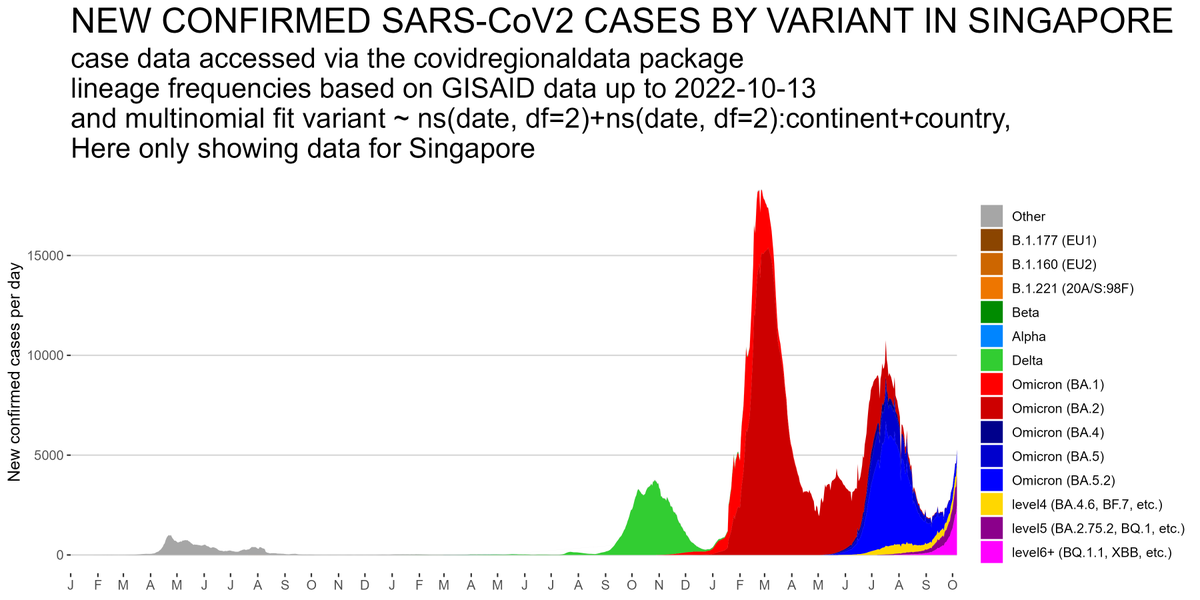
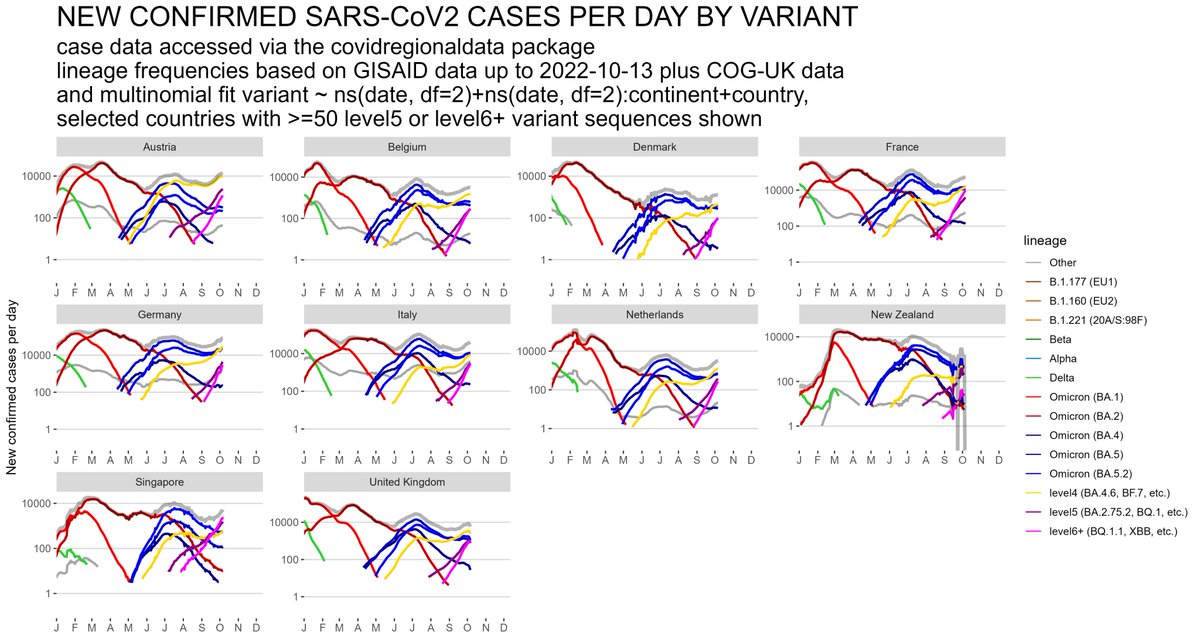
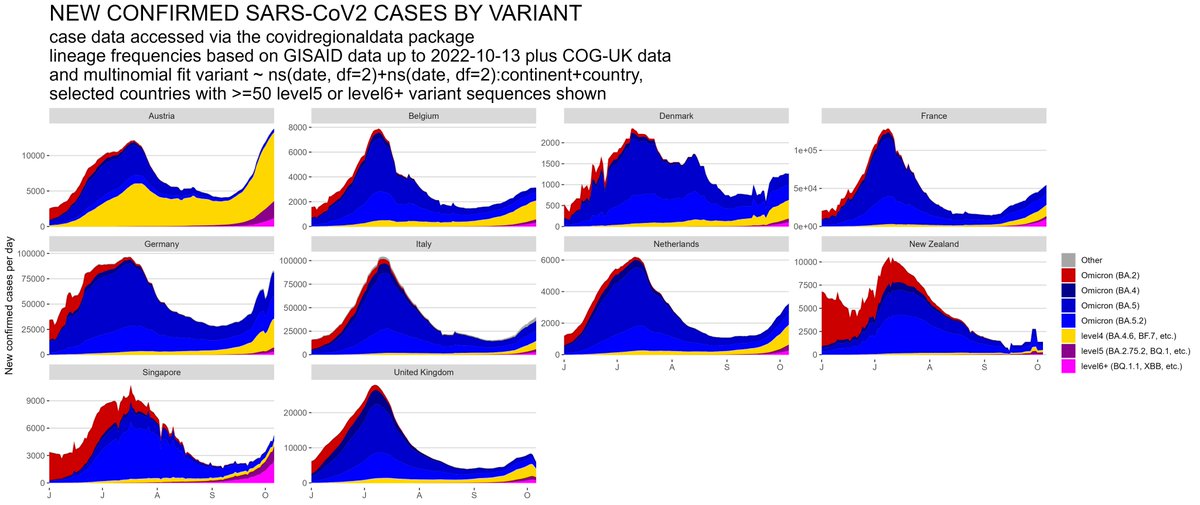
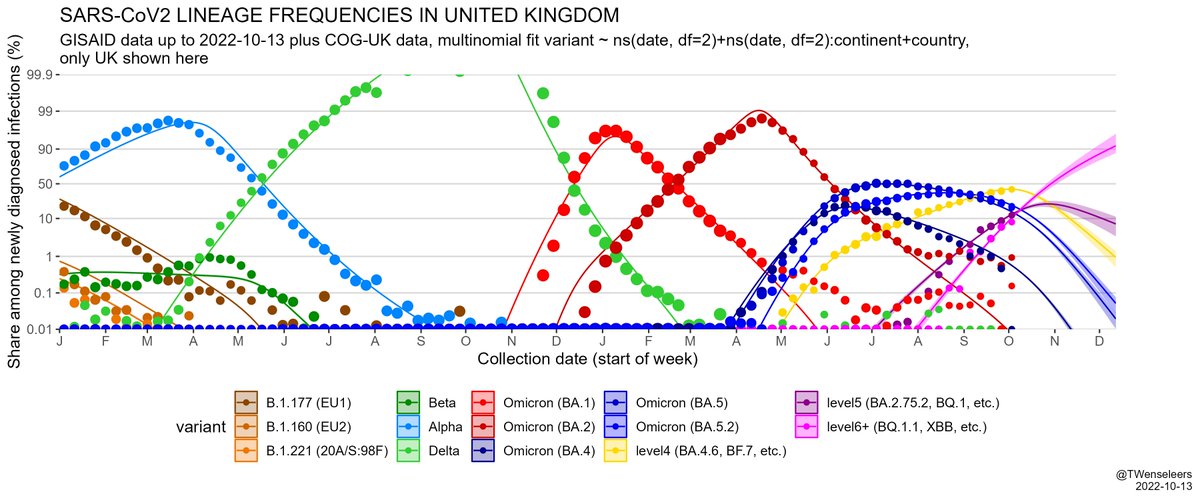
