
Cannabis Genome Project,2011. SOLiD sequencer. R&D lead Human Genome Project at MIT/WIBR. Founder- Medicinal Genomics.
How to get URL link on X (Twitter) App


https://twitter.com/newsfromscience/status/2056578113622990979First time- Really?

 This paper tries to makes claims of RT-qPCR sensitivity with a LLOQ of 372,800 molecules/ml as their lowest limit of Quantiation.
This paper tries to makes claims of RT-qPCR sensitivity with a LLOQ of 372,800 molecules/ml as their lowest limit of Quantiation.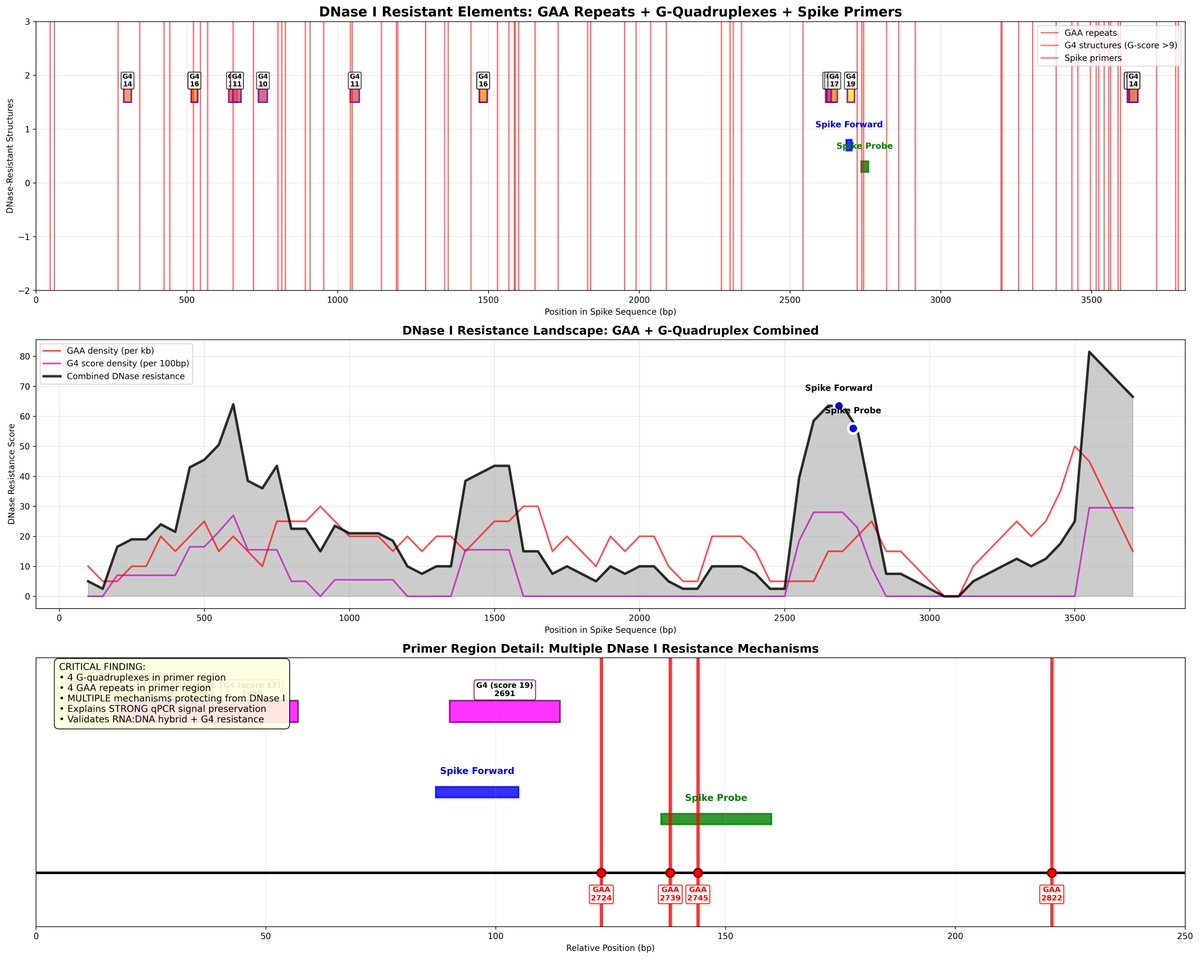
 Evans et al demonstrate DNaseI is resistant to quadruplex Gs. They also move to alternative enzymes.
Evans et al demonstrate DNaseI is resistant to quadruplex Gs. They also move to alternative enzymes. 


 journalofindependentmedicine.org/articles/v02n0…
journalofindependentmedicine.org/articles/v02n0…




 Here is the Rub.
Here is the Rub.

https://twitter.com/Kevin_McKernan/status/1997109939194511669
 Yup.
Yup.

