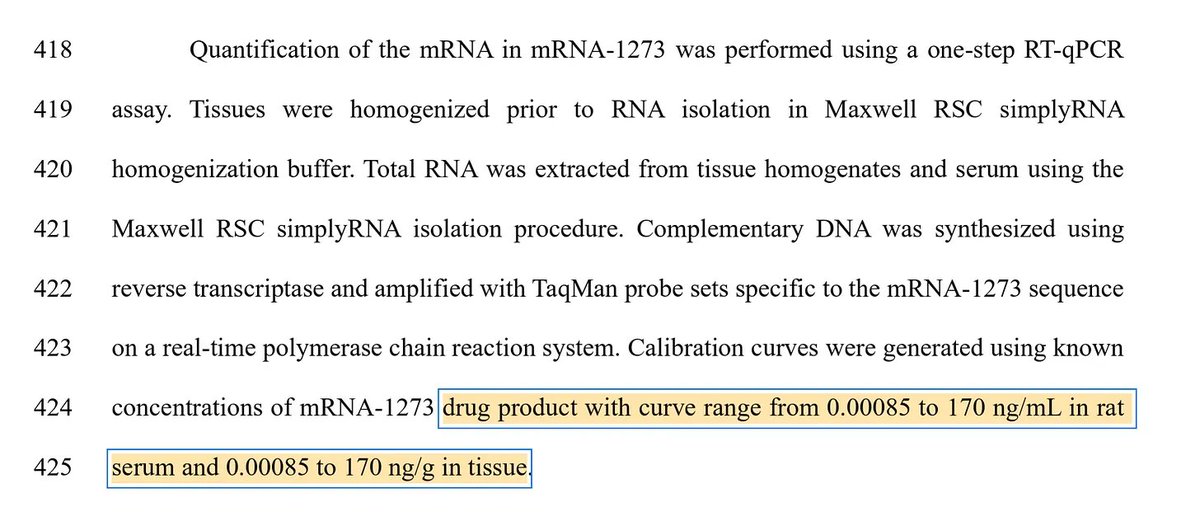
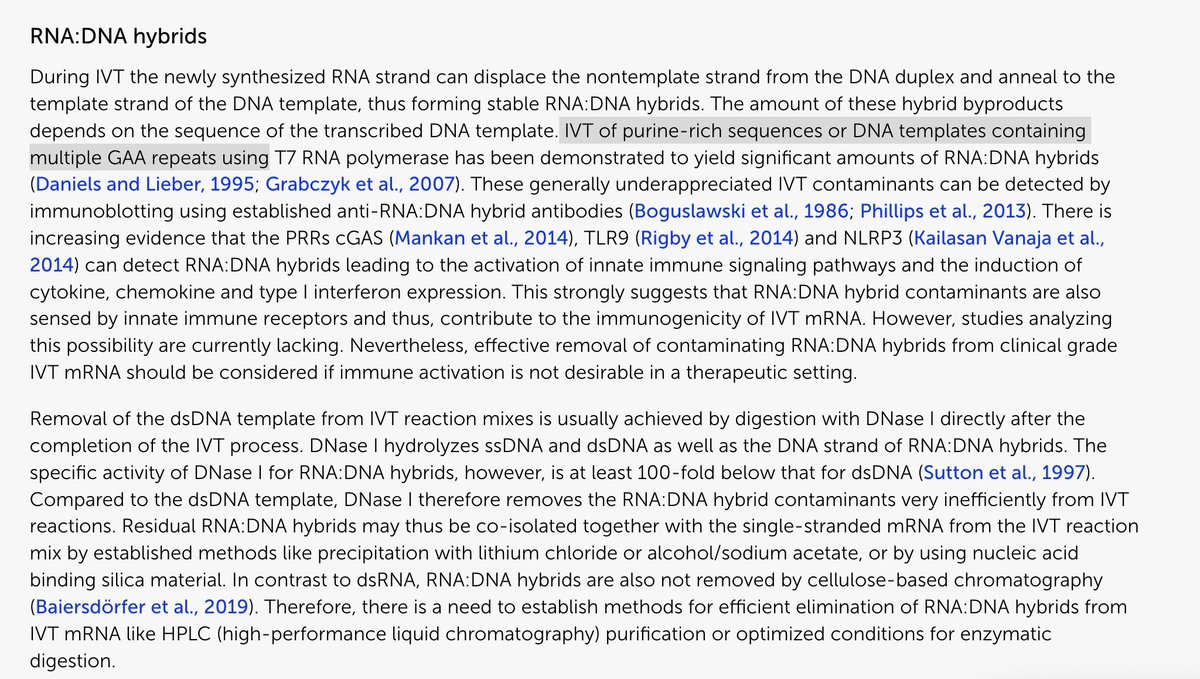
