Today we release in @biorxivpreprint a pair of related methods, DNA Ticker Tape & ENGRAM, for multiplex, temporally ordered genomic recording of cell lineage histories, enhancer & signal transduction activities, respectively led by the brilliant @_Choi_Junhong & @wchenomics 1/25 

Links to preprints on @biorxivpreprint:
DNA Ticker Tape: tinyurl.com/vdummmww
ENGRAM: tinyurl.com/2784mhkh
Let’s start w/ DNA Ticker Tape, by @_Choi_Junhong, a general system for pseudo-processive genome editing that can "write" the ORDER of molecular events. 2/25
DNA Ticker Tape: tinyurl.com/vdummmww
ENGRAM: tinyurl.com/2784mhkh
Let’s start w/ DNA Ticker Tape, by @_Choi_Junhong, a general system for pseudo-processive genome editing that can "write" the ORDER of molecular events. 2/25
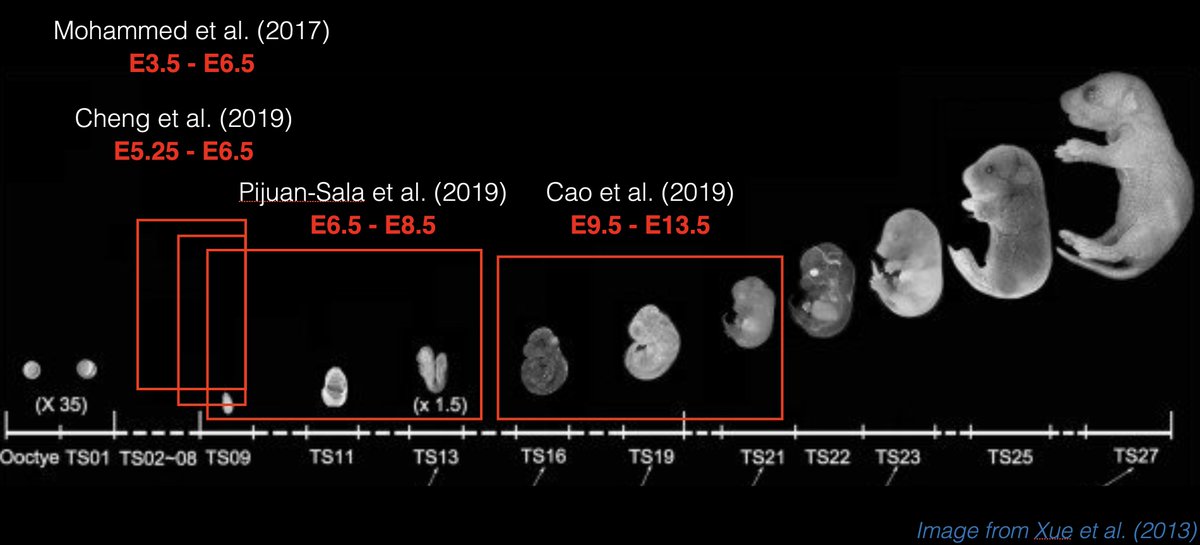
For background on our motivation, in 2016, w/ @schierlab, we developed GESTALT, a genome editing-based method for cell lineage tracing that we applied to zebrafish development; others have subsequently developed & applied similar methods in mouse, cancer & other systems. 3/25 
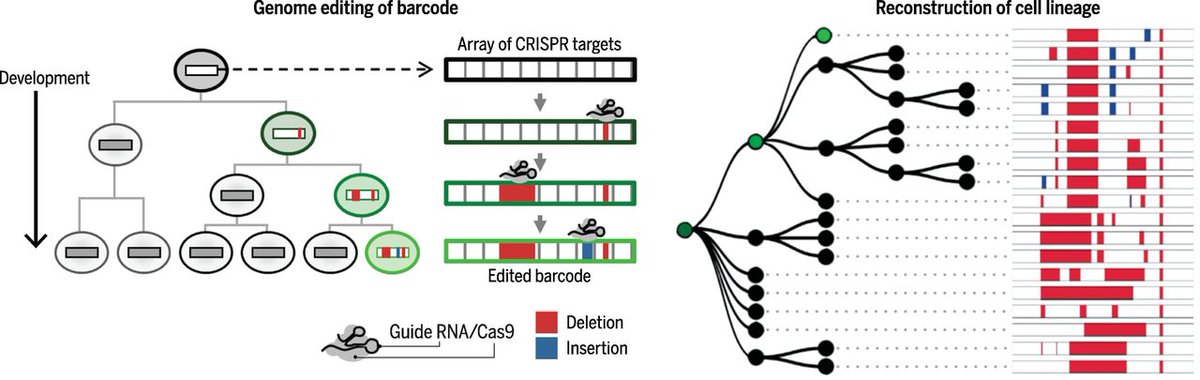