 Why this matters-
Why this matters-

https://x.com/TFTC21/status/1994441655282458913?s=20

 HT @sudokuvariante for the file forensics.
HT @sudokuvariante for the file forensics. https://twitter.com/kevin_mckernan/status/1991889175659102452


 1)Turn your PDF into a PNG file
1)Turn your PDF into a PNG file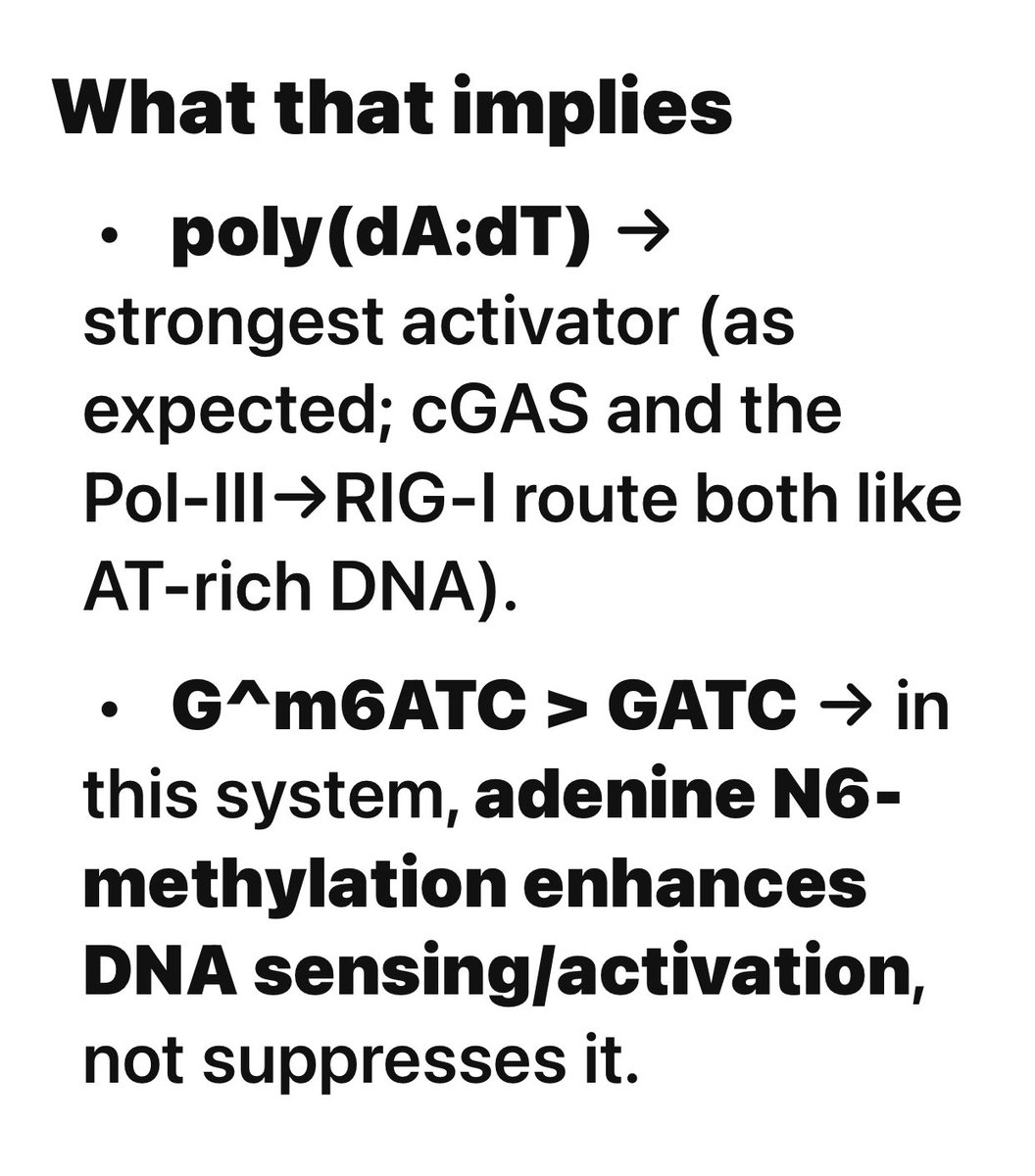
 @luijsterburglab @MaryanneDemasi @jeffreyatucker @TonyNikolic10 @ClareCraigPath @JesslovesMJK @dragonfishy @RWMaloneMD @weldeiry @Humanspective @DrEliDavid @kenjaques Note the asterisk and P values.