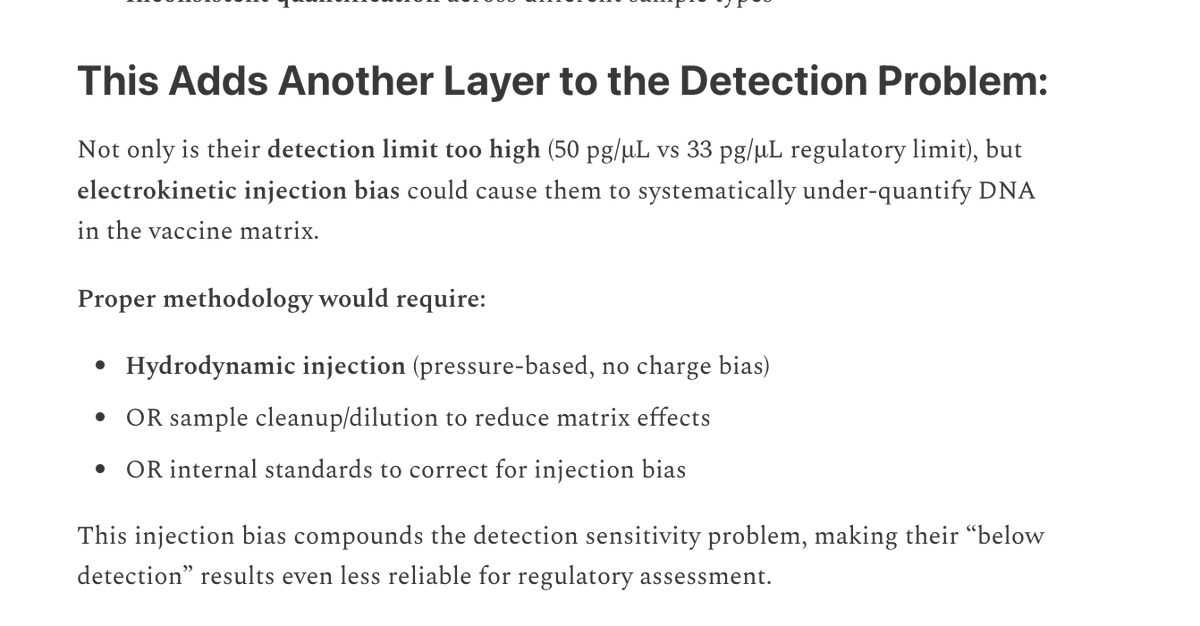
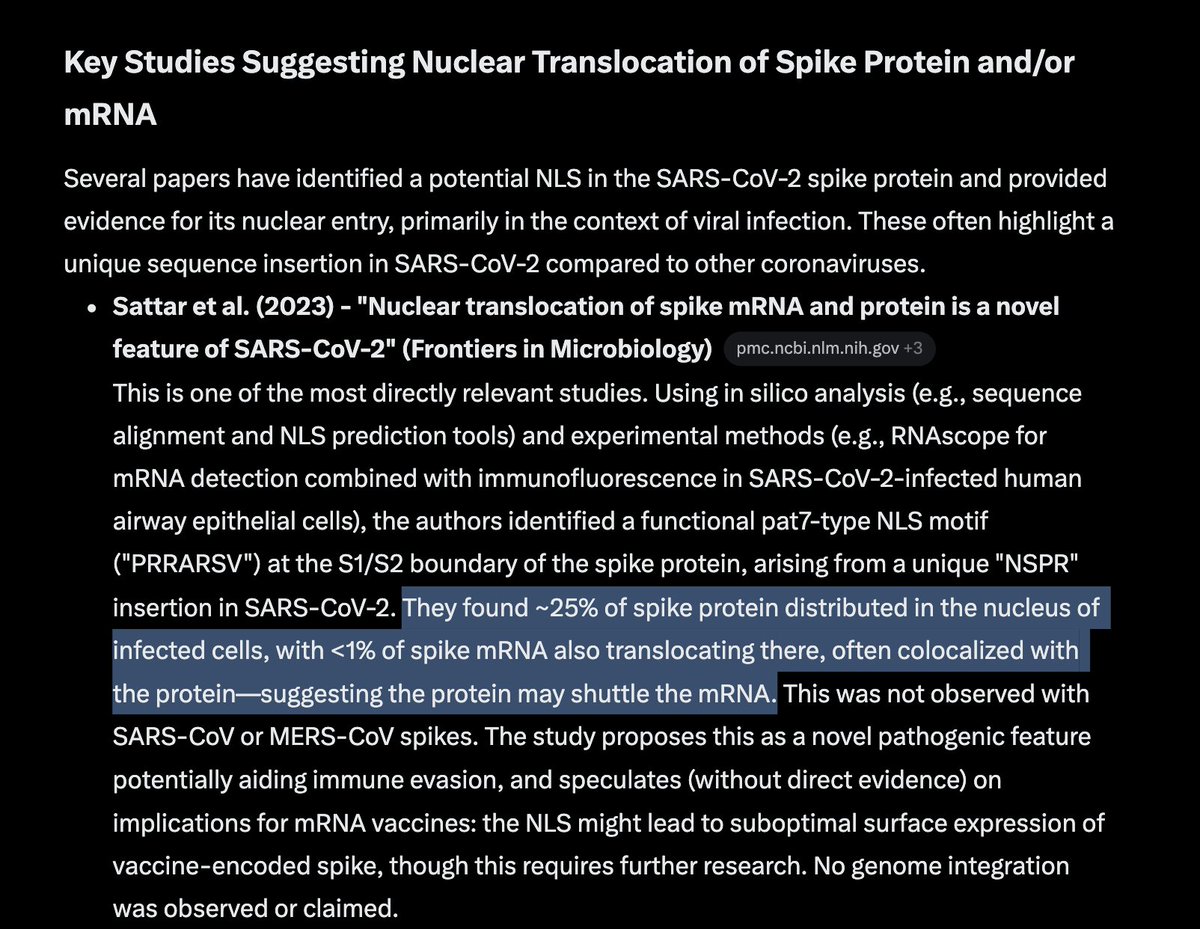
