We've created an interactive website to visualize >100,000 experimental measurements of how mutations to #SARSCoV2 RBD affect binding by antibodies & sera: jbloomlab.github.io/SARS2_RBD_Ab_e… Explore it to examine a wealth of information about the antigenic effects of viral mutations. (1/n)
Over the last 9 months, the indefatigable @tylernstarr & @AllieGreaney have used deep mutational scanning to measure how the 2,304 RBD mutations tolerated for protein folding / ACE2 binding affect recognition by 50 antibodies / sera. Data scattered across multiple papers. (2/n)
We have consolidated these data so they can be explored to understand antigenic impacts of mutations observed during genomic surveillance. Best way to look at data is to explore the website at jbloomlab.github.io/SARS2_RBD_Ab_e…, but here are some static-image summaries: (3/n)
First, we used experimental measurements to organize antibodies in the space of "viral escape." This organization mostly concordant with structural classification scheme of @cobarnes27 @bjorkmanlab; you can click on specific antibody to see its sites of escape mutations. (4/n) 

For any subset of antibodies, you can visualize mean antigenic effect of mutations at each RBD site. It's striking how peaks in this plot include so many sites of emerging mutations: E484, L452, K417, R346, etc. (5/n) 

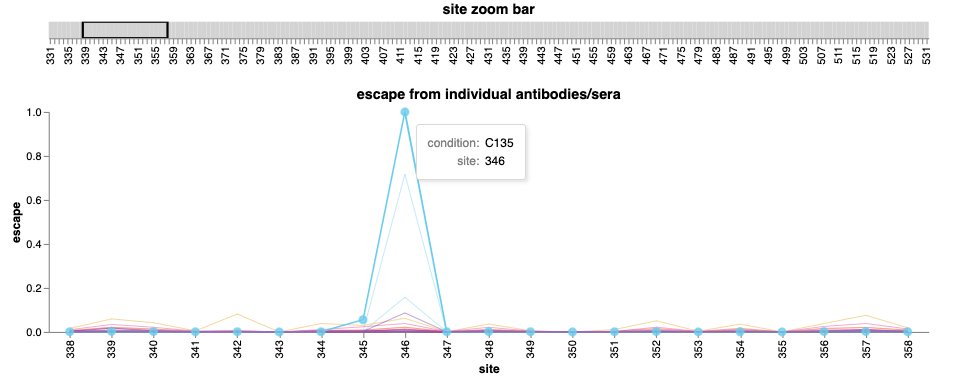
You can also zoom in on sites of interest to see which antibodies/sera are affected by mutations there. For instance, @Tuliodna recently reported a new lineage with a mutation at R346: mutations there impact several class 3 antibodies such as C135. (6/n) 
You can also select specific antibodies to see their binding escape mutations. For instance, LY-CoV555 (bamlanivimab) is unfortunately affected by mutations at E484 and L452, which is why US government recently halted distribution of this therapeutic antibody. (7/n) 

You can also visualize sera binding-escape mutations. For instance, below shows mutations that reduce binding by convalescent sera from @HelenChuMD's HAARVI cohort are most similar to ones (eg, E484K) that affect class 2 antibodies. (8/n) 

We've also included a detailed technical explanation of data, link to CSV file with raw data for all 100,000+ measurements (raw.githubusercontent.com/jbloomlab/SARS…), and GitHub repo with code to make visualizations (github.com/jbloomlab/SARS…). (9/n)
In addition to scientists tagged above, this work derives from generous sharing/help from many collaborators including @seth_zost @VUMC_Vaccines @AdamDingens @DrJLi @Dr_MChoudhary @NussenzweigL @c_gaebler @PaulBieniasz @theodora_nyc (10/n)
Finally, the interactive visualizations are all built with the amazing altair-viz.github.io plotting package created by @jakevdp (11/n)
• • •
Missing some Tweet in this thread? You can try to
force a refresh