@luijsterburglab @MaryanneDemasi @jeffreyatucker @TonyNikolic10 @ClareCraigPath @JesslovesMJK @dragonfishy @RWMaloneMD @weldeiry @Humanspective @DrEliDavid @kenjaques Note the asterisk and P values.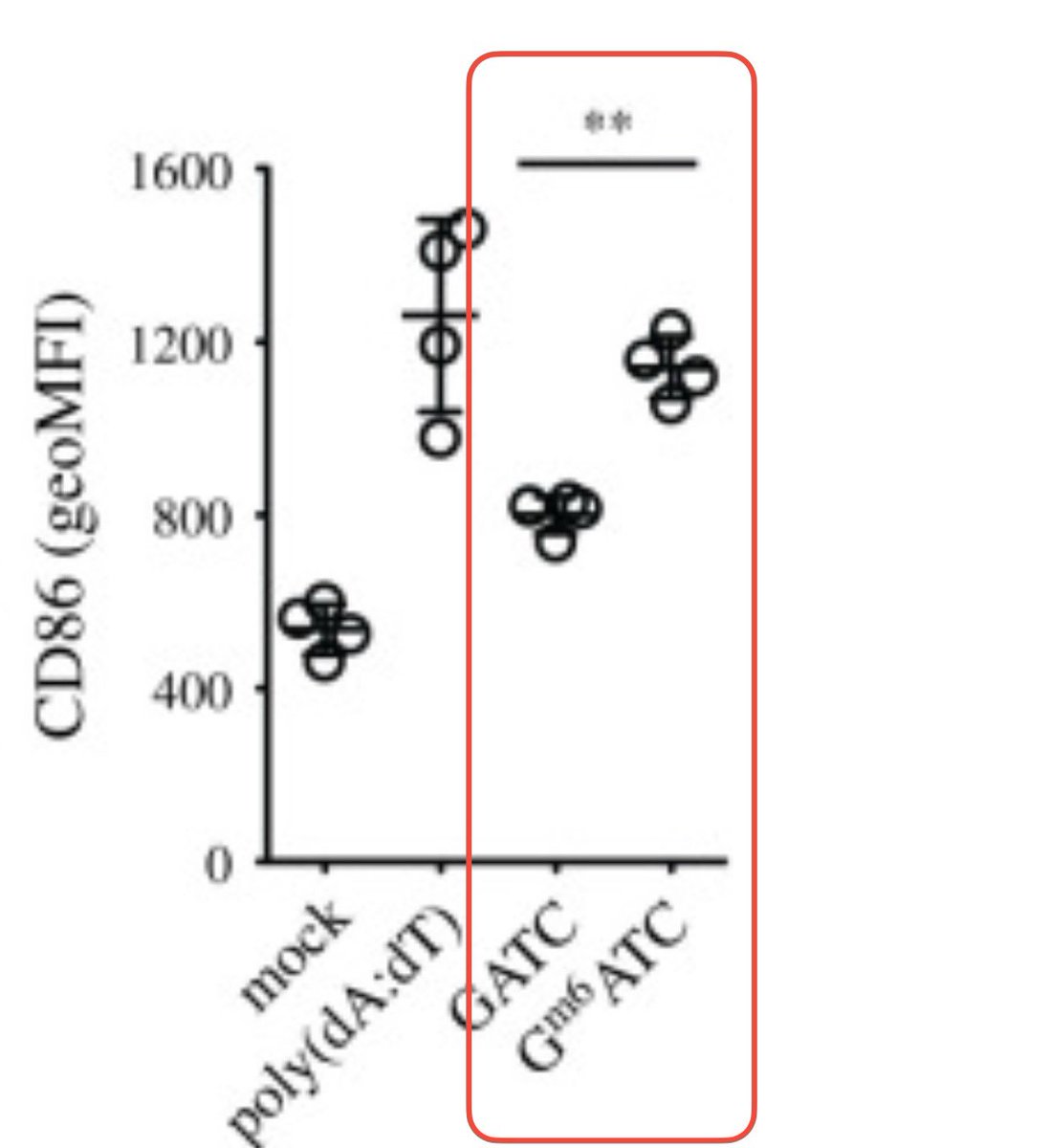

 Oxford Nanopore Sequencing reveals Pfizer didnt use a Dam knock out E.coli strain and the resulting plasmid DNA contamination is hyper stimulatory to the cGAS-STING pathway.
Oxford Nanopore Sequencing reveals Pfizer didnt use a Dam knock out E.coli strain and the resulting plasmid DNA contamination is hyper stimulatory to the cGAS-STING pathway. 
 You asked for it. This is an Algo experiment.
You asked for it. This is an Algo experiment.