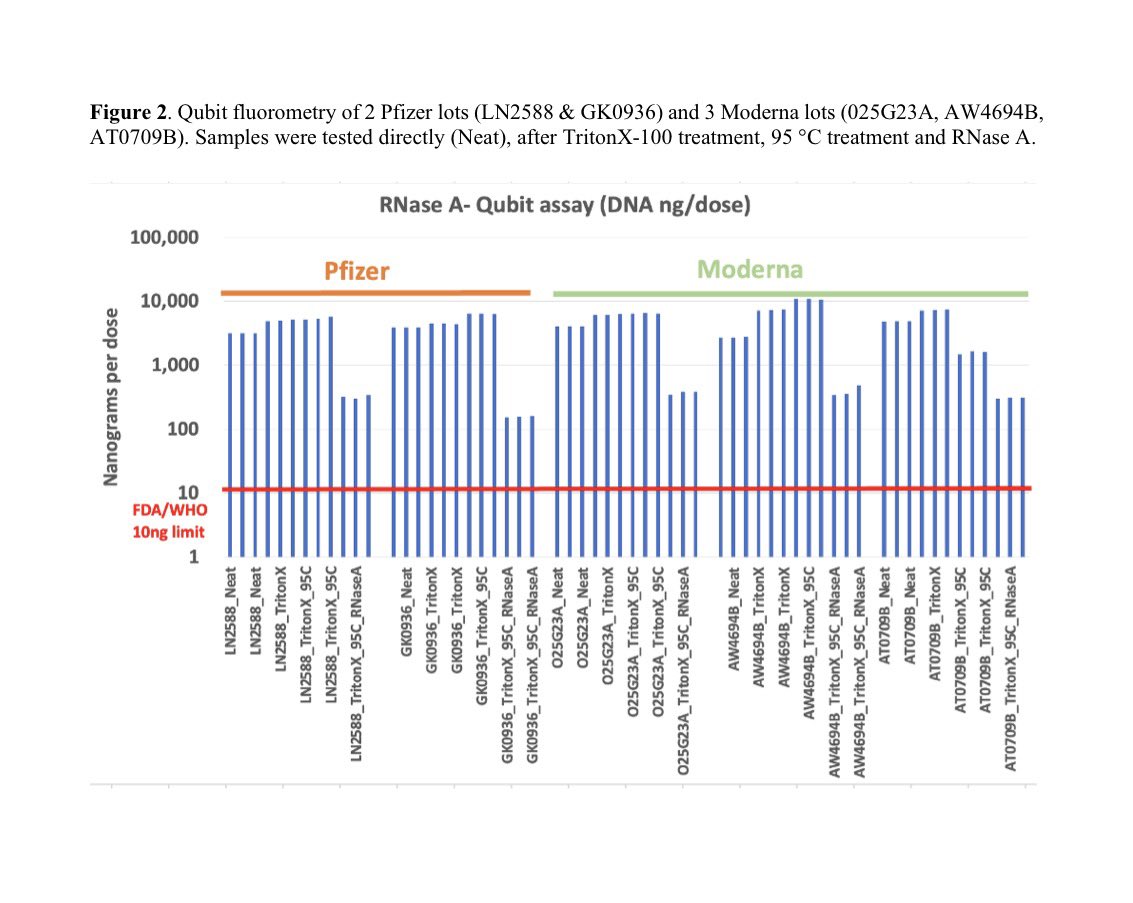
Many people ask me about this Moderna patent sequence.
