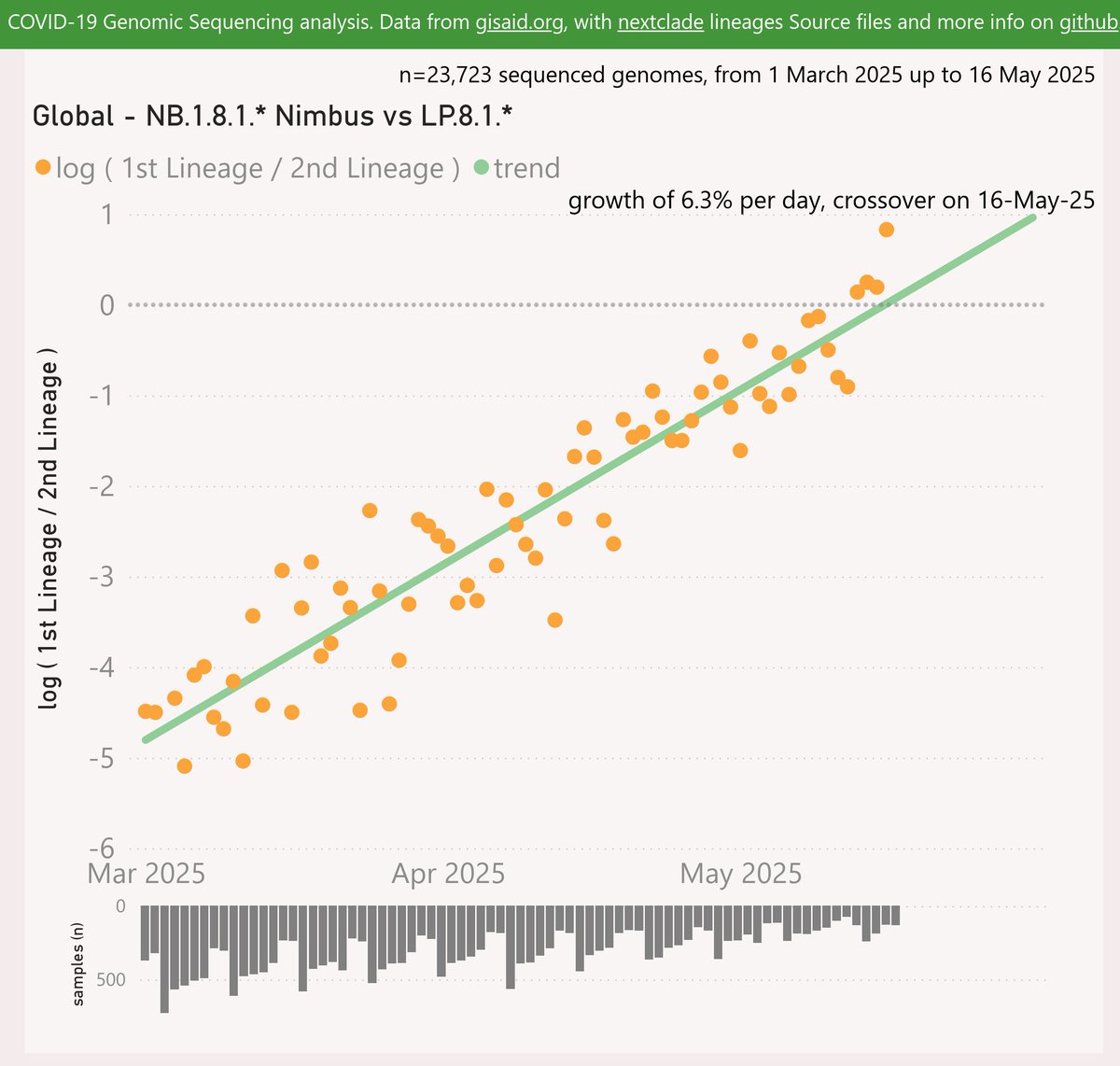
XFP is the latest recombinant variant to be classified and incredibly it is the third of a recent set of recombinants with identical spike mutations. The earlier ones were XFJ and XFM.
I think of them as the "Doppelgängers".
#COVID19 #XFJ #XFM #XFP #Doppelgängers
🧵
I think of them as the "Doppelgängers".
#COVID19 #XFJ #XFM #XFP #Doppelgängers
🧵

Note "Doppelgänger" is not an agreed variant nickname, nor should it be. There doesn’t seem to be anything stopping other sets of variants or recombinants from ending up the same way, as multiple Doppelgänger packs.
🧵
🧵
Also note their non-Spike mutations do vary. But the Spike proteins of SARS-CoV-2 are the focus of much study and explain the success of most variants.