However, GESTALT was limited by: 1) a failure to explicitly record the ORDER of edits, which renders phylogenetic reconstruction very, very challenging; 2) a reliance on cytotoxic DSBs and unpredictable NHEJ; 3) recording only lineage & not other biological information. 4/25
DNA Ticker Tape solves these challenges. Blank tape = tandem array of 14-bp monomer. These are partial genome editing targets, all but first of which are 5’ truncated. Prime editing (h/t @davidrliu!) of ‘active site’ inserts informative barcode, but also activates next site! 5/25
As the active site or “write head” moves along the monomeric array, the order in which barcodes were written to DNA is simply their linear order! Although prime editing efficiency is issue, we’ve recorded as many as 14 consecutive events onto a single tape (w/ PE2). 6/25
To demonstrate the method, we performed serial transfections w/ 16 pegRNAs (encoding different barcodes) & asked if we could infer their order out of 16! (20 trillion) possibilities by sequencing the ticker tape. Indeed, we could! We can also deconvolve more complex patterns 7/25 
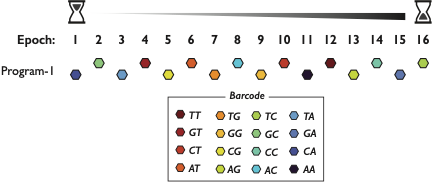