Some calc the odds of a 19mer by chance as 4^19.
A big number if life were truly random.
But evolution is a preservation of those random words that improve fitness so we have to ask, are there similarities to common words?
Some calc the odds of a 19mer by chance as 4^19.
A big number if life were truly random.
But evolution is a preservation of those random words that improve fitness so we have to ask, are there similarities to common words?

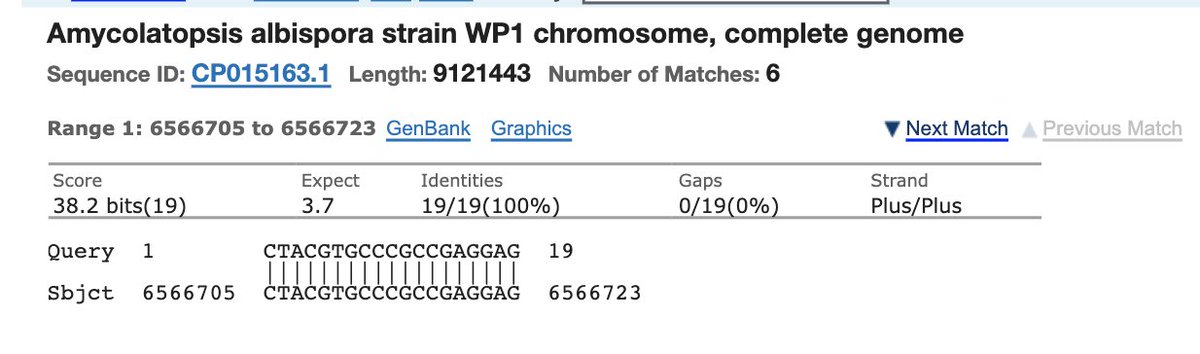
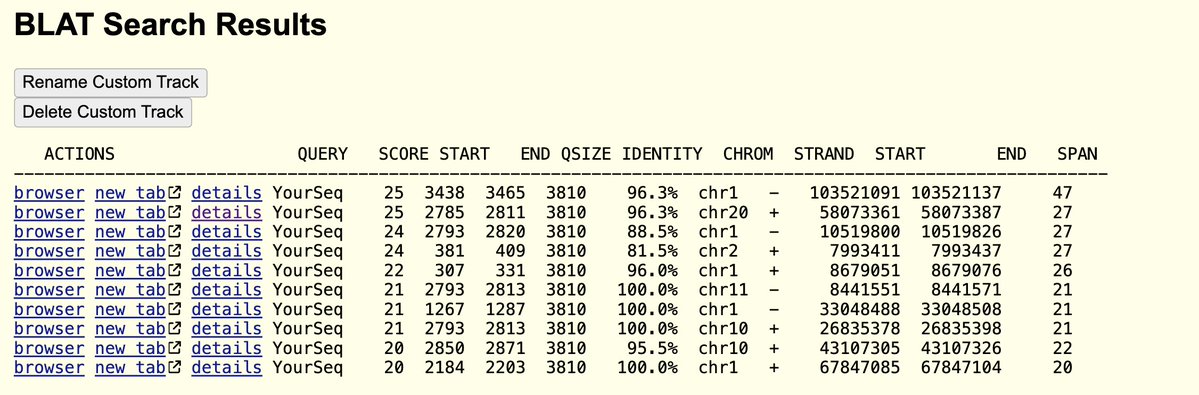
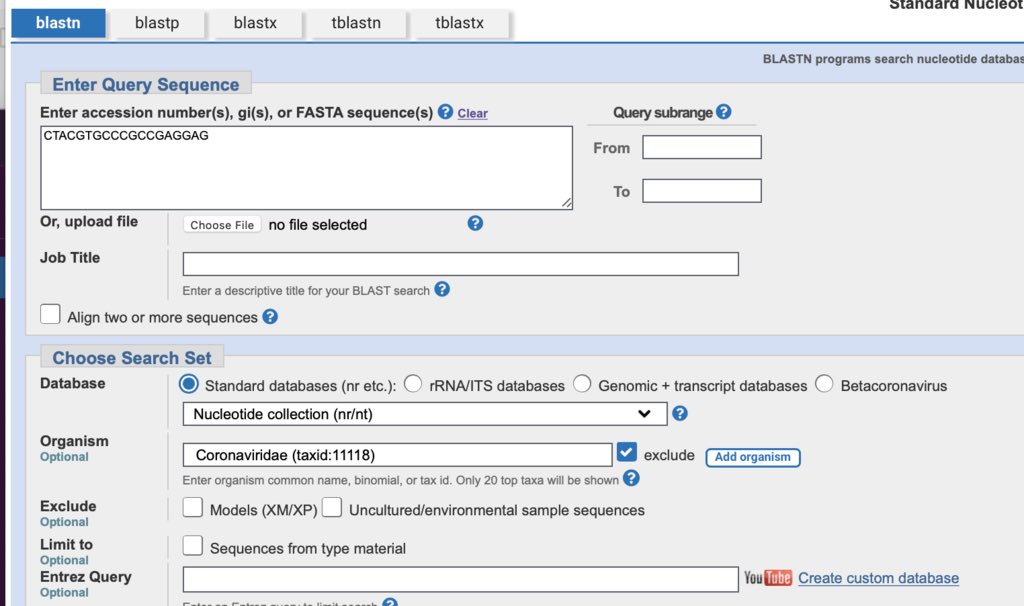
Take the 19mer sequence and plug it into NCBI BLASTn against the Nr database.
Check ‘exclude’ and enter coronaviridae.
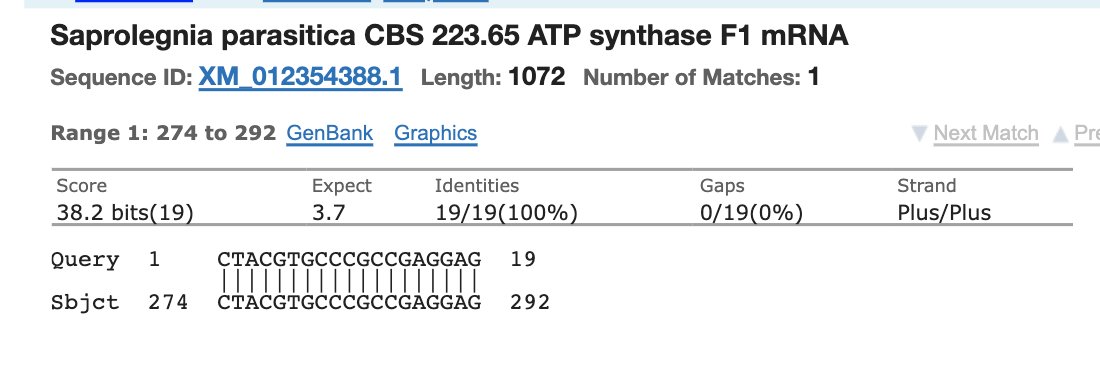
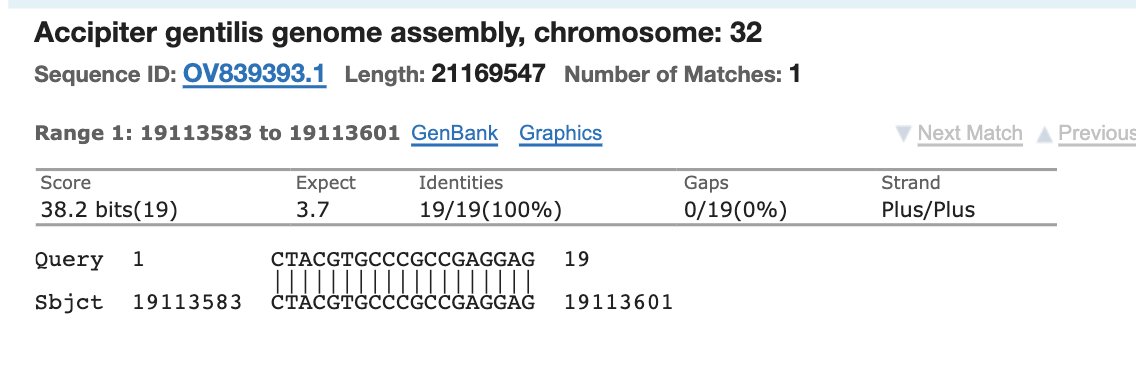
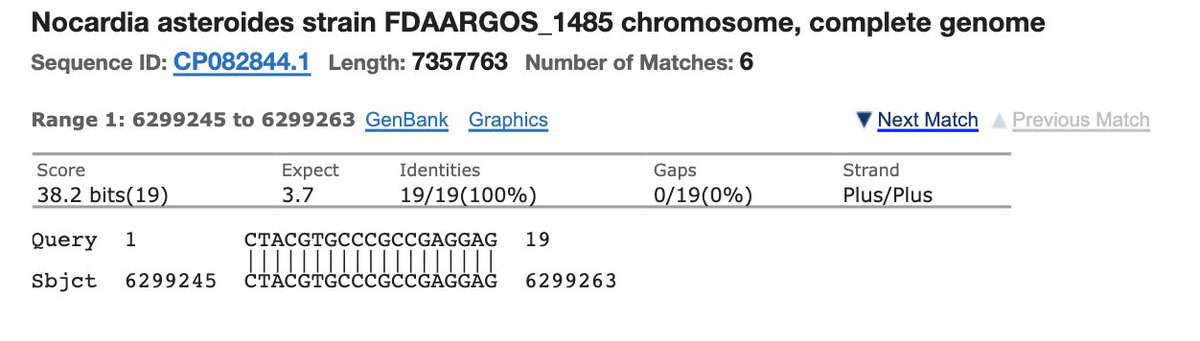
You’ll get microbial hits like this.
Check the E-Value.
What’s the E-Value?
Check ‘exclude’ and enter coronaviridae.
You’ll get microbial hits like this.
Check the E-Value.
What’s the E-Value?
Q: What is the Expect (E) value?
