BREAKING🔔 A new study from G2P-Japan🇯🇵 is out! The preprint will be out so soon, but first we show our results on BA.2.86 - transmissibility, infectivity and immune resistance. This would be the 4th non-peer-reviewed🗒, but includes some new stuff. Please retweet/repost🔥 1/
Cf. Non-peer-reviewed works on BA.2.86:
The 1st🗒 from @yunlong_cao :
The 2nd🗒 from @BenjMurrell :
The 3rd🗒 from @BarouchLab :
Here I summarized the mutations detected in BA.2.86. 2/

The 1st🗒 from @yunlong_cao :
The 2nd🗒 from @BenjMurrell :
The 3rd🗒 from @BarouchLab :
Here I summarized the mutations detected in BA.2.86. 2/
https://twitter.com/yunlong_cao/status/1697318194976010446?s=20
https://twitter.com/BenjMurrell/status/1697751445351575656?s=20
https://twitter.com/BarouchLab/status/1698850719959544271?s=20

1⃣Transmissibility/fitness (Re):
Analysis by Jumpei @jampei2 using the data from 🇩🇰 showed that BA.2.86's Re is greater than XBB.1.5 and comparable to or even greater than EG.5.1. 3/
Analysis by Jumpei @jampei2 using the data from 🇩🇰 showed that BA.2.86's Re is greater than XBB.1.5 and comparable to or even greater than EG.5.1. 3/

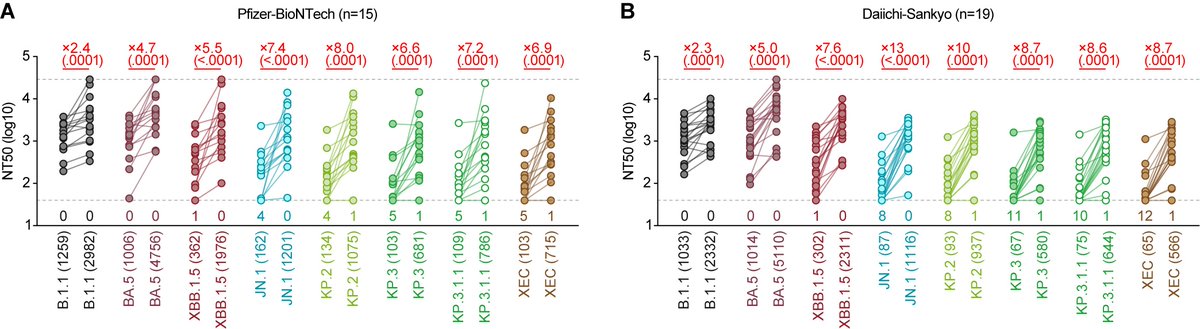
3⃣Immune resistance:
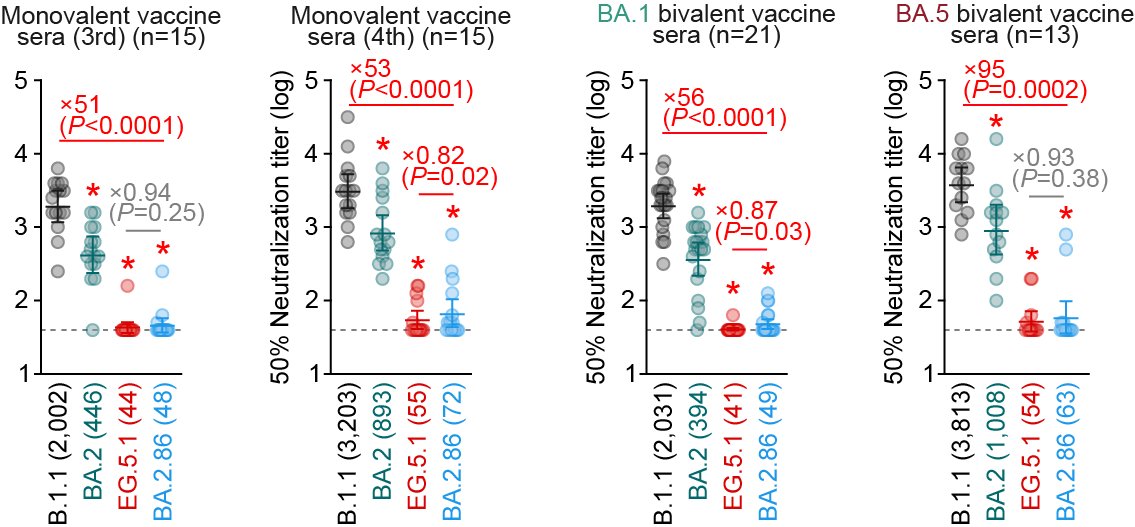
All vaccine sera tested (monox3, monox4, BA.1 bivalent, and BA.5 bivalent) showed NO neutralizing activity against BA.2.86 (and EG.5.1). 5/
All vaccine sera tested (monox3, monox4, BA.1 bivalent, and BA.5 bivalent) showed NO neutralizing activity against BA.2.86 (and EG.5.1). 5/
Three monoclonal antibodies (Bebtelovimab, Sotrovimab, Tixagevimab) that were antiviral against the parental BA.2 (see the G2P-Japan 9th paper) did NOT work against BA.2.86 AT ALL. 6/

https://twitter.com/SystemsVirology/status/1535191118354481152?s=20

Finally, the anti-BA.2.86 activity of XBB breakthrough infection sera was PRETTY POOR, and BA.2.86 was significantly (1.6-fold) more resistant to these sera than EG.5.1. 7/ 

Altogether, it is suggested that BA.2.86 is one of the most highly immune evasive variants ever and should have the potential to be considered as a variant of interest. 8/
Btw, 2 inturnship students from 🇮🇩 and 🇬🇧, Olivia and Yoonjin, did a great job in this urgent project. A great summer labwork🌊!! 9/
• • •
Missing some Tweet in this thread? You can try to
force a refresh