🧵
🧵
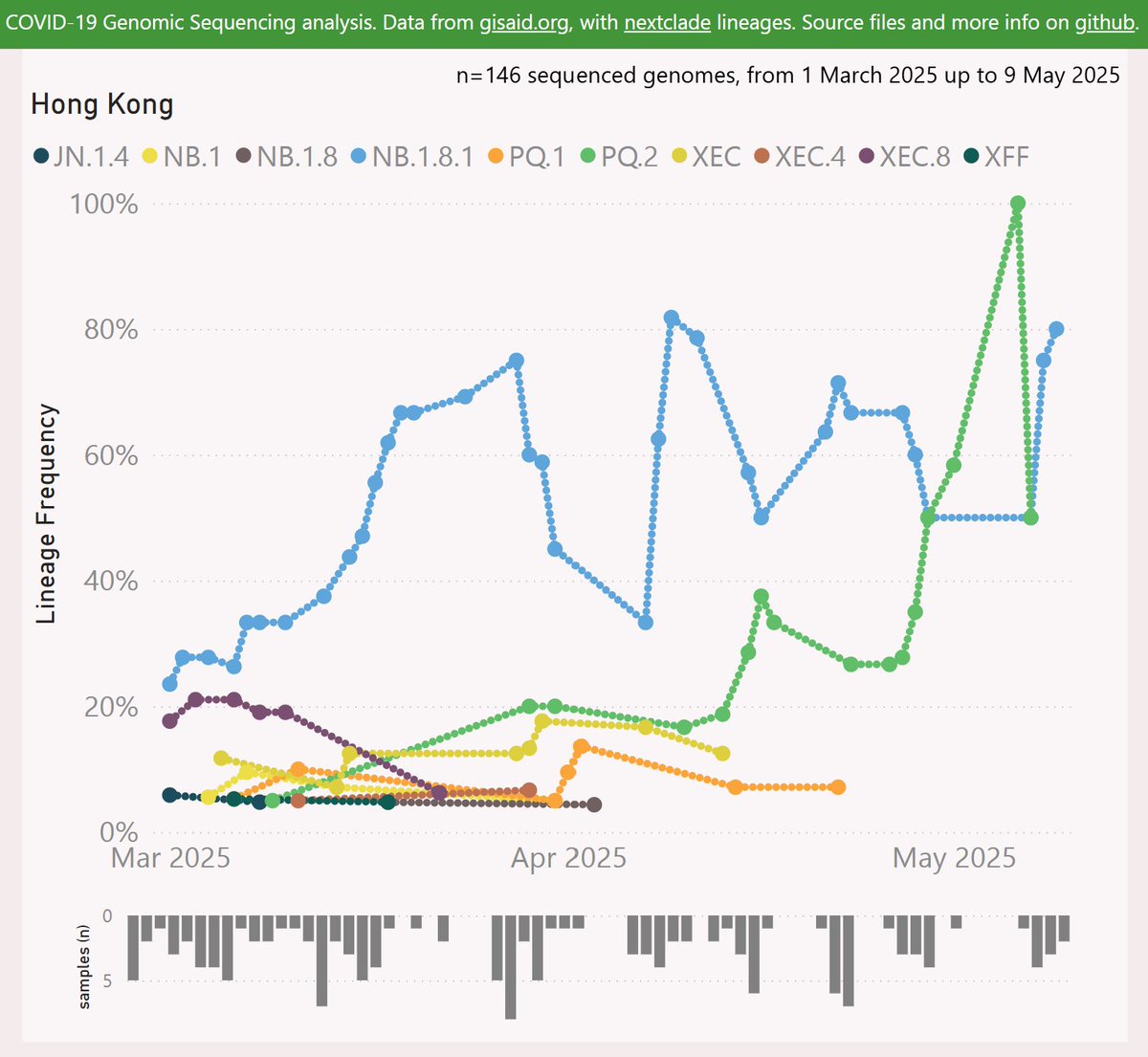
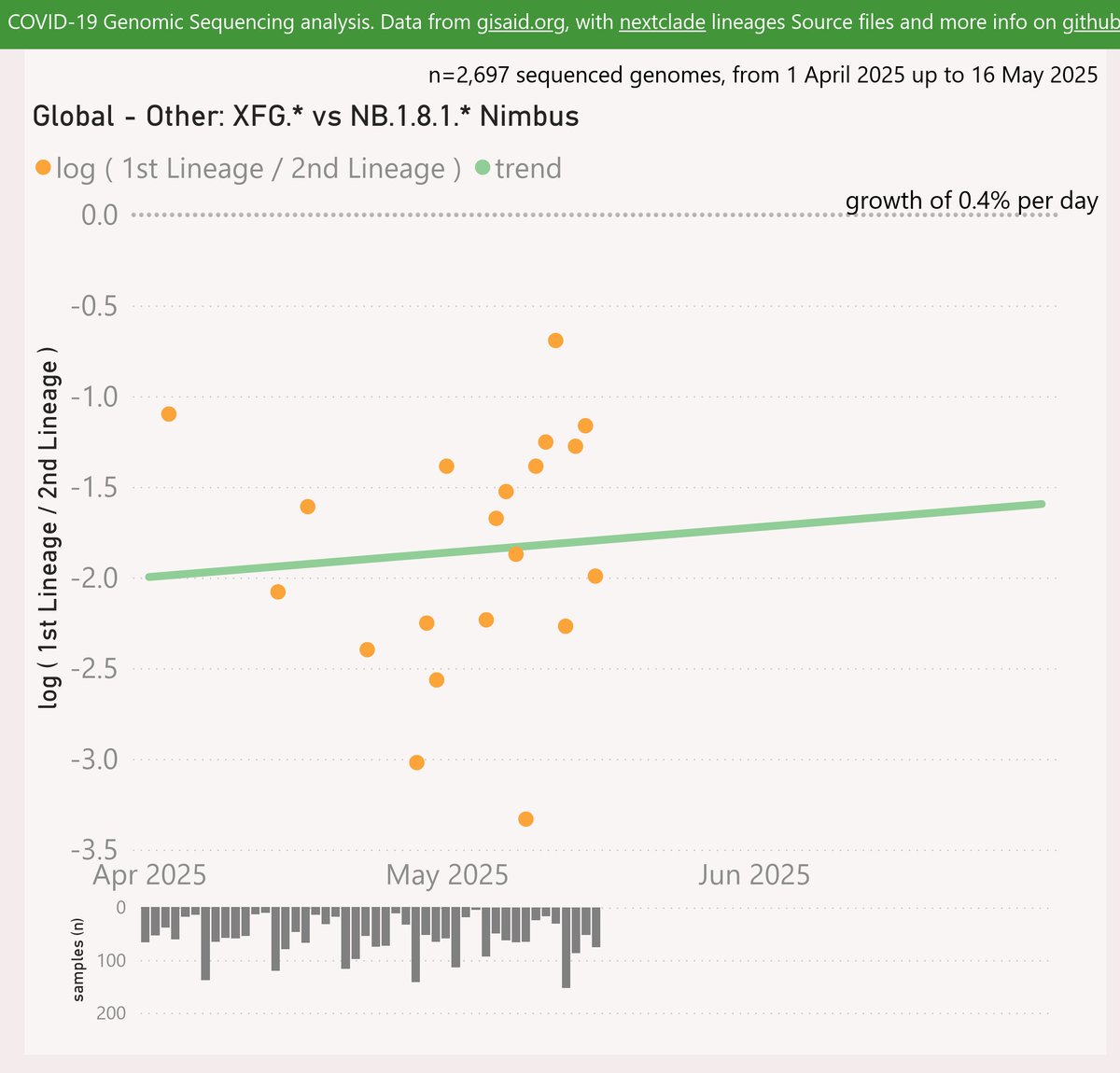