The Expect value (E) is a parameter that describes the number of hits one can “expect” to see by chance when searching a database of a particular size. It decreases exponentially as the Score (S) of the match increases.
The Expect value (E) is a parameter that describes the number of hits one can “expect” to see by chance when searching a database of a particular size. It decreases exponentially as the Score (S) of the match increases.
Essentially, the E value describes the random background noise. For example, an E value of 1 assigned to a hit can be interpreted as meaning that in a database of the current size one might expect to see 1 match with a similar score simply by chance.
The lower the E-value, or the closer it is to zero, the more “significant” the match is. However, keep in mind that virtually identical short alignments have relatively high E values. This is because the calculation of the E value takes into account the length of the query seq
These high E values make sense because shorter sequences have a higher probability of occurring in the database purely by chance. For more details please see the calculations in the BLAST Course.
/end of NCBI description
/end of NCBI description
If we Steel man the argument, Moderna attorneys would point to all the microbes that have the same 19bp sequence and ask if they are guilty of making that sequence 50 million years ago.
It’s smoke, not fire.
But who built the P4 lab in Wuhan and why is that CEO now at Moderna?
It’s smoke, not fire.
But who built the P4 lab in Wuhan and why is that CEO now at Moderna?
We’re usually excited about E-values <10-4. E Values above 1 implies you’ll get many sequences back by pure chance.
I don’t think those will hold up in court alone.
We need more evidence.
I don’t think those will hold up in court alone.
We need more evidence.
If you take Seq ID 11652 (~3.3kb) and blast (megablast) it against NR, you get 77% hits to PREDICTED mRNAs in Chinchilla. These are found with Whole genome shotgun and automatic annotation pipelines. 

Using discontinuous BLAST you can capture Human mRNAs with E values of Zero and 72% identical. These are much stronger hits than the Evalue we get blasting to SARs (100% for only 19bp, E= 31. They were chasing cancer vaccines back then. This would explain the MutS kitchen sink. 
