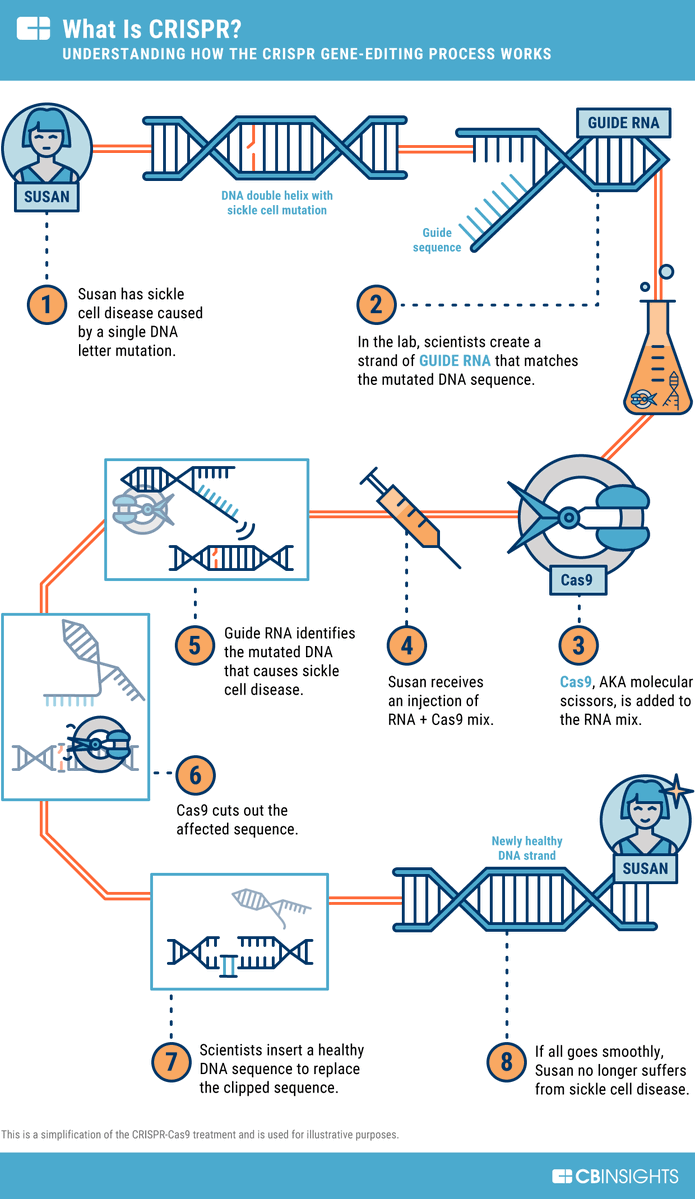
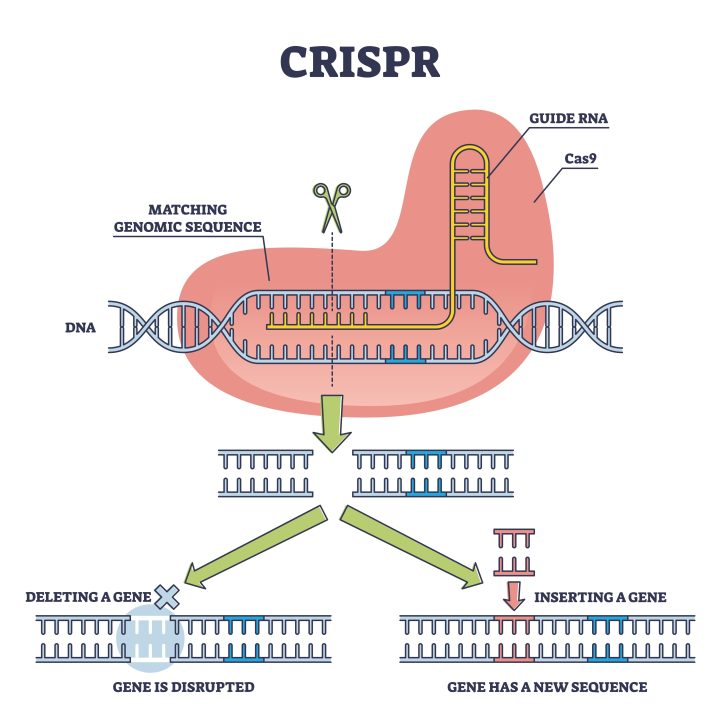
2/ China has a growing “CRISPR + hantavirus” literature—but it’s mainly **diagnostics**:
CRISPR-Cas systems used to detect hantavirus RNA quickly/sensitively.
Useful for surveillance, but it’s not editing the virus genome. Example: 2025 J Med Virol CRISPR/Cas12a work. (2)
CRISPR-Cas systems used to detect hantavirus RNA quickly/sensitively.
Useful for surveillance, but it’s not editing the virus genome. Example: 2025 J Med Virol CRISPR/Cas12a work. (2)
3/ **Reverse genetics** is closest to “editing hantavirus”:
Recover virus from cloned genome segments. Milestone: “Rescue of Hantaan virus minigenomes” (2003). A 2023 JID review notes the field long lacked full-length clone systems—so true genome-editing papers are scarce. (3)(4)
Recover virus from cloned genome segments. Milestone: “Rescue of Hantaan virus minigenomes” (2003). A 2023 JID review notes the field long lacked full-length clone systems—so true genome-editing papers are scarce. (3)(4)
4/ Citations:
(1) journals.plos.org/plospathogens/…
(2) onlinelibrary.wiley.com/doi/10.1002/jm…
(3) sciencedirect.com/science/articl…
(4) academic.oup.com/jid/article/22…
(1) journals.plos.org/plospathogens/…
(2) onlinelibrary.wiley.com/doi/10.1002/jm…
(3) sciencedirect.com/science/articl…
(4) academic.oup.com/jid/article/22…
5/ We are giving you the breadcrumbs in hopes of experts connecting the dots with their extensive relevant knowledge and credibility.
We looked for hantavirus + CRISPR work with American and Chinese scientists since 1993.
The strongest public match is not edited hantavirus — it is CRISPR editing of host genes / animal models to study how hantavirus enters cells.
We looked for hantavirus + CRISPR work with American and Chinese scientists since 1993.
The strongest public match is not edited hantavirus — it is CRISPR editing of host genes / animal models to study how hantavirus enters cells.
6/ The key 2018 Nature paper identified PCDH1 as an entry factor for Andes & Sin Nombre hantaviruses.
U.S. labs included Einstein, USAMRIID, and Utah State; Chinese-born USU scientist Zhongde Wang co-led the work, with Rong Li in the Wang lab.
U.S. labs included Einstein, USAMRIID, and Utah State; Chinese-born USU scientist Zhongde Wang co-led the work, with Rong Li in the Wang lab.

