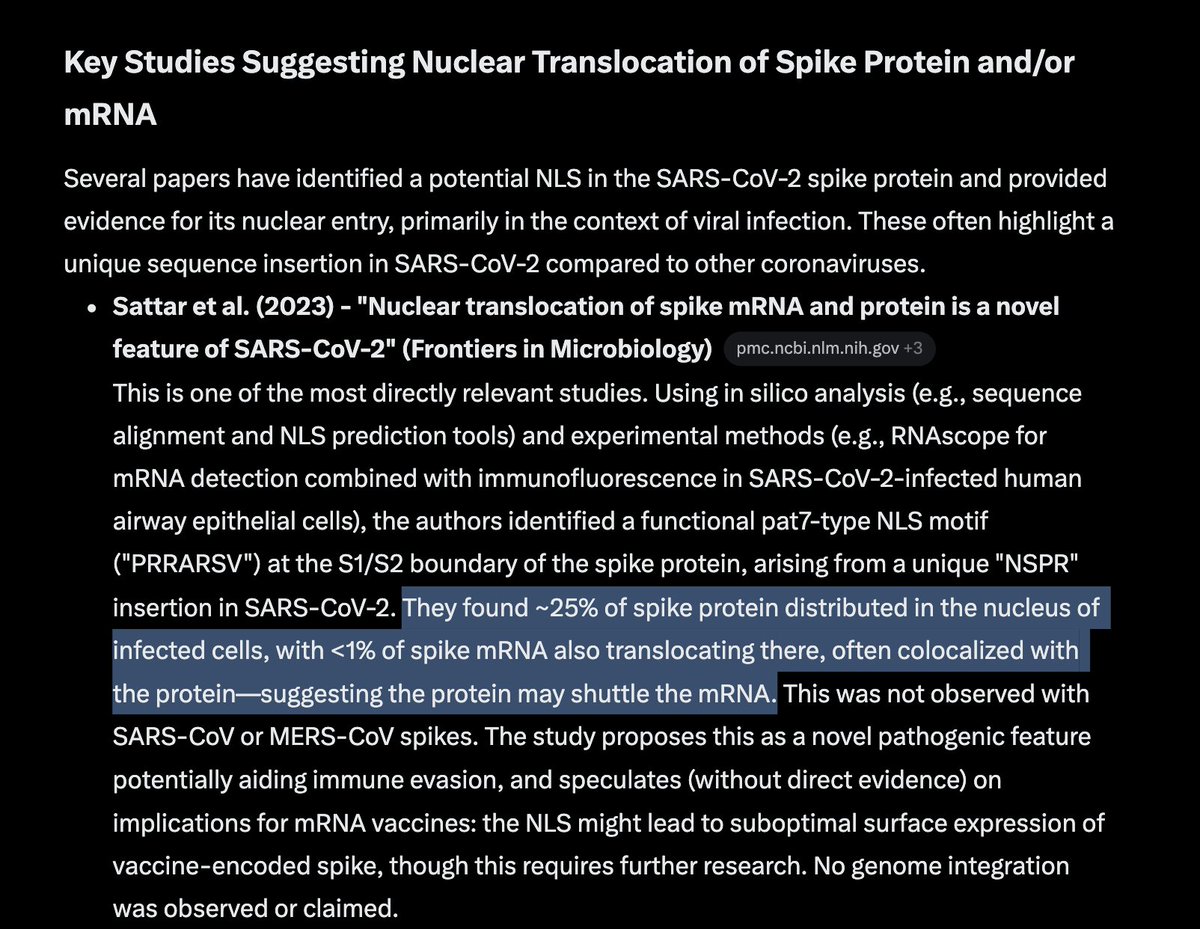
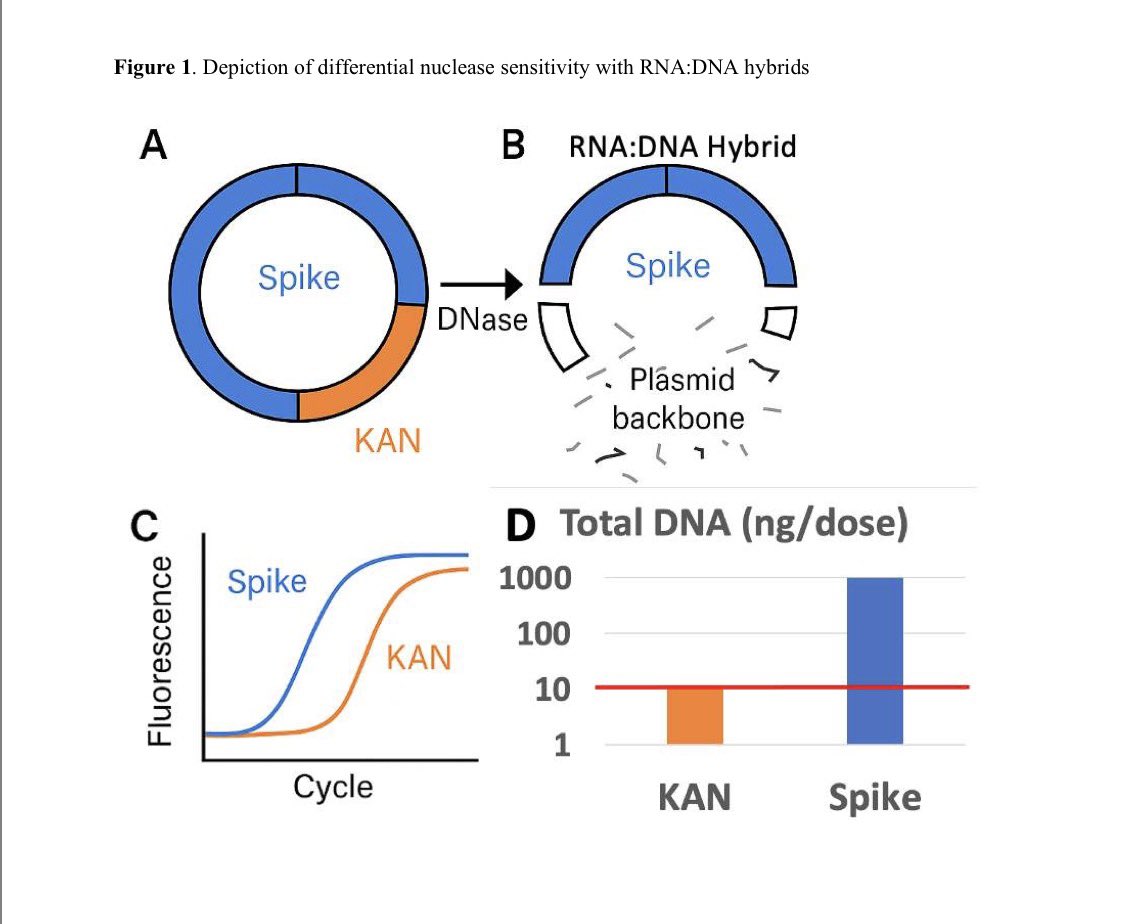
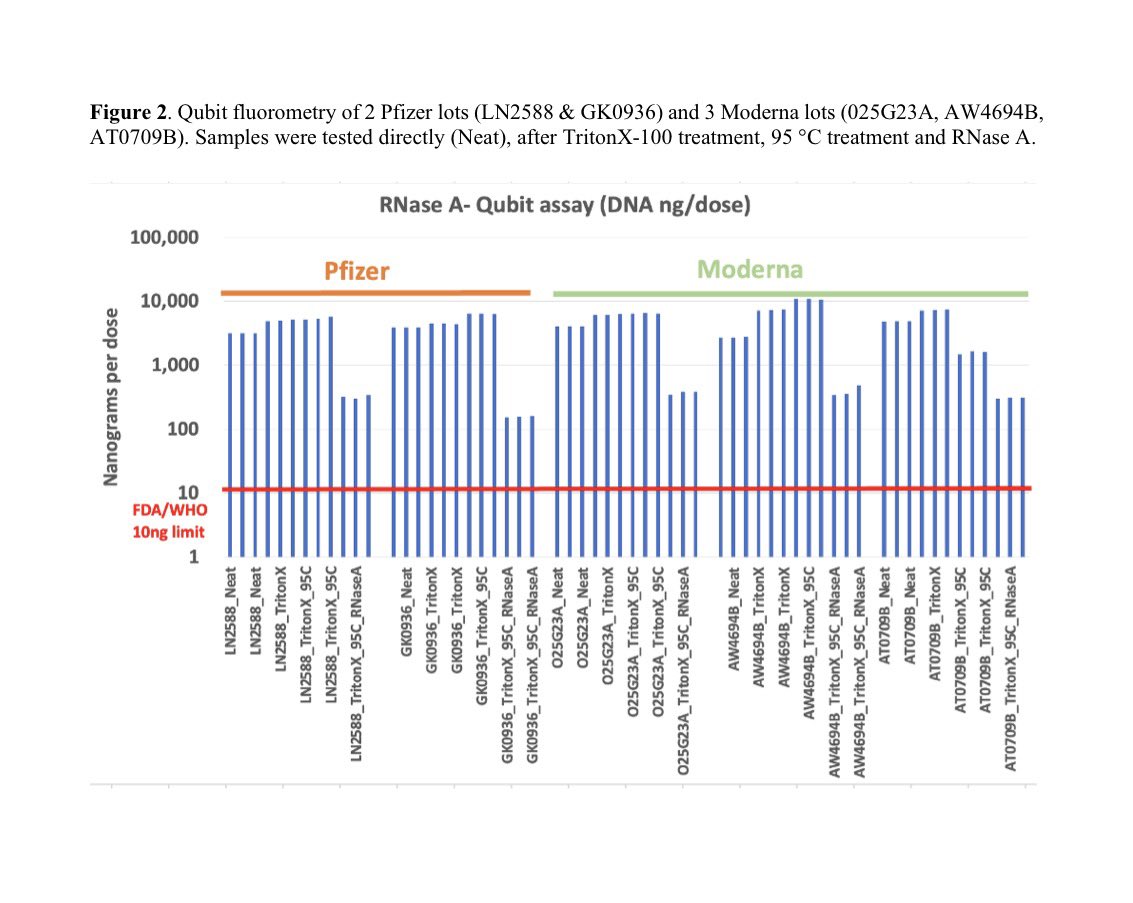
Of the many mutations in omicron the most interesting two are in the SEB domain.
Staphylococcus Enterotoxin B is a peptide that is classified as a bioweapon.
When ingested it creates symptoms very similar to COVID and it’s know to trigger cytokine storms
Staphylococcus Enterotoxin B is a peptide that is classified as a bioweapon.
When ingested it creates symptoms very similar to COVID and it’s know to trigger cytokine storms

It’s well described in Cheng et al.
This known toxin was warp sped into spike only Vaccines.
SEB is directly adjacent to the FCS.
Let’s have a look at what Ahanotu et al has to say about SEB exposure and it’s capacity as a bioweapon.
pnas.org/content/117/41…
This known toxin was warp sped into spike only Vaccines.
SEB is directly adjacent to the FCS.
Let’s have a look at what Ahanotu et al has to say about SEB exposure and it’s capacity as a bioweapon.
pnas.org/content/117/41…
Exposure to these peptides has been known to cause cytokine storms.
They are superantigens.
liebertpub.com/doi/pdfplus/10…
They are superantigens.
liebertpub.com/doi/pdfplus/10…

Now look at Omicron spike protein 671-692.