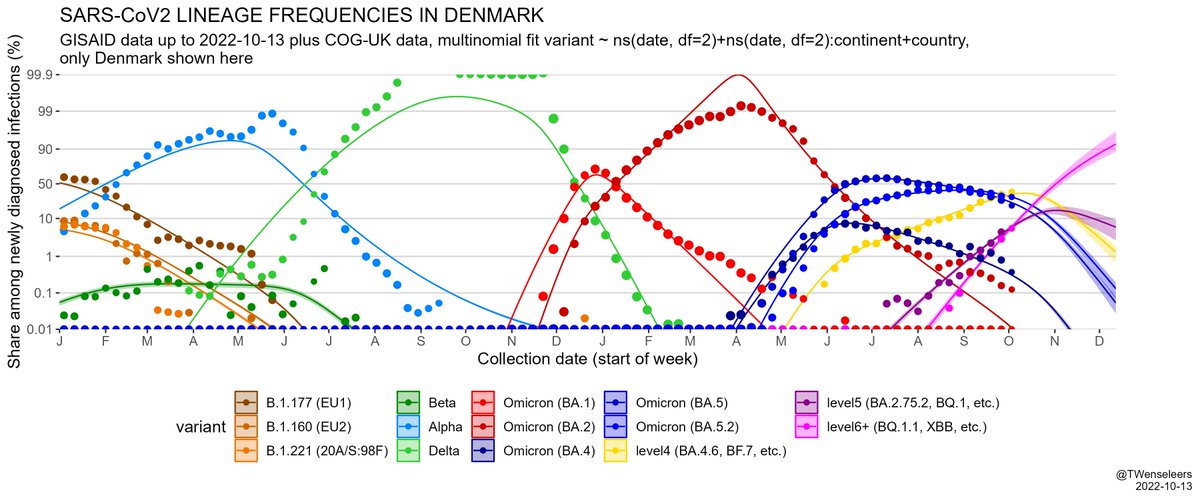
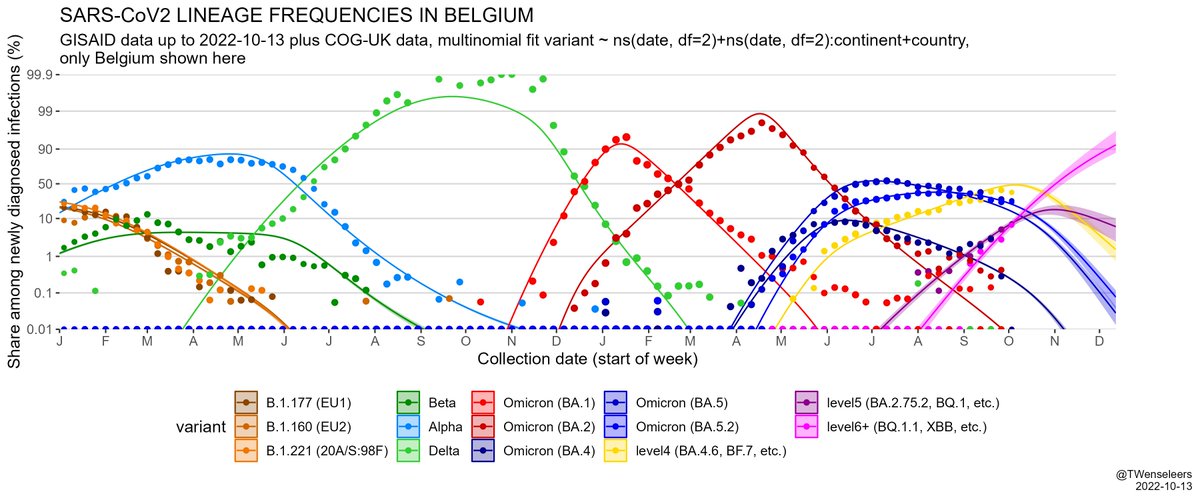
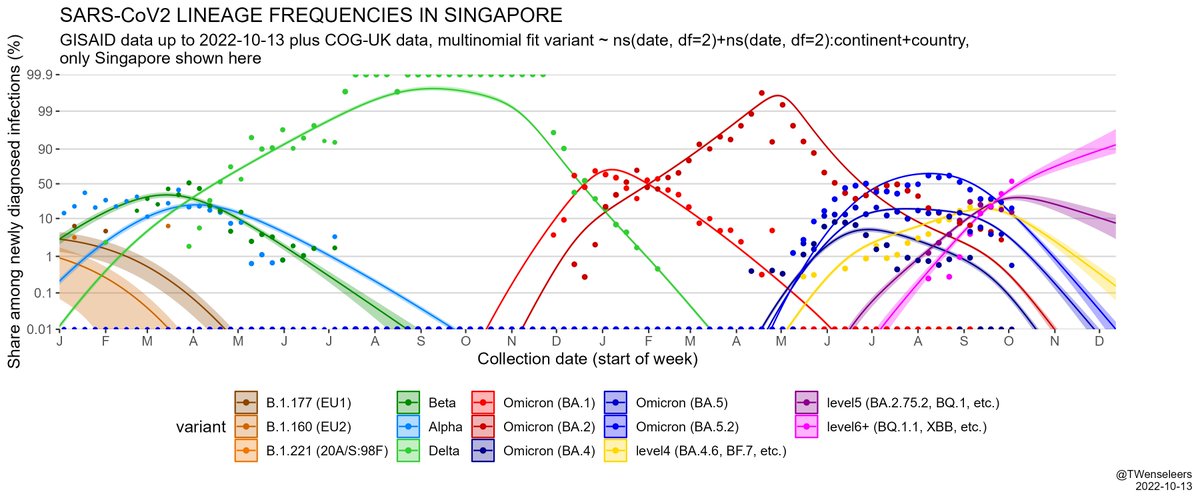
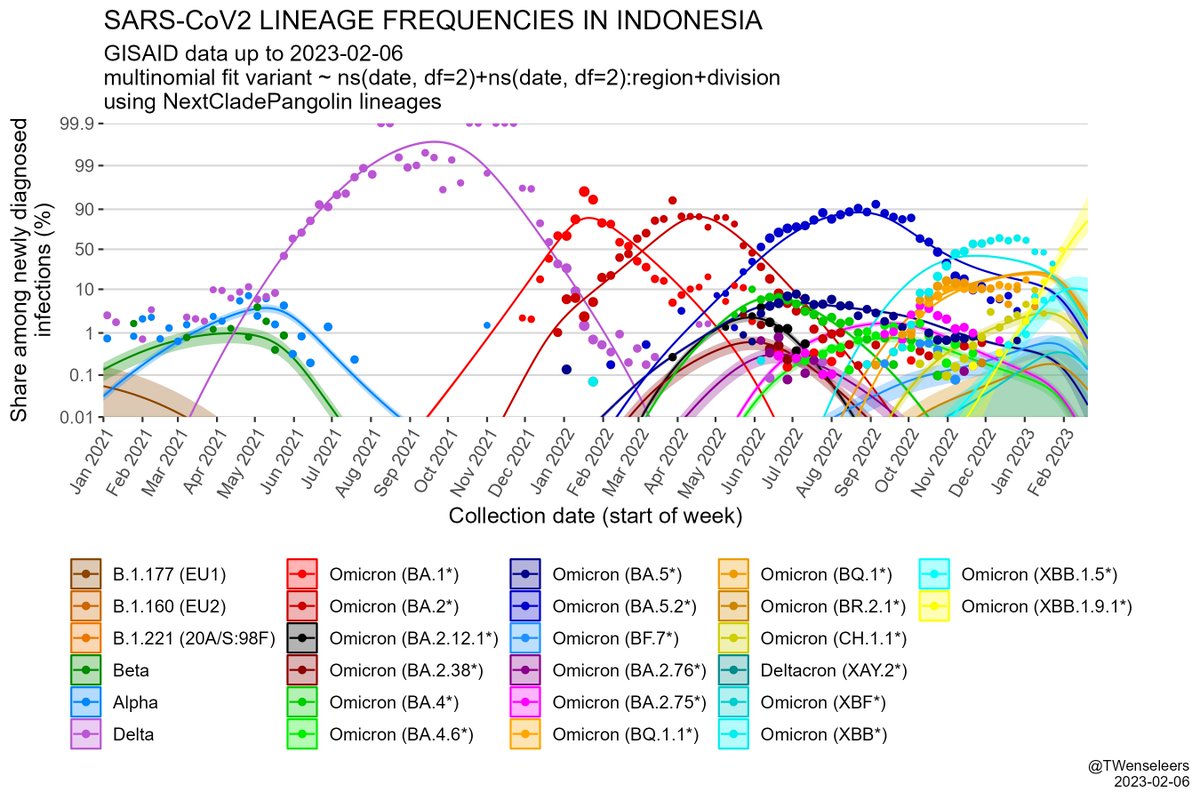
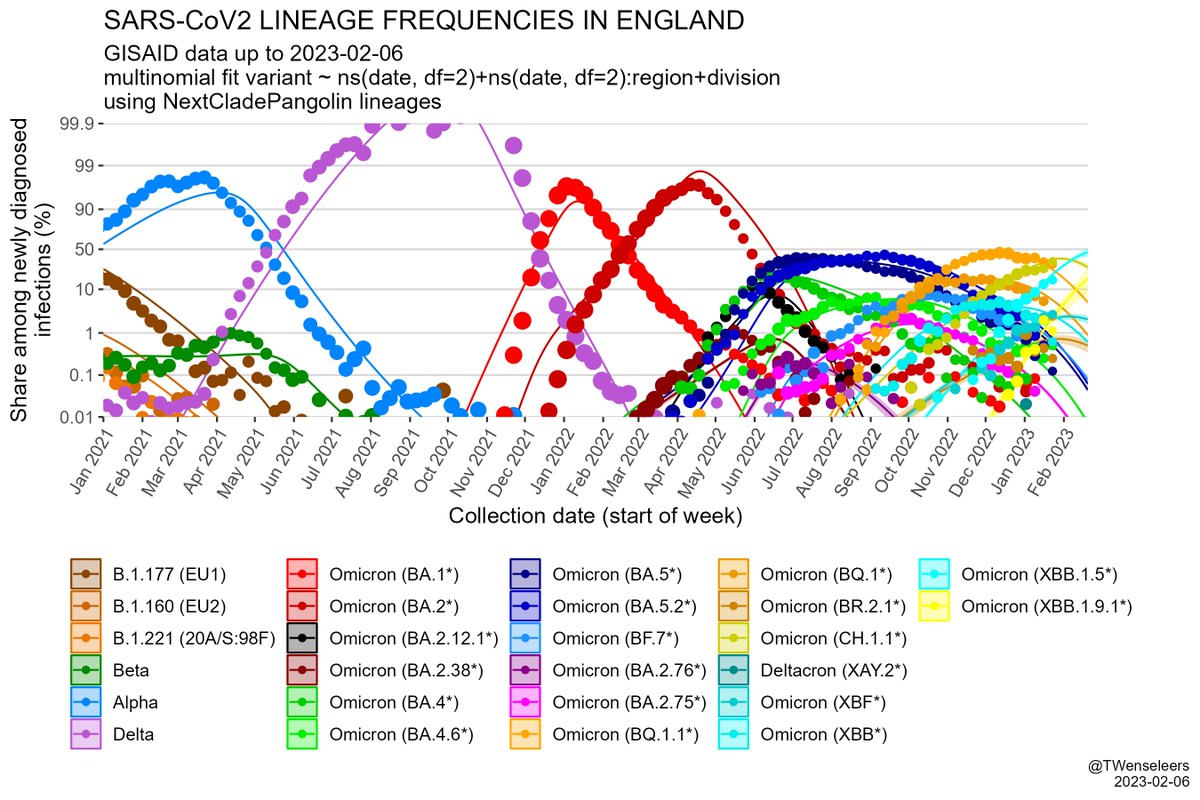
