BREAKING NEWS 2025 💣💯
SARS-CoV-2 variants are not simply competing with each other, but are in fact WORKING TOGETHER in a COOPERATIVE MANNER !!! 🤔
pmc.ncbi.nlm.nih.gov/articles/PMC10…
SARS-CoV-2 variants are not simply competing with each other, but are in fact WORKING TOGETHER in a COOPERATIVE MANNER !!! 🤔
pmc.ncbi.nlm.nih.gov/articles/PMC10…

2) This study looked at how different versions of the COVID-19 virus, called variants, have evolved over time. The researchers found that these variants are not just competing with each other, but are actually working together. 
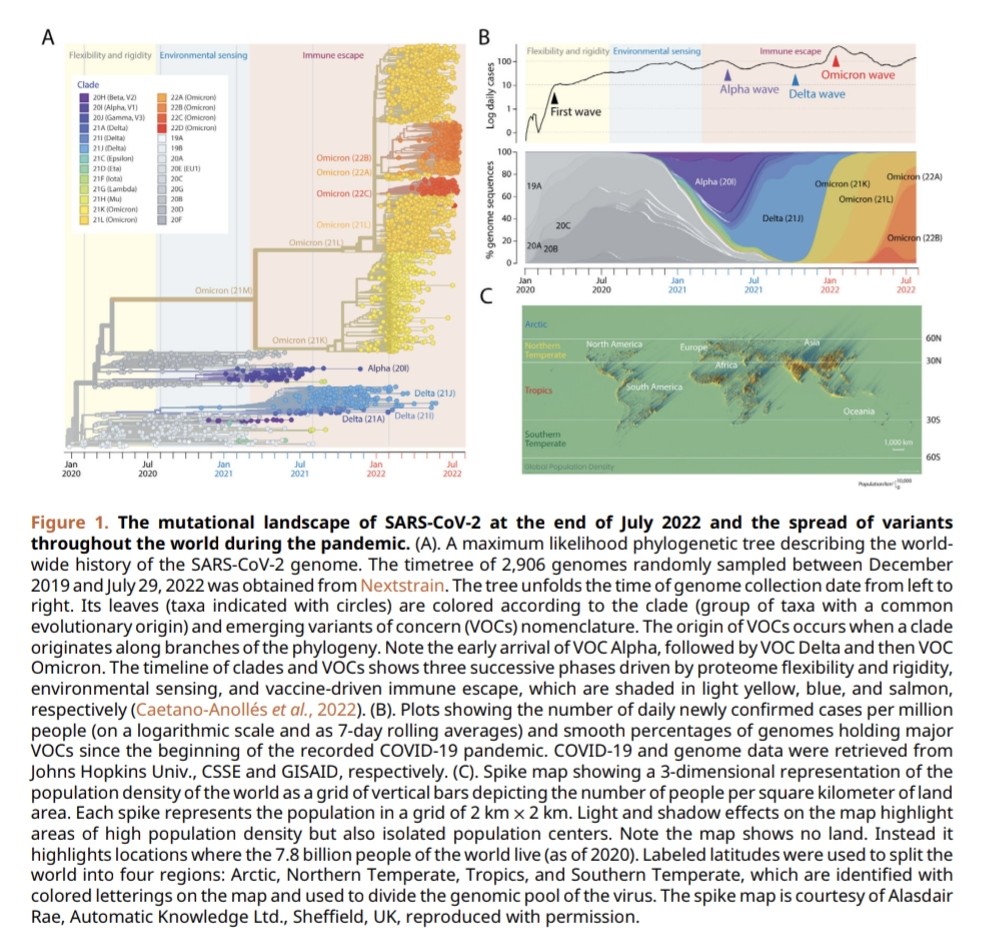
3) As new variants emerge, they take in and use the genetic changes (mutations) from previous variants, rather than completely replacing them. This suggests the variants are building on each other's changes, not just trying to beat each other. 