Two amino acid changes and one is the infamous proline change near the FCS. Prolines are right angle brackets in proteins. When they change, they alter structure.
P681H
N679K
This may attenuate the SEB toxicity. Fewer Ckine storms.
Two amino acid changes and one is the infamous proline change near the FCS. Prolines are right angle brackets in proteins. When they change, they alter structure.
P681H
N679K
This may attenuate the SEB toxicity. Fewer Ckine storms.
Some of this is discussed in our preprint.
I was not aware of these SEB mutations in omicron when I wrote this.
I was C19 +ve when I submitted this, ironically.
1K downloads…
osf.io/bcsa6/
I was not aware of these SEB mutations in omicron when I wrote this.
I was C19 +ve when I submitted this, ironically.
1K downloads…
osf.io/bcsa6/
It is early with Omicron so this is speculation. When the dust settles, I’d like to know if there are any cases of cytokine storms with this variant.
https://twitter.com/Covid19Crusher/status/1469786948000694274
Some critiques of the Cheng paper
A) lack of replication
B) in silico modeling of short peptides
Until october 2021 where Brogna et al find similar peptides circulating in the blood and urine of C19 patients.
@federicolois sent me this..
f1000research.com/articles/10-550
A) lack of replication
B) in silico modeling of short peptides
Until october 2021 where Brogna et al find similar peptides circulating in the blood and urine of C19 patients.
@federicolois sent me this..
f1000research.com/articles/10-550

Of note, They find a pango-lineage (B.1.596 - not to be confused with Omicron B.1.1.529) that has these mutations that would make for higher homology to the toxins.
This lineage broke out in the US and Kenya. It didn't get very far.
cov-lineages.org/lineage.html?l…
This lineage broke out in the US and Kenya. It didn't get very far.
cov-lineages.org/lineage.html?l…
You can see its prevalence in blue. Delta replaced it.
Wonder if these were sicker cases? Anyone have IFR by Pango-lineage? Is there even any signal with all the confounders?
We keep debating the IFR of the most prevalent strains but should we?
outbreak.info/location-repor…
Wonder if these were sicker cases? Anyone have IFR by Pango-lineage? Is there even any signal with all the confounders?
We keep debating the IFR of the most prevalent strains but should we?
outbreak.info/location-repor…
The most prevalent strains are unlikely to be the most lethal. Note the Proline mutation in question is flagged (yellow) as Variants Of Interest (VOI) (P681H). As is N679K. Both are are not observed in the other lineages on display. 
Cheng et al followed up with a paper in Cell showcasing an antibody that blocked SEB/FCS.
cell.com/structure/full…
cell.com/structure/full…

Look at how sensitive the FCS codons are. By using the rarest codon in human cells, stack two rare events next to each other and you get ribosomal pausing which in turn is required for spike to fold properly.
Now imagine changing all the codons.
mdpi.com/1422-0067/22/1…
Now imagine changing all the codons.
mdpi.com/1422-0067/22/1…
Physician on the Ground in South Africa is not seeing cytokine storms.
@WoodReporting
podcasts.apple.com/us/podcast/tri…
@WoodReporting
podcasts.apple.com/us/podcast/tri…
Preprint on the SEB domain and it’s importance to virulence.
Authors never mention SEB.
Seems to be a zoonosis pimping paper failing to reference prior art and funded by Fauci.
bit.ly/3q9cebx
Authors never mention SEB.
Seems to be a zoonosis pimping paper failing to reference prior art and funded by Fauci.
bit.ly/3q9cebx
They should at least reference the Cheng et al paper that raised an antibody to this exact domain.
And RatG13 doesn’t shoe horn SEB into natural origins.
That’s a lab construct.
And RatG13 doesn’t shoe horn SEB into natural origins.
That’s a lab construct.

Some background on RatG13 odd origins.
https://twitter.com/stevenemassey/status/1464214480942997508
• • •
Missing some Tweet in this thread? You can try to
force a refresh