To further show that DNA Ticker Tape constitutes a digital system (information represented by the content and order of discrete symbols), we asked whether we could record and recover short text messages. First, we constructed an alphabet conversion based on a 3 bp code. 8/25 
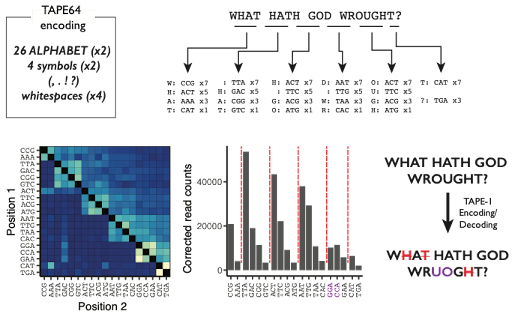
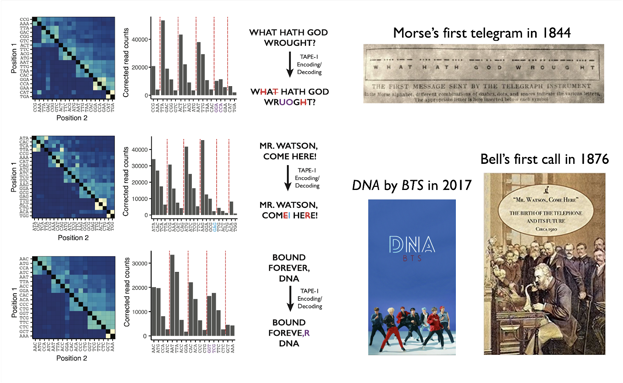
Within genomes of living, mammalian cells, we recorded & recovered short text msgs from Samuel Morse, Alexander Graham Bell & K-Pop group #BTS. Info in barcode order & abundance. Some errors but not too shabby. K-Pop lyrics from “DNA” translated to English by @_Choi_Junhong 9/25 
Returning to our original motivation, we also show that we can record lineage histories to genomic DNA in manner much more straightforward to interpret than GESTALT & similar methods, because order of stochastic edits is explicitly represented 5’ to 3’ along DNA Ticker Tape 10/25 
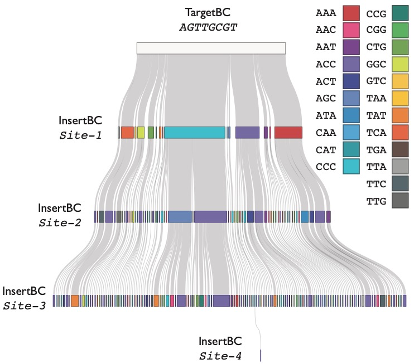
In summary, as a pseudo-processive system for inscribing discrete "symbols" to DNA in an ordered fashion, DNA Ticker Tape is essentially an artificial “writing system” that is operational within living mammalian cells. 11/25 

OK, now that we have an in vivo writing system, what do we want to write about? Let’s turn to ENGRAM (ENhancer-driven Genomic Recording of transcriptional Activity in Multiplex), a signal conversion system developed by @wchenomics. 12/25
