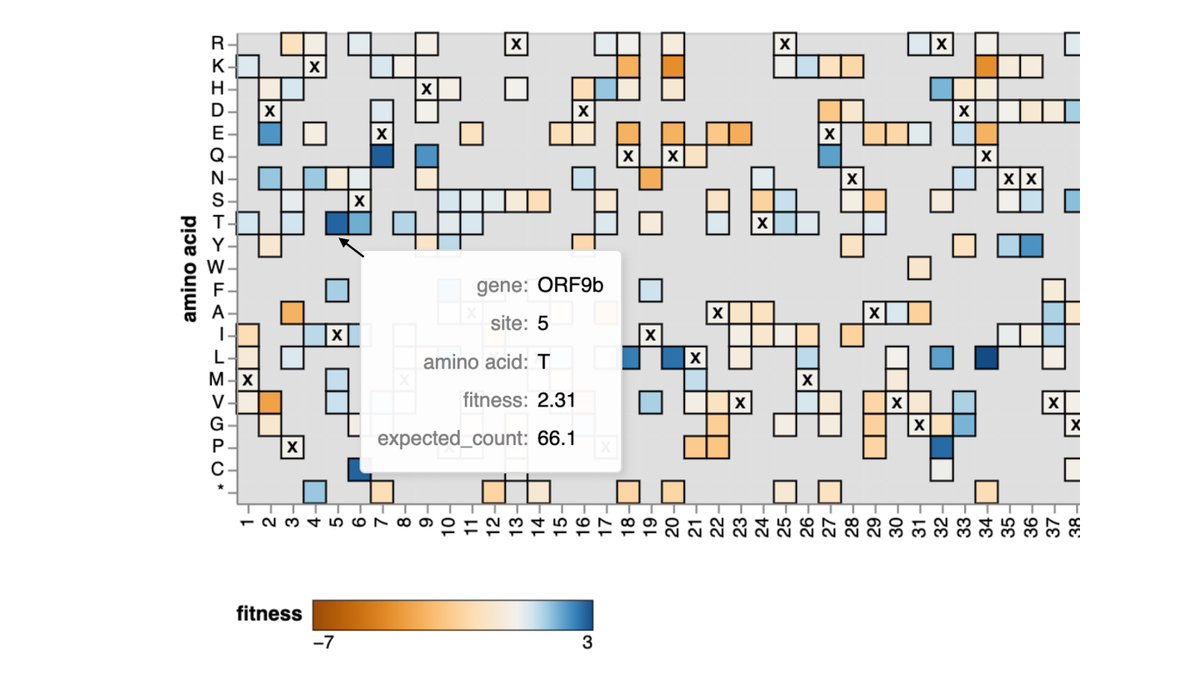
I wanted to provide assessment of spike amino-acid mutations in new highly mutated 2nd generation BA.2 SARS-CoV-2 variant
This is analysis for those (like me) scientifically interested in SARS2 evolution
(Anyone else is completely within reason to ignore this variant right now)
This is analysis for those (like me) scientifically interested in SARS2 evolution
(Anyone else is completely within reason to ignore this variant right now)
https://twitter.com/LongDesertTrain/status/1690989522035617792
Full analysis of the mutations is in these slides:
Analysis is based mostly on deep mutational scanning experiments
TLDR: lots of antigenic change, and some interesting RBD mutations (addition of N-linked glycan & deletion in receptor-binding motif)slides.com/jbloom/new_2nd…
Analysis is based mostly on deep mutational scanning experiments
TLDR: lots of antigenic change, and some interesting RBD mutations (addition of N-linked glycan & deletion in receptor-binding motif)slides.com/jbloom/new_2nd…
First, to emphasize, only THREE sequences of variant identified so far. There is not currently evidence of wide transmission.
As this thread outlines, people who study SARS2 evolution may want to pay attention to features of this variant. Everyone else can ignore if they wish.
As this thread outlines, people who study SARS2 evolution may want to pay attention to features of this variant. Everyone else can ignore if they wish.

As has been noted already, this variant has lots of amino-acid mutations in spike: 33 relative to its putative ancestor BA.2. It is also very different from XBB.1.5.
This makes it an evolutionary jump comparable in size to that which originally gave rise to Omicron.
This makes it an evolutionary jump comparable in size to that which originally gave rise to Omicron.