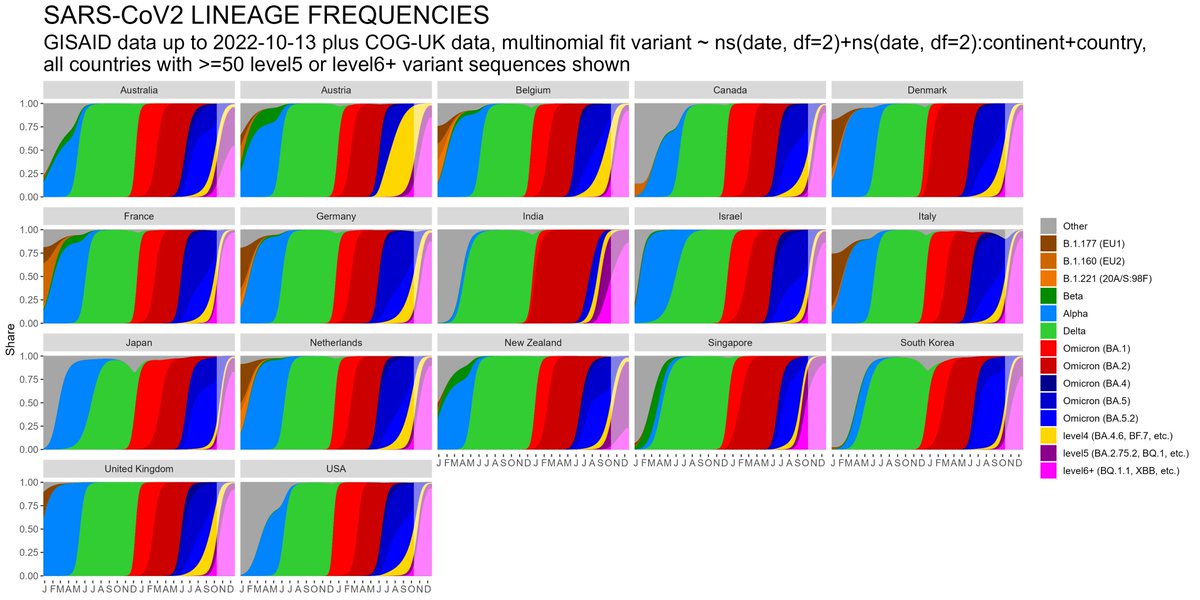
The SARS-CoV2 lineage soup was becoming a bit too complex to plot nicely, so here some simplified plots with lineages grouped by number of key mutations present, as suggested by @CorneliusRoemer (with some help from @rquiroga777). 

With the exception of Singapore, that is experiencing an XBB driven wave now, level6+ variants (BQ.1.1, XBB, etc) would appear to still be at a relatively low level & not to exert a lot of pressure on case nrs yet, but this might soon change,... 