ENGRAM is a method that enables the multiplex, DNA-based recording of the strength, duration and, potentially, the order of multiple enhancer and signal transduction activities (as well as anything else one might convert to Pol-2 mediated transcription) 13/25 

Other DNA recorders rely on coupling a signal of interest to Pol-2 mediated activation of enzyme (e.g. Cre, Cas9, BE) that mediates specific change to a DNA target sequence. But this doesn’t multiplex well. One would need a different enzyme for each signal of interest. 14/25
Other DNA recorders rely on coupling a signal of interest to Pol-2 mediated activation of enzyme (e.g. Cre, Cas9, BE) that mediates specific change to a DNA target sequence. But this doesn’t multiplex well. One would need a different enzyme for each signal of interest. 15/25 

To address this, we leveraged Csy4-based guide release, which allows functional pegRNAs to be released from Pol-2 transcripts. ENGRAM 2.0 designs implement various strategies for suppressing background. 16/25 
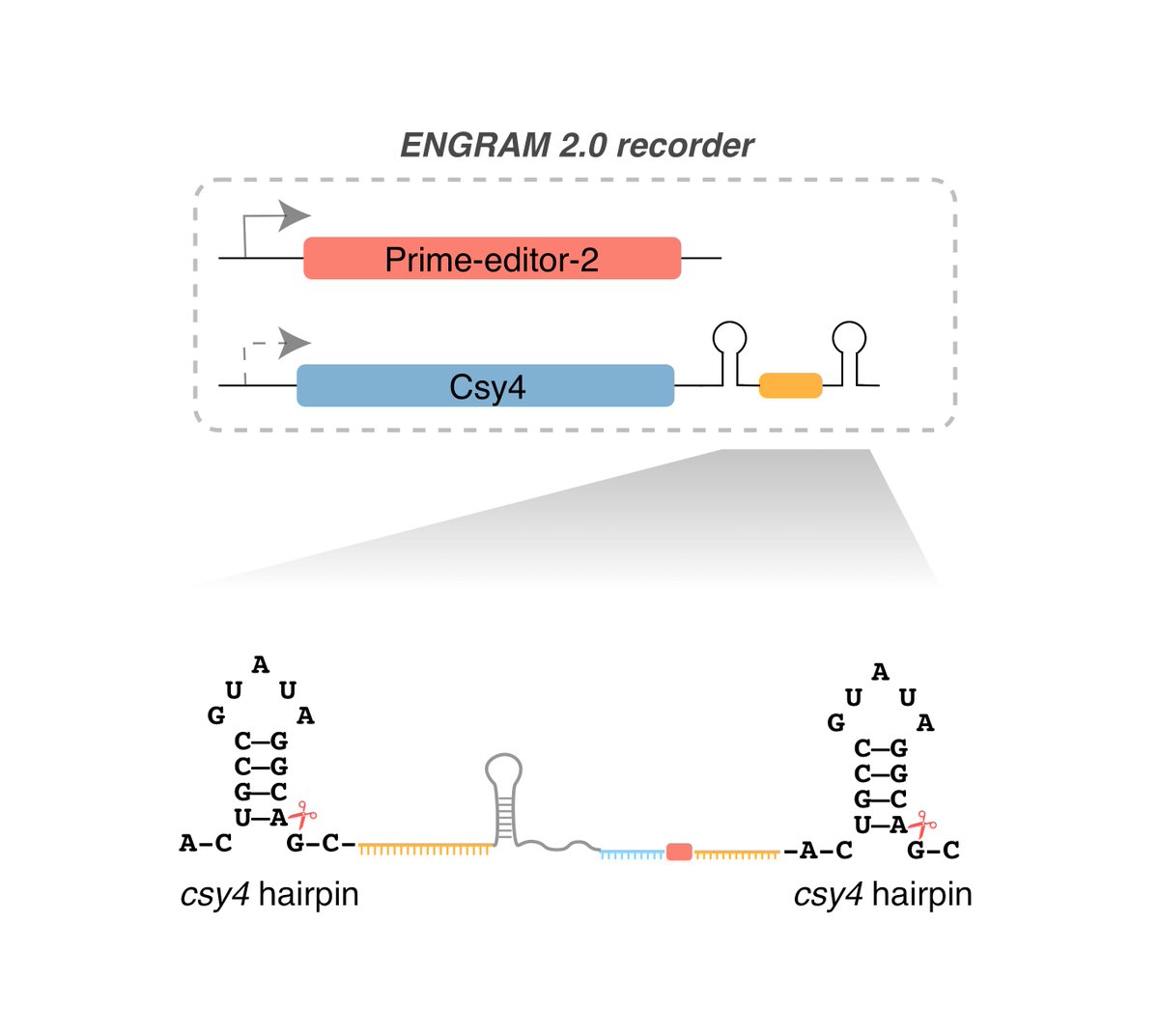
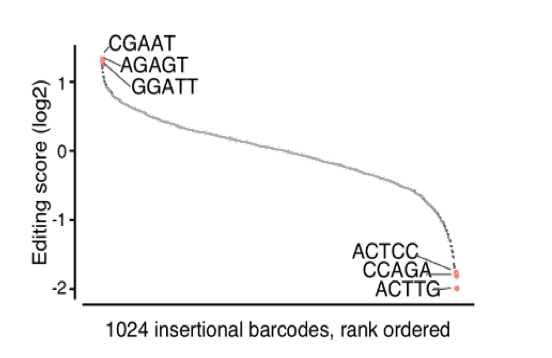