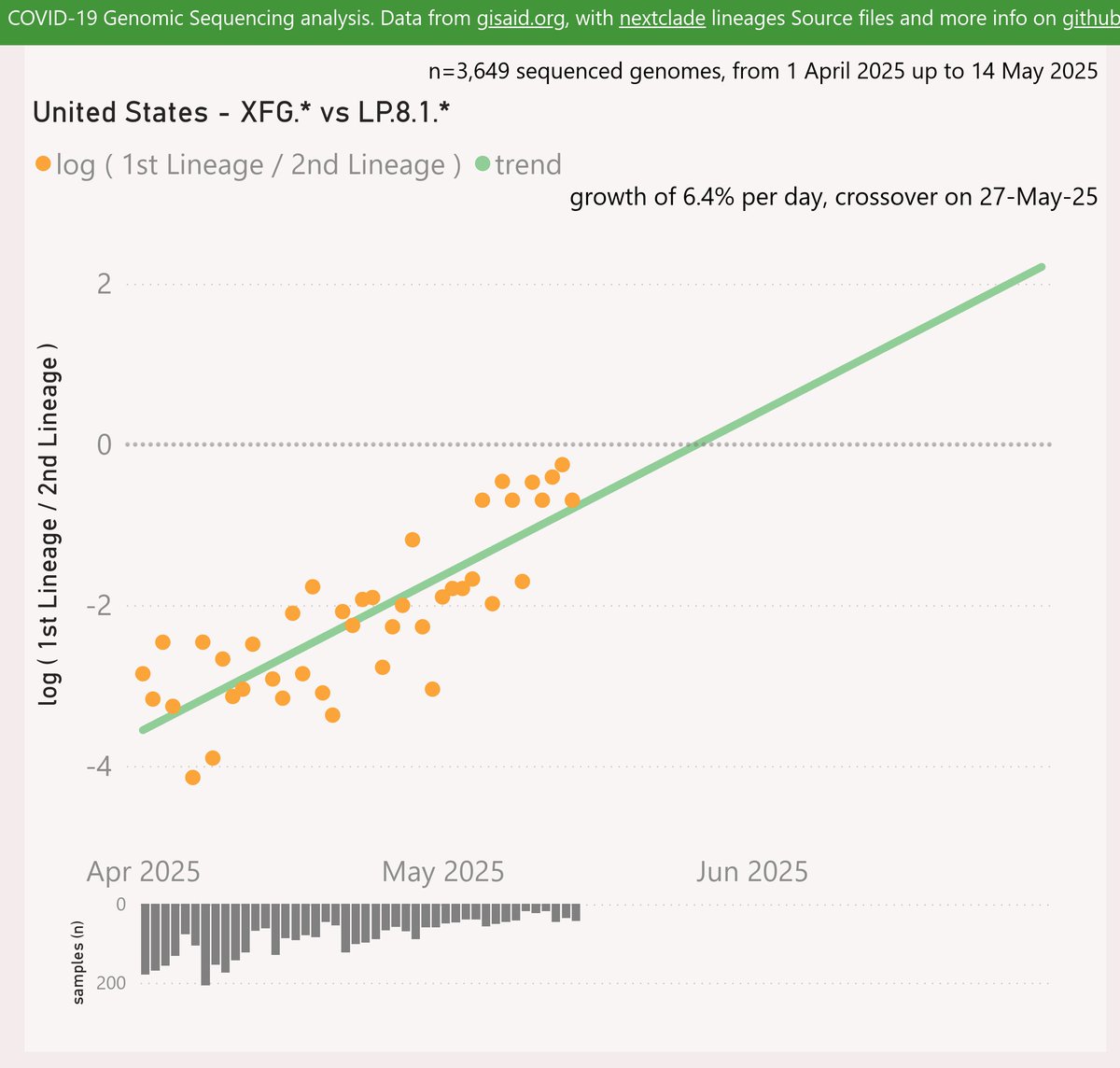
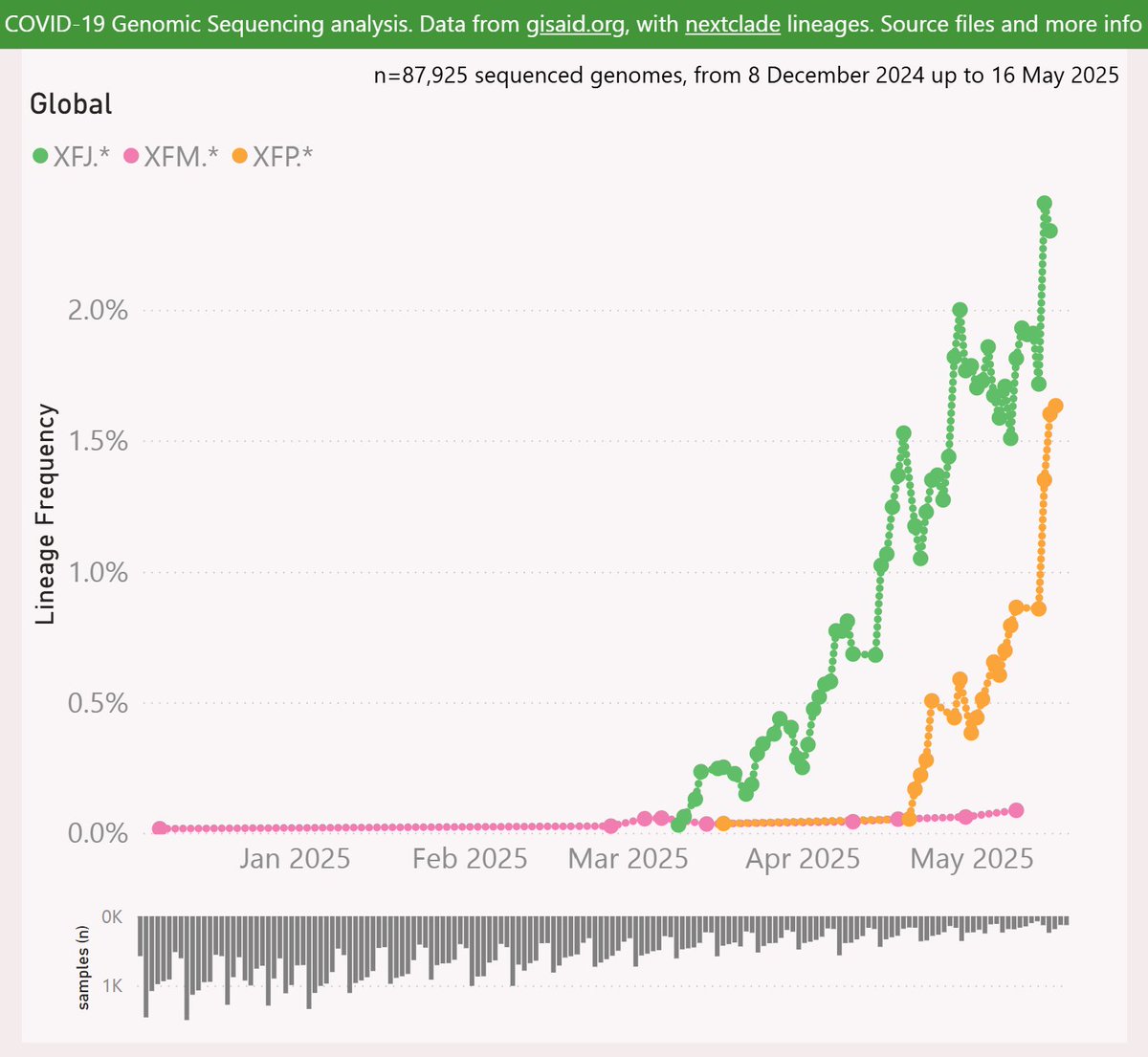
Here’s the growth shown by the 3 Doppelgängers (in this pack). XFM has the earliest detection, but it has not been successful.