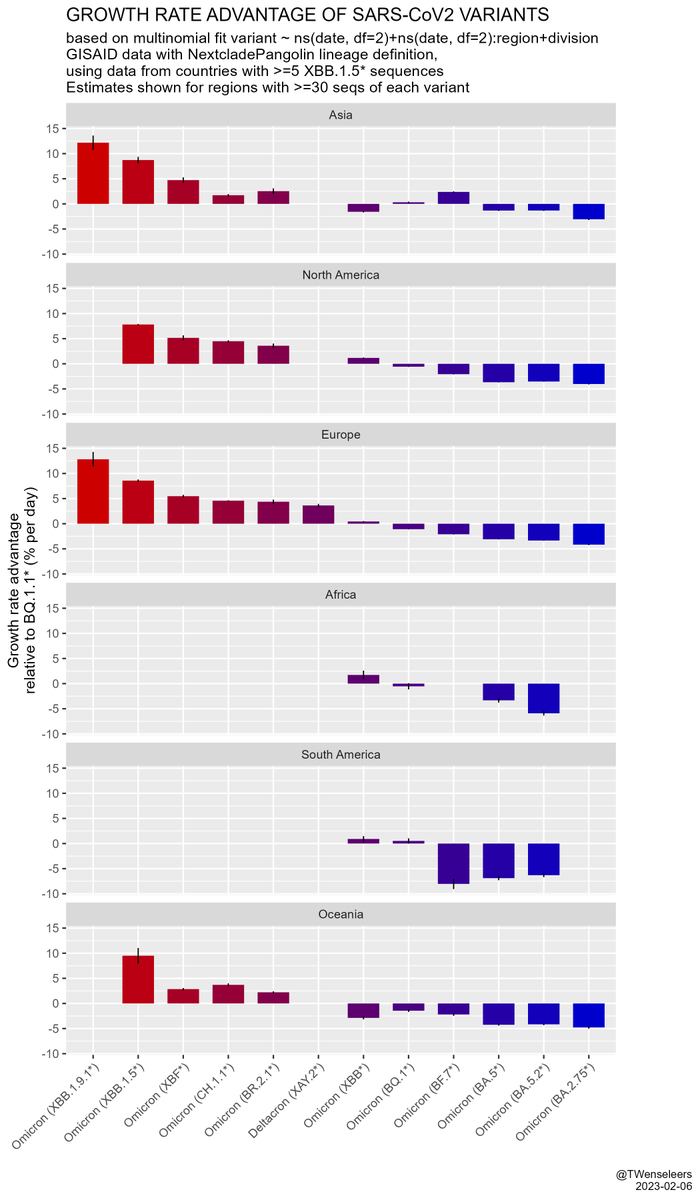
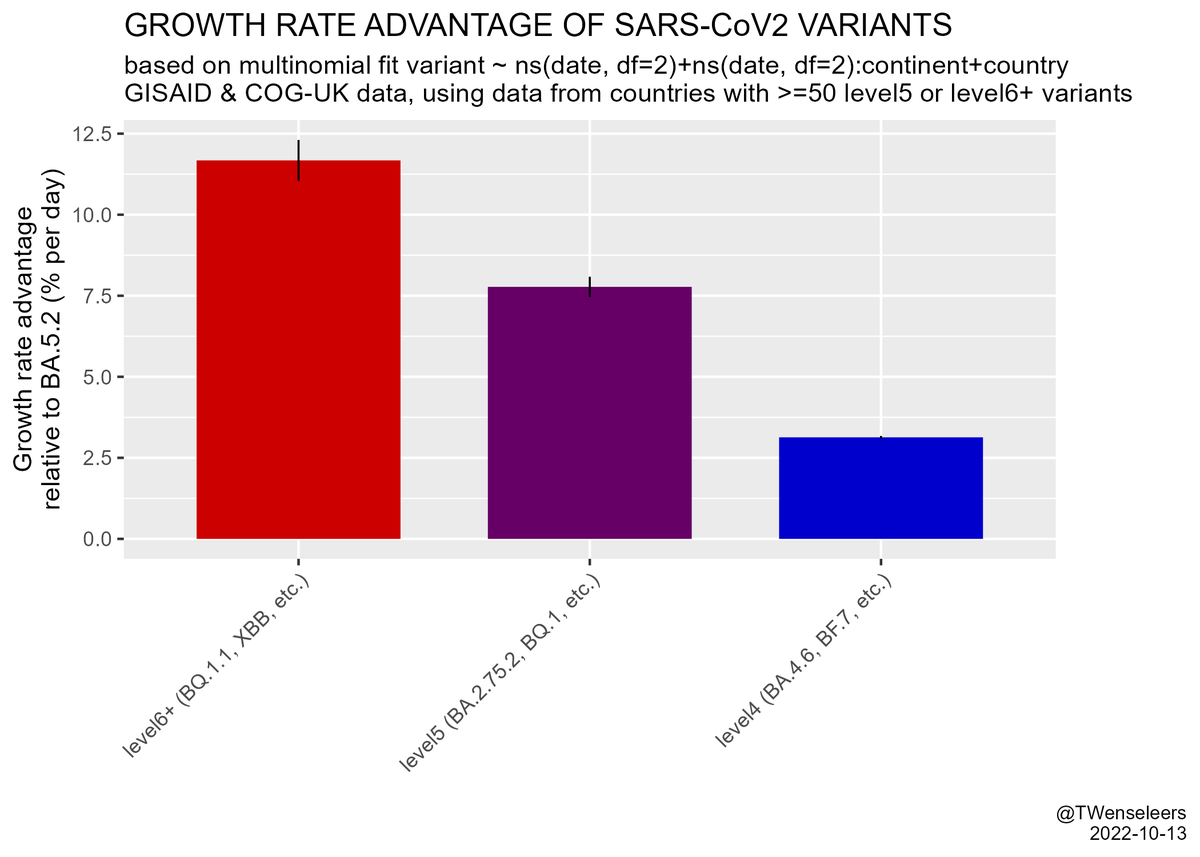
and that the estimated growth rate advantage of level6+ variants (BQ.1.1, XBB, etc) over resident type BA.5.2 is in excess of 10% per day. All this means a resurgence is likely later in November, even if cases currently start to flatten a bit... 