Coupling libraries of enhancers to ENGRAM 2.0, we can “record” via enhancer-specific insertional barcodes , analogous to MPRAs, except we are quantitatively writing their activities to genomic DNA, rather than reading from RNA. ENGRAM & MPRA results are well correlated 17/25 
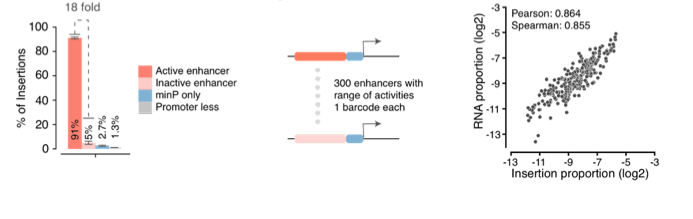
OK, great, but with an eye to applying this to developmental biology, can we also record the activity of signal transduction pathways? To test this, we constructed ENGRAM 2.0 recorders for TetON, NF-KB and Wnt signaling. 18/25 
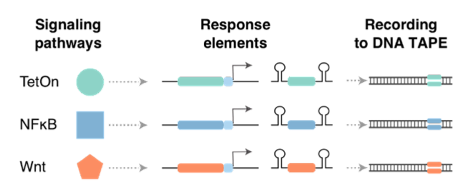
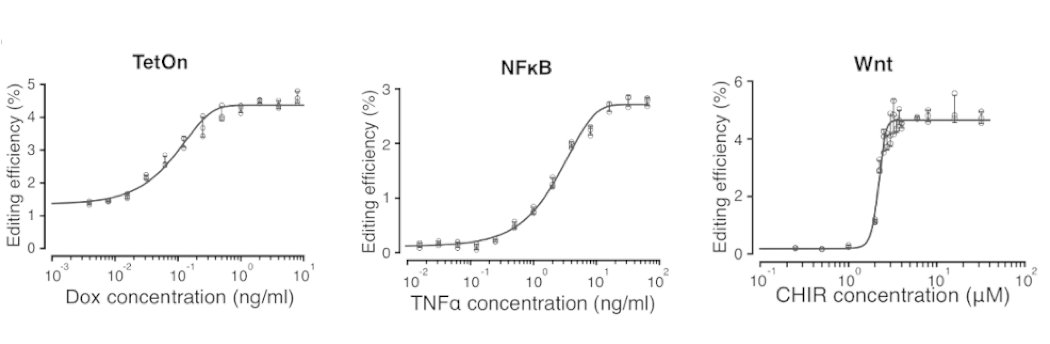
For each of these signal-responsive ENGRAM recorders, genome editing of DNA Tape with signal-specific barcodes exhibited a strikingly sigmoidal dependence on the log[conc.] of the corresponding stimulant. Dynamic ranges up to 3, 19 & 27-fold for Tet, NF-KB and Wnt recorders 19/25 
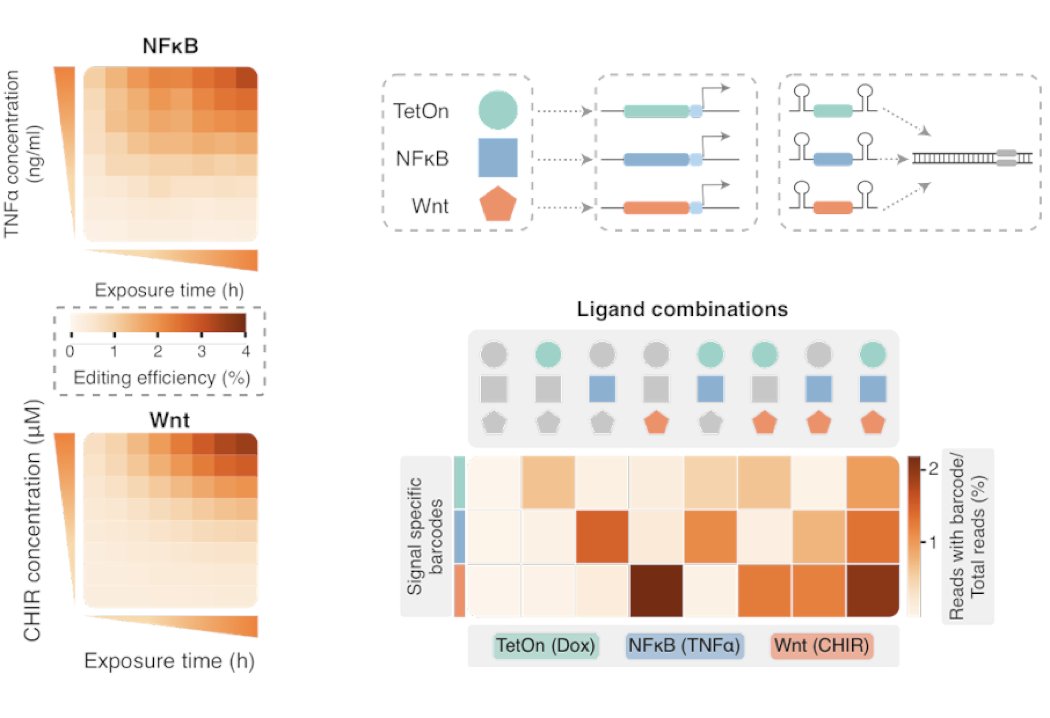
NF-KB & Wnt ENGRAM recorders are sensitive to both the intensity and duration of stimulation. When multiple signal-responsive recorders are deployed together, they exhibit minimal cross-talk, consistent with expectation that these signaling pathways are largely orthogonal 20/25 