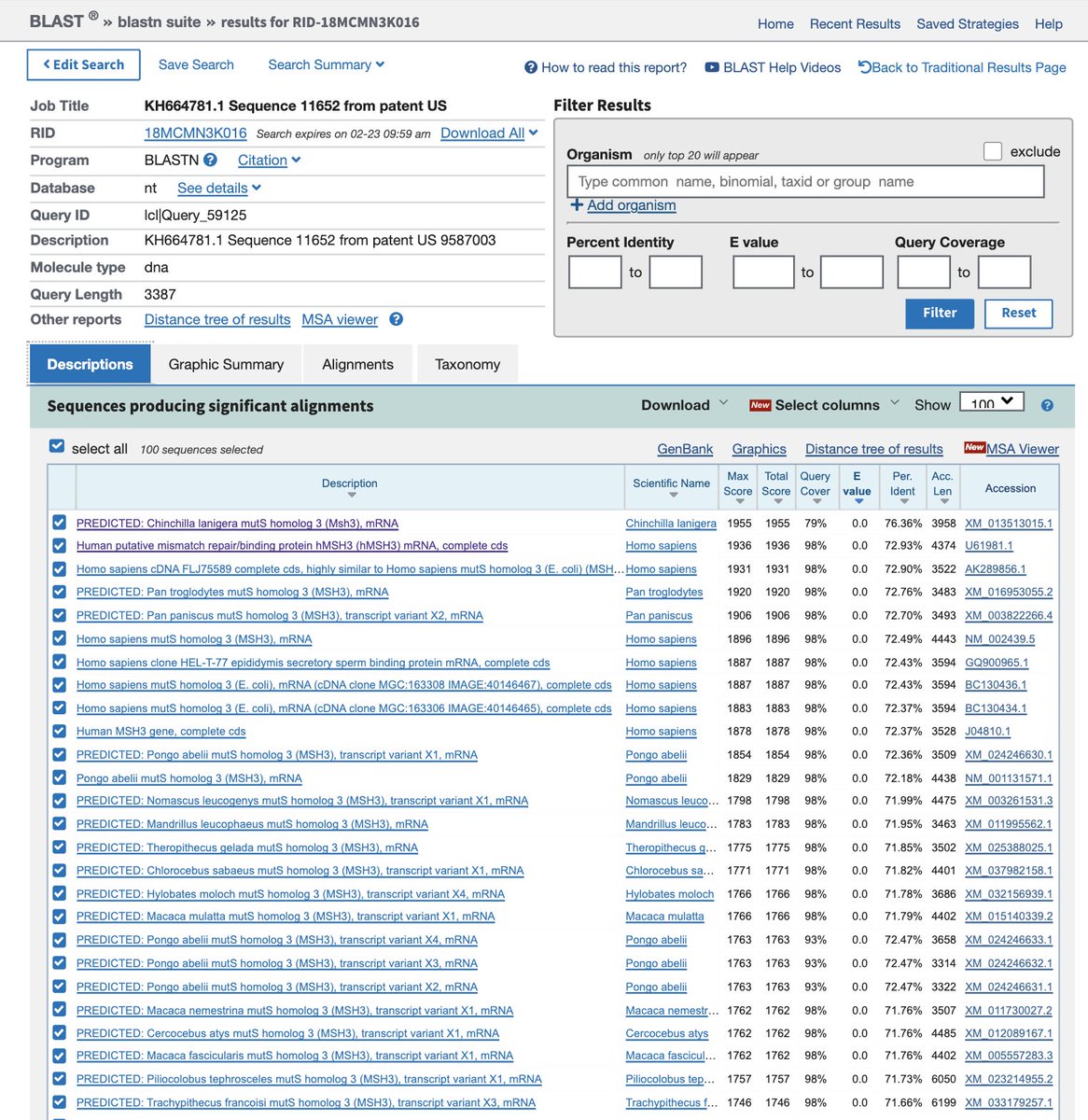
A list of other organisms this 19bp sequence exists in.
Nature has played this tune many times before.
Nature has played this tune many times before.

I am not suggesting bacteria has recombined this sequence into SARs-CoV-2. I am suggesting that Moderna didn't invent this sequence. Mother Nature did, and someone likely emulated it into SARs-CoV-2. A kitchen sink patent that has all human genes in the specification..
Isn't a smoking a gun for where this sequence was derived from. It demonstrates desperation of Biotech company, trying to perform an IP land grab on any human genome put into their platform for any reason. They can make additional claims from anything in the specification through
A Continuation in Part (CIP) or a divisional patent application. Hence, they threw everything into their specifications, including human MSH3 (similar to E.coli MutS).
A pertinent question is why did a paper publish that buried all of these obvious hits?
Why did they Re-Invent the E-value with a new way of calculating these odds when time tested BLAST offers odds they chose to not highlight?
Why did they Re-Invent the E-value with a new way of calculating these odds when time tested BLAST offers odds they chose to not highlight?
Some have accused me of carrying water for Moderna. I carry water for the truth and if the opposition can easily dismantle an argument with whats posted in this thread, it is better to hash it out now than later.
So what would be convincing evidence?
So what would be convincing evidence?
In any synthetic biology project DNA oligos that code for the construct are synthesized. These are usually outsourced to companies like IDT. Strong evidence would be Moderna ordering oligos that consist of SARs-CoV-2 sequence and this FCS. That would demonstrate intent.
This would avoid 'Bible Codes' like associations of kitchen sink patent publication to SARs-CoV-2.