XFJ was next and has grown quite strongly to over 2% globally.
XFP has shown even faster growth to over 1.5%.
🧵
XFJ was next and has grown quite strongly to over 2% globally.
XFP has shown even faster growth to over 1.5%.
🧵
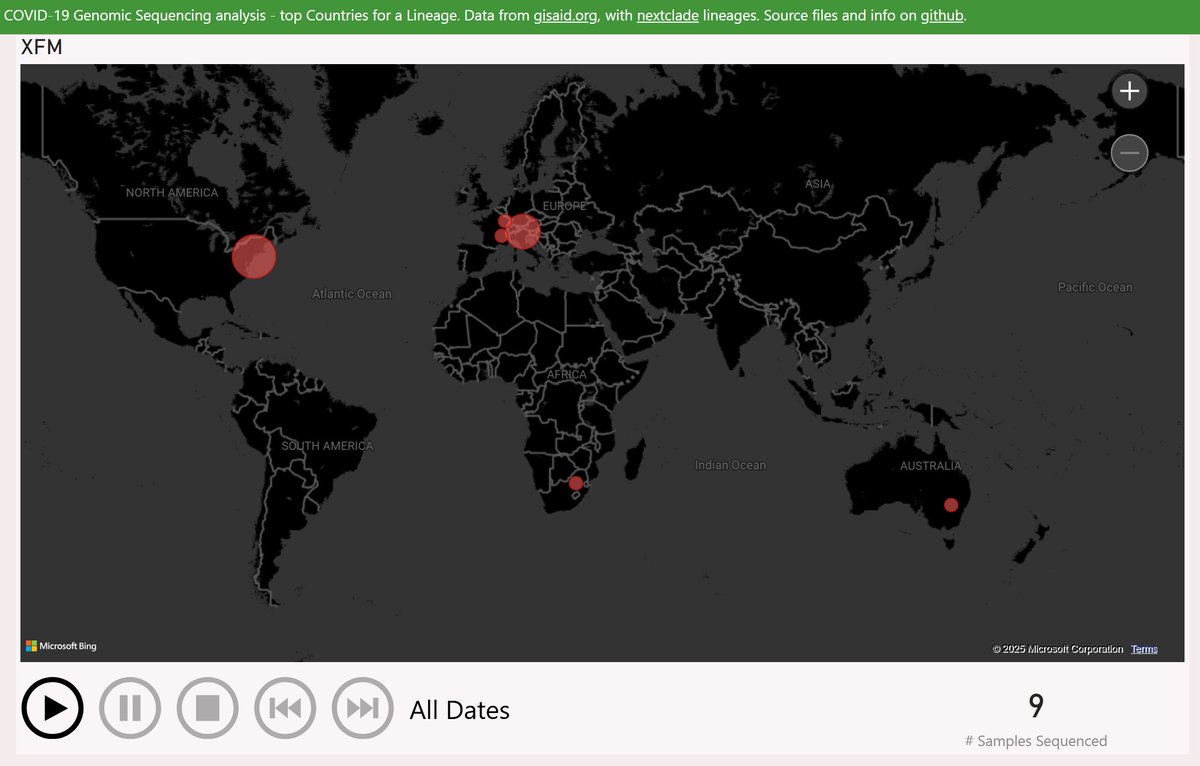
XFM is a recombinant of LF.7.6.2 (descended from JN.1.16.1) and LS.2 (descended from JN.1.18.5).
The earliest XFM sample was reported from South Africa in December.
After a long gap, the remaining samples have mostly been from New York and alpine regions of France and Italy.
🧵
The earliest XFM sample was reported from South Africa in December.
After a long gap, the remaining samples have mostly been from New York and alpine regions of France and Italy.