In summary, ENGRAM is a method that enables the multiplex conversion of biological signals, including but not limited to enhancer and signal transduction pathway activities, to signal-specific barcodes that are written to a DNA Tape. 21/25 
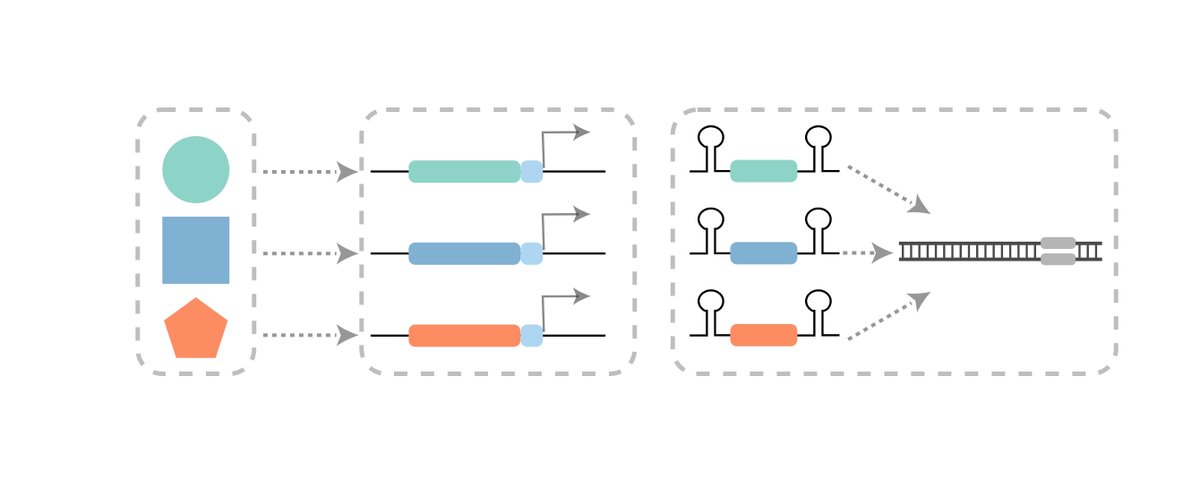
Although still in progress, the combination of DNA Ticker Tape and ENGRAM has the potential to enable the highly multiplex, temporally ordered, in vivo recording of diverse classes of biological information to genomic DNA. 22/25
ENGRAM and DNA Ticker Tape were developed by @wchenomics and @_Choi_Junhong, respectively, who have built up quite a synergy. It’s impossible to capture in a tweet what an adventure with them this has been. 23/25 

Many thanks to co-authors on one or both papers, including @Anna_Minkina @FloChardon @Chase_C_Suiter @SRegalad0 @sdomcke @Nobu_Hamazaki Charlie Lee @bethkarenmartin Riza Daza @vagar112 @evaknichols & Anh Leith! 24/25
And of course thank you to our funders and institutions, as well, especially @AllenFrontiers, @HHMINEWS, @UWMedicine, @uwgenome, @BrotmanBaty, @DamonRunyon. Finis! 25/25 

• • •
Missing some Tweet in this thread? You can try to
force a refresh