4) The study also showed that the combination of mutations in each variant helps balance out the extreme effects that individual mutations can have. This indicates the variants are working together to maintain a healthy balance, not just one variant dominating. 
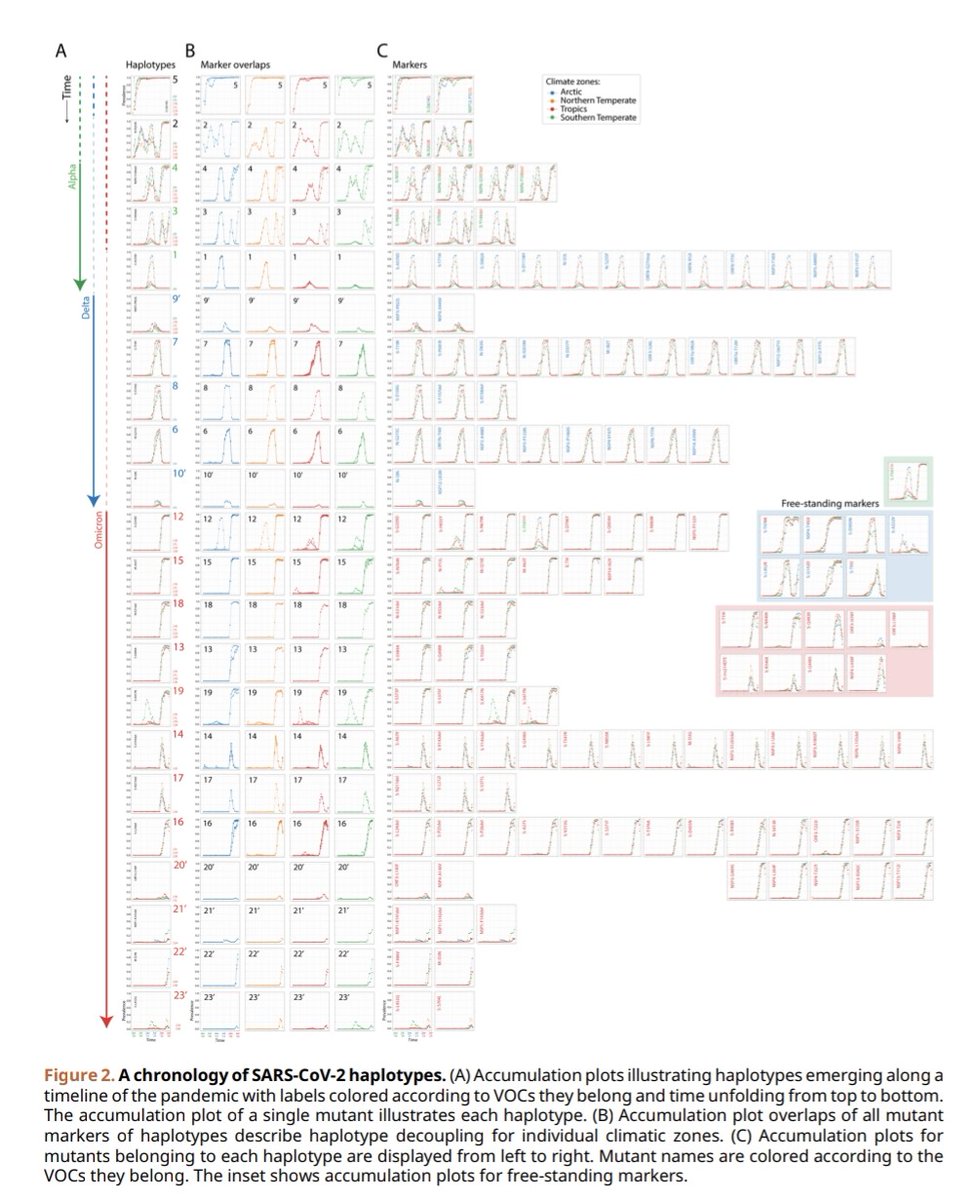
5) Another key finding was that the variants adapt to different seasons and locations in a coordinated way. The genetic signatures (haplotypes) of the variants became disconnected across regions due to seasonal changes. 

6) This implies the variants are adjusting to environmental conditions together, not competing for a single environment.
Overall, the researchers conclude the variants exhibit "cooperative evolution." The accumulation of mutations creates a complex but balanced landscape ...
Overall, the researchers conclude the variants exhibit "cooperative evolution." The accumulation of mutations creates a complex but balanced landscape ...

7) ...suggesting the variants are evolving together as a connected population, not just outcompeting each other.
In summary, the COVID-19 virus variants are not simply competing. They are working together by building on each other's changes, balancing extreme effects, and ...
In summary, the COVID-19 virus variants are not simply competing. They are working together by building on each other's changes, balancing extreme effects, and ...

8) ... adapting to different environments in a coordinated way. This cooperative evolution allows the virus to rapidly diversify and adapt as a united population.
Thanks for reading 🙏 and happy safe year 2025 😷

Thanks for reading 🙏 and happy safe year 2025 😷


• • •
Missing some Tweet in this thread? You can try to
force a refresh






