🧵
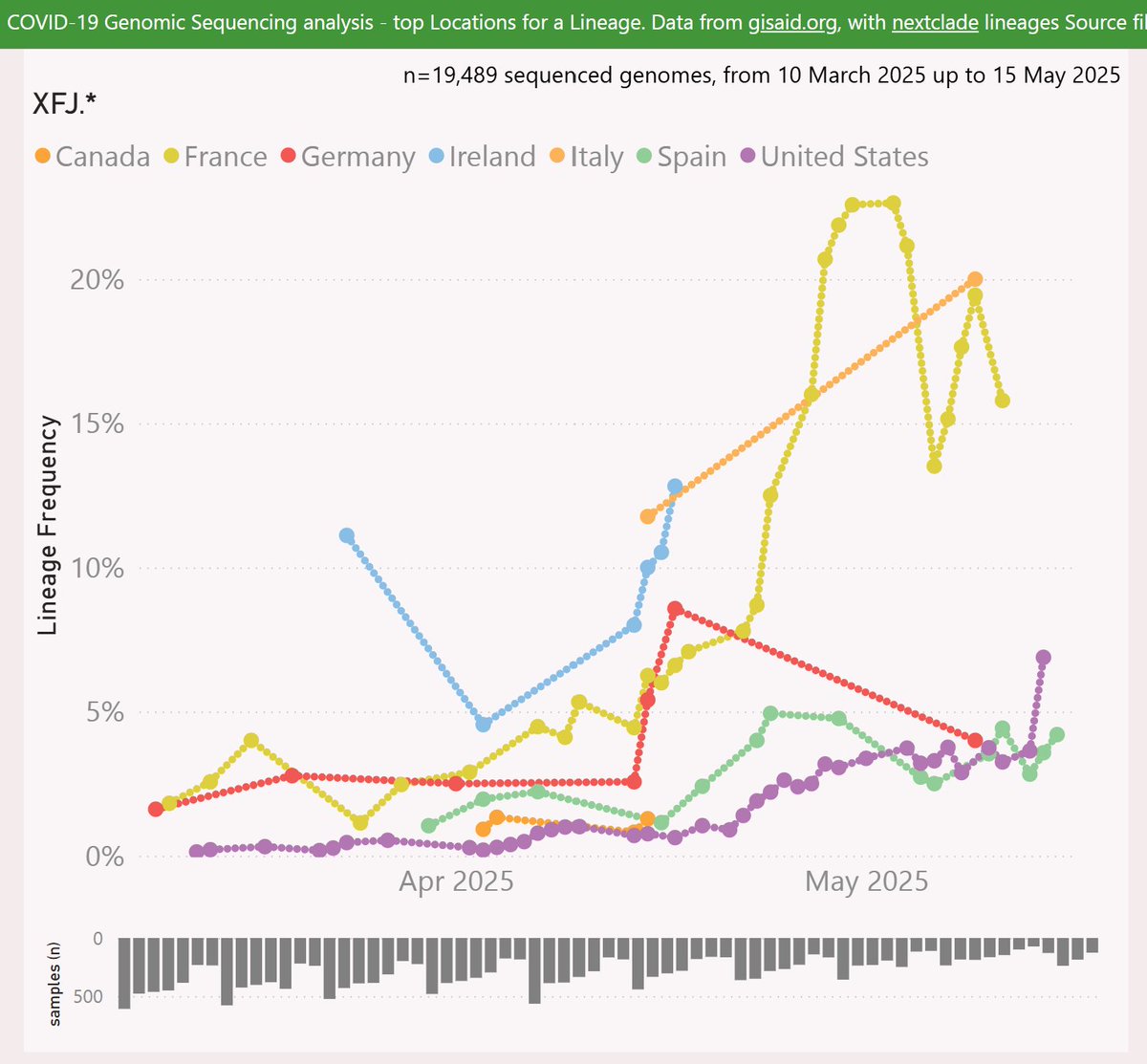
XFJ is a recombinant of LF.7.2 (descended from JN.1.16.1) and LS.2 (descended from JN.1.18.5).
The earliest XFJ sample was reported from Germany, in mid-March.
The samples reported have mostly been from Europe and the North-East US.
🧵
The earliest XFJ sample was reported from Germany, in mid-March.
The samples reported have mostly been from Europe and the North-East US.
🧵
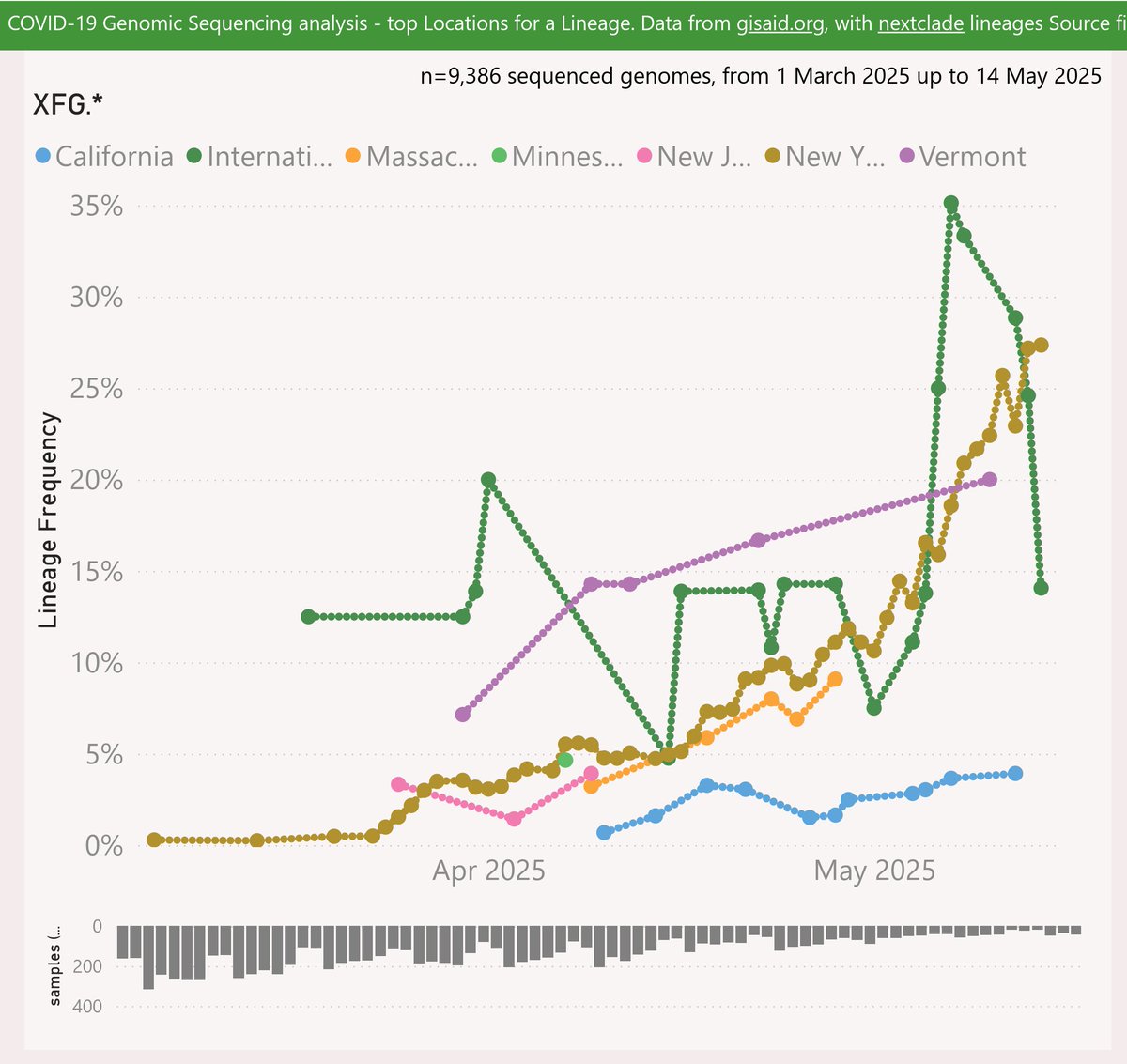
XFP is a recombinant of LF.7 (descended from JN.1.16.1) and LS.2 (descended from JN.1.18.5).