Below is list of spike amino-acid mutations relative to BA.2, w my annotations of likely effects based on experimental data from @bdadonaite @tylernstarr @yunlong_cao @Dr_MattMcCallum @veeslerlab
One thing that is obvious is that many of these mutations cause antibody escape.
One thing that is obvious is that many of these mutations cause antibody escape.

I’m going to call out three mutations that I think are especially interesting.
K356T creates a N-linked glycosylation site (at N354) in the RBD, with the resulting glycan likely to completely mask that antibody epitope.
K356T creates a N-linked glycosylation site (at N354) in the RBD, with the resulting glycan likely to completely mask that antibody epitope.

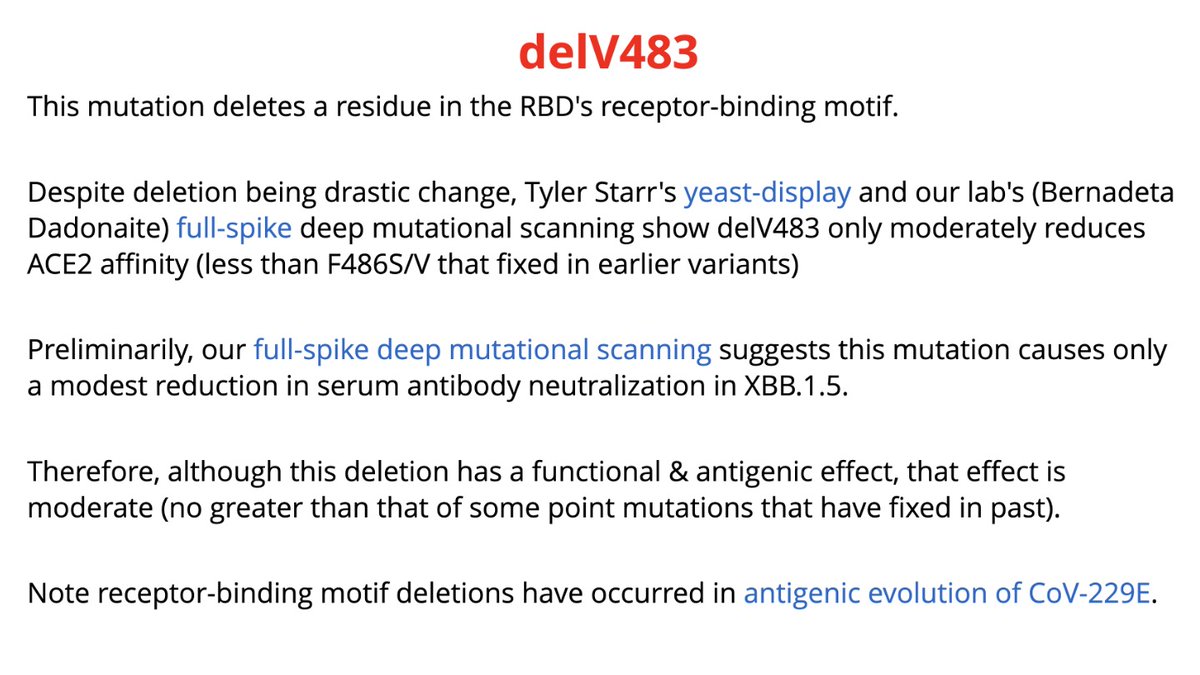
There is deletion of V483 in RBD’s receptor-binding motif
Unpublished data from @tylernstarr & @bdadonaite (linked in slides) indicate delV483 will moderately reduce ACE2 affinity & antibody recognition, but effect will be no bigger than some point mutations we’ve seen.
Unpublished data from @tylernstarr & @bdadonaite (linked in slides) indicate delV483 will moderately reduce ACE2 affinity & antibody recognition, but effect will be no bigger than some point mutations we’ve seen.