7/ The CRISPR piece was on the host side:
USU says its team used CRISPR/Cas9 to create the first PCDH1-knockout hamster model. Removing that receptor made hamsters largely resistant to Andes virus lung injury.
USU says its team used CRISPR/Cas9 to create the first PCDH1-knockout hamster model. Removing that receptor made hamsters largely resistant to Andes virus lung injury.

8/ A 2023 follow-up pushed further: researchers used CRISPR/Cas9 genome engineering to introduce two PCDH1 mutations in hamsters, mapping a receptor surface used by hantavirus glycoproteins.
Still: host receptor editing, not editing hantavirus itself.
Still: host receptor editing, not editing hantavirus itself.

9/ Citations:
2018 Nature PCDH1 paper: nature.com/articles/s4158…
USU CRISPR/hamster summary: usu.edu/today/story/st…
Zhongde Wang bio: usu.edu/today/story/fr…
2023 Nat Comm follow-up: nature.com/articles/s4146…
eLife CRISPR receptor-depletion near-miss: elifesciences.org/articles/69708
China-only CRISPR diagnostic example: sciencedirect.com/science/articl…
2018 Nature PCDH1 paper: nature.com/articles/s4158…
USU CRISPR/hamster summary: usu.edu/today/story/st…
Zhongde Wang bio: usu.edu/today/story/fr…
2023 Nat Comm follow-up: nature.com/articles/s4146…
eLife CRISPR receptor-depletion near-miss: elifesciences.org/articles/69708
China-only CRISPR diagnostic example: sciencedirect.com/science/articl…
@threadreaderapp unroll please
• • •
Missing some Tweet in this thread? You can try to
force a refresh