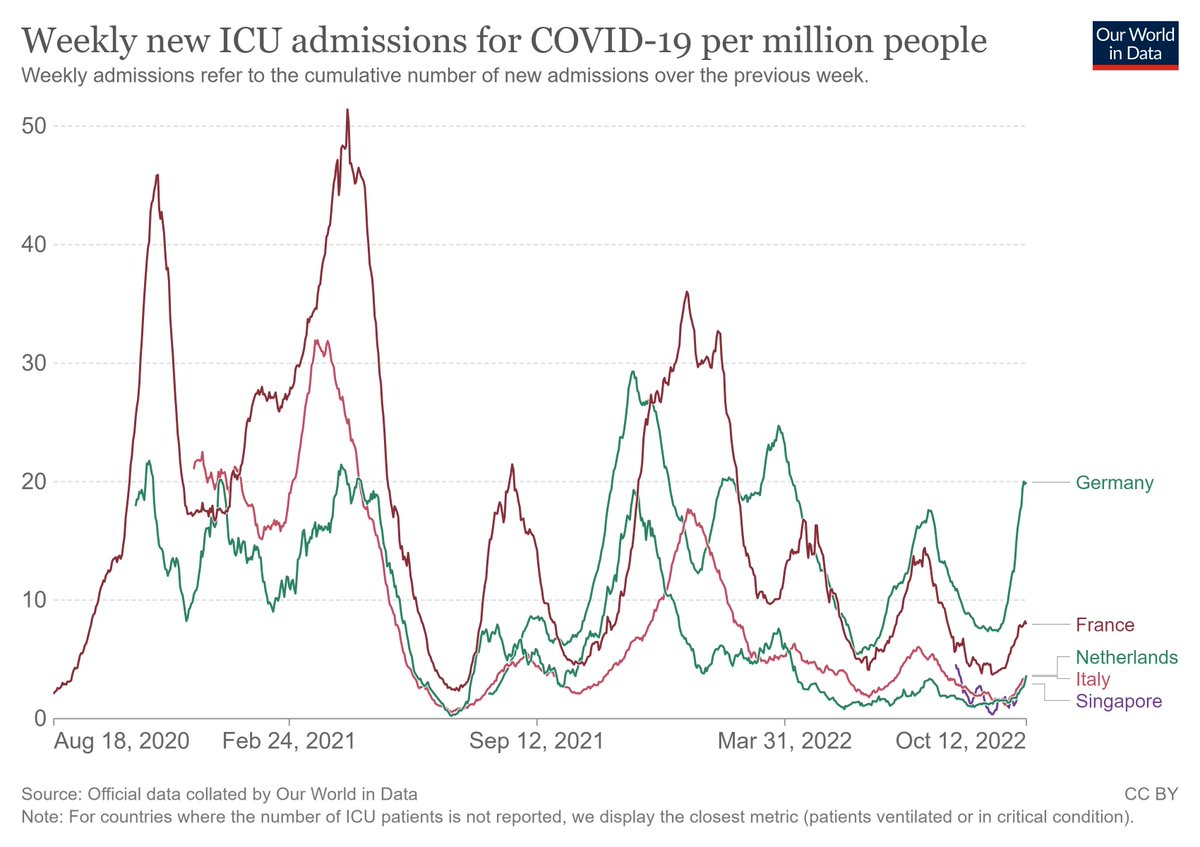
Western European countries are seeing rising hospital and ICU admissions, with especially Germany already seeing figures that are higher than expected. If you get your invitation for a booster it might be a good idea to go and get it... 