Importantly, sequences listed in patent specifications that are not in the claims, are not owned nor invented by the patent applicant.
ams.org/notices/199708…
Importantly, sequences listed in patent specifications that are not in the claims, are not owned nor invented by the patent applicant.
ams.org/notices/199708…
Specification are like citations in a paper that can be used in the future to potentially expand claims if they can pass the 101, 102 ,and 103 tests at the USPTO for novelty, inventiveness and utility. Seq ID 11652 was never put to this test as its not in the claims.
More useful resources.
If you want to dig into this patent-
You can download the 33,915 sequences included in the specifications at this link.
It is a 92Mb file.
seqdata.uspto.gov/?pageRequest=d…
If you want to dig into this patent-
You can download the 33,915 sequences included in the specifications at this link.
It is a 92Mb file.
seqdata.uspto.gov/?pageRequest=d…
Only 2 Seq ID are mentioned in the claims. 7572 and 16557. The first sequence is an amino acid sequence. The other 33,913 sequences are not claimed. 

This is a kitchen sink that becomes a CIP (continuation in part ) pool. Once these seqs are on record in the specifications, they can draw from them and file additional claims later. This leads to a long network of intertwined IP.I see no evidence that Seq ID 11652 is claimed. 

If it is not claimed, it is not proprietary. The paper used this language. There is no test for inventiveness which should not surprise anyone as people were speaking openly about FCS sites in 2015.
This isn't 'their' sequence
This isn't 'their' sequence
https://twitter.com/jhas5/status/1496478686240980996?s=21
Correction.. not all of these sequences have human as the top hit. Many are just computer predicted genes in genome projects. One of the claimed sequences is from water buffalo and picked up by NCBIs automated annotation pipeline. 

If you are interested in further reading on gene patents and the tangled web they weave....
researchgate.net/publication/25…
researchgate.net/publication/25…
Here is the patent. Try searching it for Seq ID 11652. You wont find it as it is so buried, its only in the downloadable file above.
patents.google.com/patent/US95870…
patents.google.com/patent/US95870…
What you can find if you BLAST this against all patents are patents from Dow in 2001 that have 17bp of the 19.
Has the complete 12bp FCS!!!
Get your Pitch Forks and Bible Codes!

Has the complete 12bp FCS!!!
Get your Pitch Forks and Bible Codes!

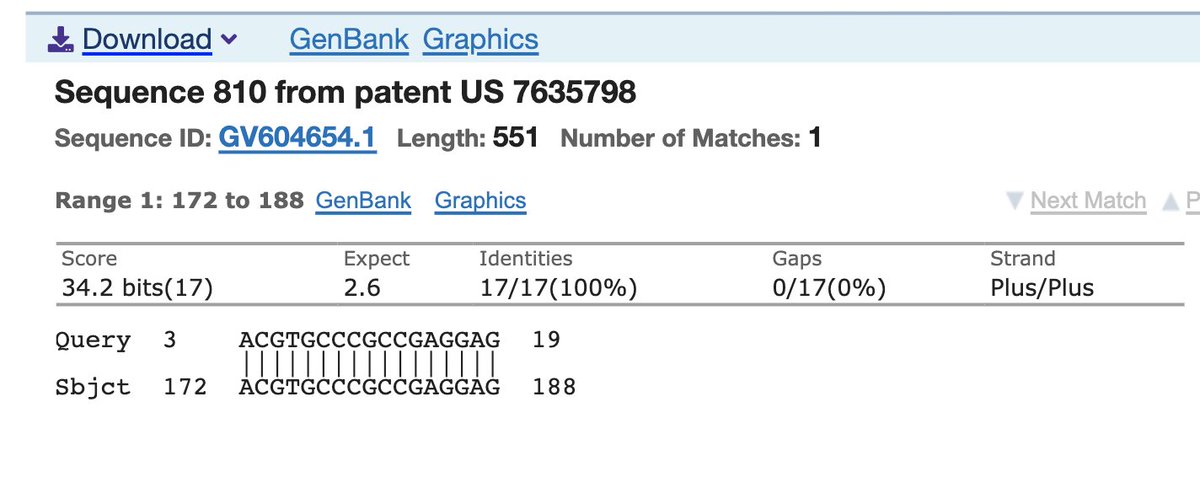
Or this Chlamydomonas
18/19bp.
You can see how sloppy of a signature this is. There is a reason Next Generation sequencers tend to have at least 25bp reads.
18/19bp.
You can see how sloppy of a signature this is. There is a reason Next Generation sequencers tend to have at least 25bp reads.
• • •
Missing some Tweet in this thread? You can try to
force a refresh