There is P1143L at beginning of S2 stem helix
In our spike deep mutational scanning (), we found mutating P1143 improves infection by pseudotypes.
We speculated just cell-culture phenomenon, but here is mutation in virus that transmitted at least a bit.cell.com/cell/fulltext/…
In our spike deep mutational scanning (), we found mutating P1143 improves infection by pseudotypes.
We speculated just cell-culture phenomenon, but here is mutation in virus that transmitted at least a bit.cell.com/cell/fulltext/…
To assess overall antigenic of mutations, we can use antibody-escape calculator () informed by data from @yunlong_cao
New variant has at least as much antigenic change relative to BA.2 as does XBB.1.5. https://t.co/z9ma98lY6Njbloomlab.github.io/SARS2-RBD-esca…

New variant has at least as much antigenic change relative to BA.2 as does XBB.1.5. https://t.co/z9ma98lY6Njbloomlab.github.io/SARS2-RBD-esca…
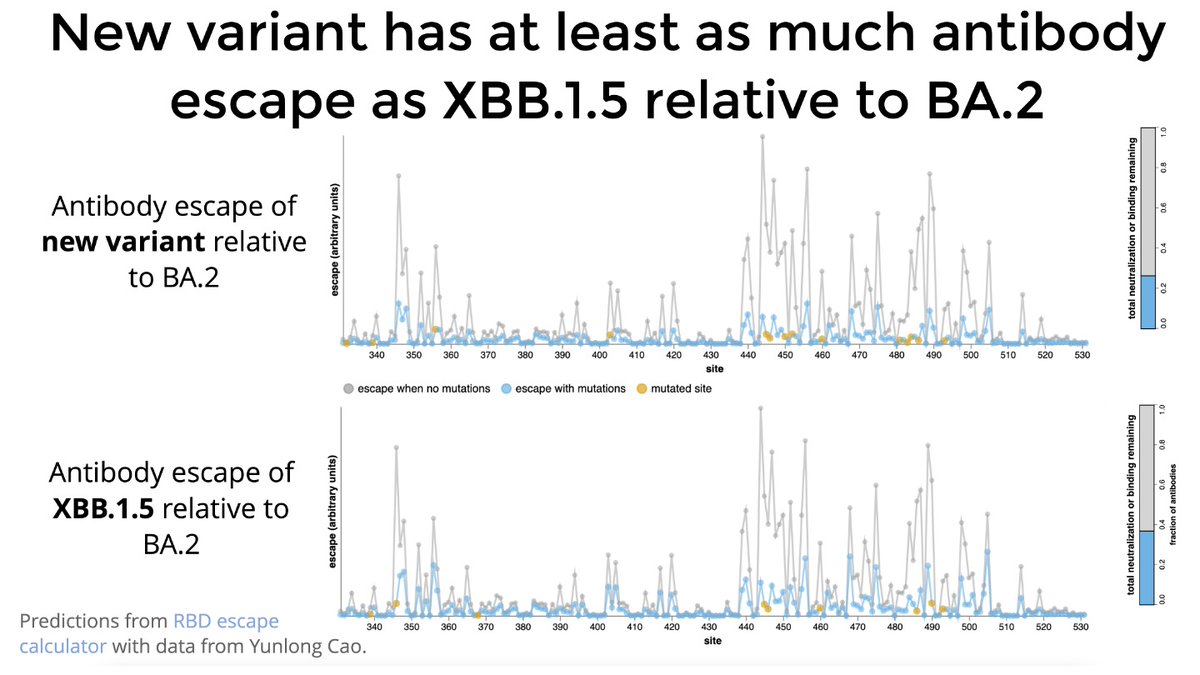
In fact, antigenic advantage of new variant over XBB is likely larger than indicated above, as many people have recently had XBB-specific antibodies boosted by infection. New variant differs from XBB at many key antigenic sites (see details)slides.com/jbloom/new_2nd…
Of course, to spread widely, antigenic advantage would need to be combined w inherent transmissibility close to best XBB variants
With only three sequences so far, there isn’t currently evidence that is case
But important to monitor if more sequences of new variant appear
With only three sequences so far, there isn’t currently evidence that is case
But important to monitor if more sequences of new variant appear
Thanks to:
Labs that sequenced variant:
Variant trackers who spotted sequences: @shay_fleishon, @LongDesertTrain, et al
Scientists doing deep mutational scanning on SARS2 spike mutations: @tylernstarr, @bdadonaite, @yunlong_cao, et alepicov.org/epi3/epi_set/2…
Labs that sequenced variant:
Variant trackers who spotted sequences: @shay_fleishon, @LongDesertTrain, et al
Scientists doing deep mutational scanning on SARS2 spike mutations: @tylernstarr, @bdadonaite, @yunlong_cao, et alepicov.org/epi3/epi_set/2…
• • •
Missing some Tweet in this thread? You can try to
force a refresh

 Read on Twitter
Read on Twitter