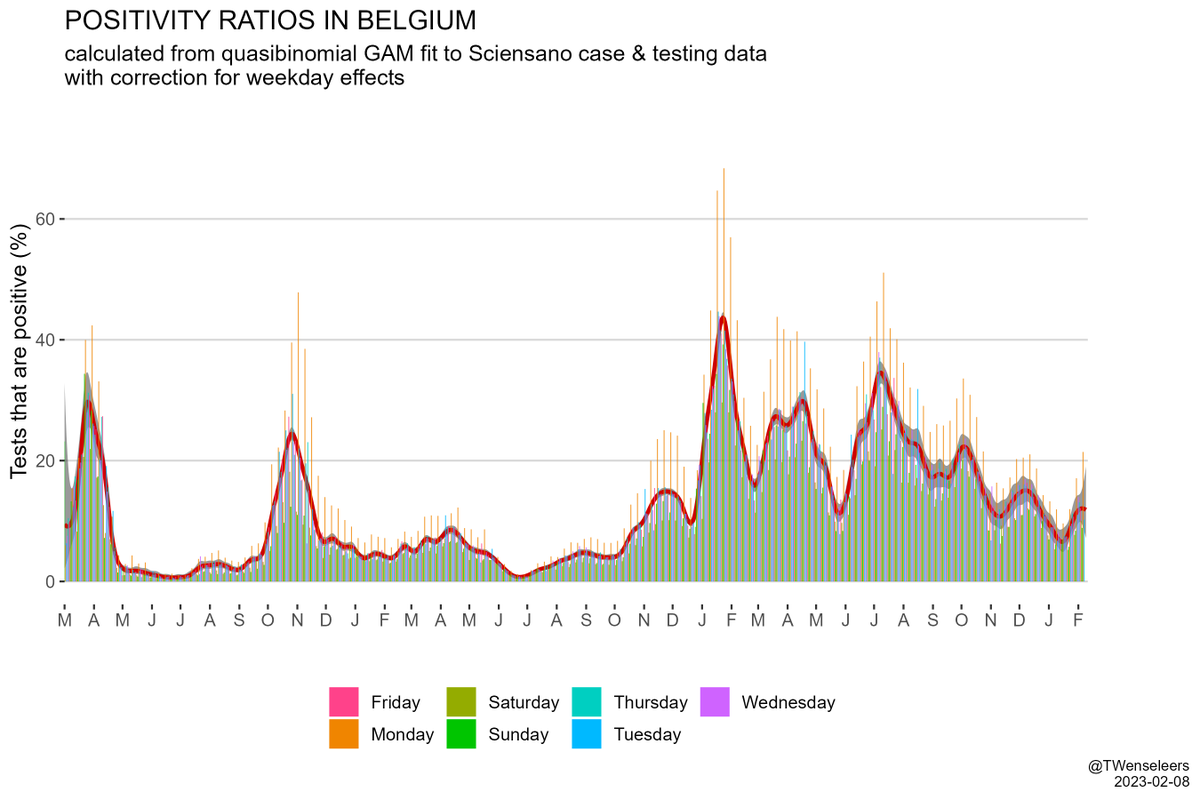
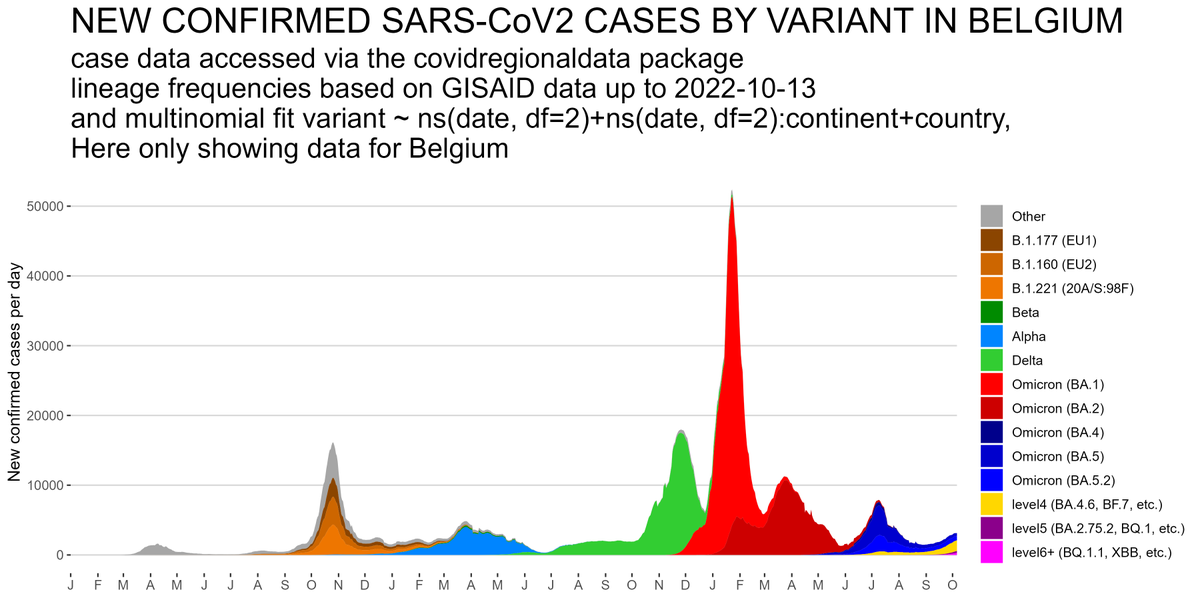
In W Europe, level 6+ lineages are still too rare to have much of an effect on case nrs - the resurgence we saw so far was mostly driven by waning immunity & a change in contact patterns. Only in Belgium BF.7 might have played some role as well. But effect of BQ.1.1 yet to come. 
R code of plots & analysis of global @GISAID & @CovidGenomicsUK data can be found here: github.com/tomwenseleers/…. Some more plots can be found here github.com/tomwenseleers/….
• • •
Missing some Tweet in this thread? You can try to
force a refresh