The earliest XFP sample was reported from Washington DC, in mid-March. Almost all the US samples detected since have been International Travellers arriving from India.
🧵
The earliest XFP sample was reported from Washington DC, in mid-March. Almost all the US samples detected since have been International Travellers arriving from India.
🧵
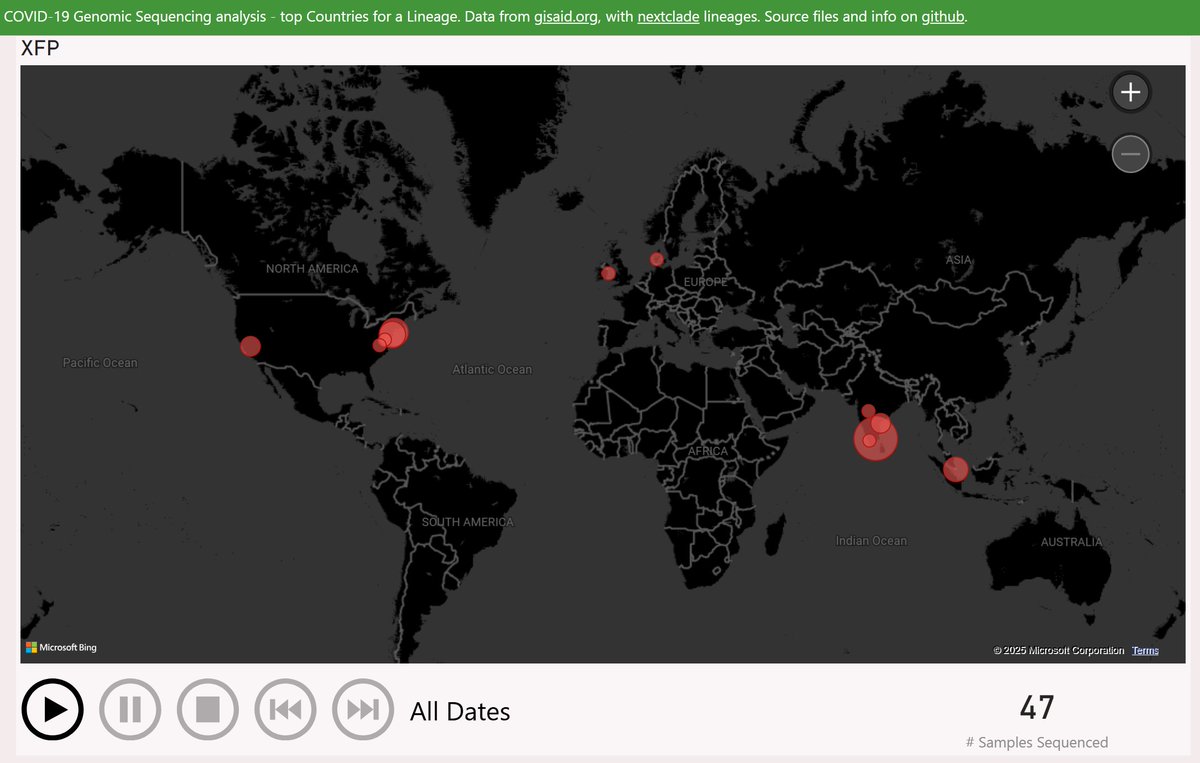
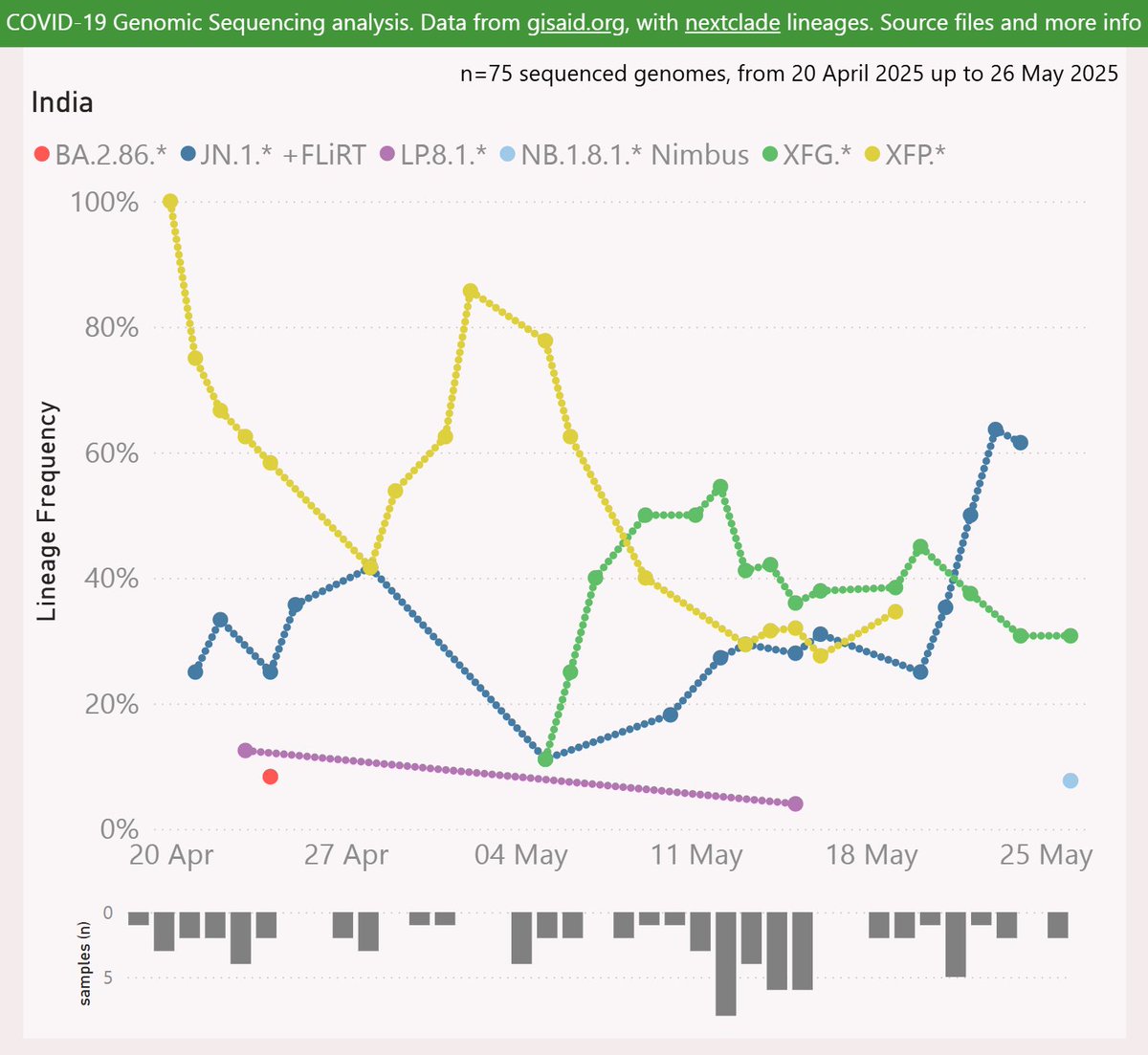
When India itself resumed sharing samples recently, XFP appeared to be dominant there from at least mid-April. So XFP is presumed to have originated in India, during the many months when there was no sample data shared.
🧵
🧵
The Doppelgängers offer a window into the historically unprecedented global variant generation engine we have constructed. I think of it as a flywheel, with tremendous momentum from the billions of infections and millions of chronic cases that are churning on year after year.
🧵
🧵
It’s not a totally random process, but I find it breathtaking to ponder the odds that 3 identical recombinations were produced in 3 separate places, 3 different ways, over just a few months.
🧵
🧵
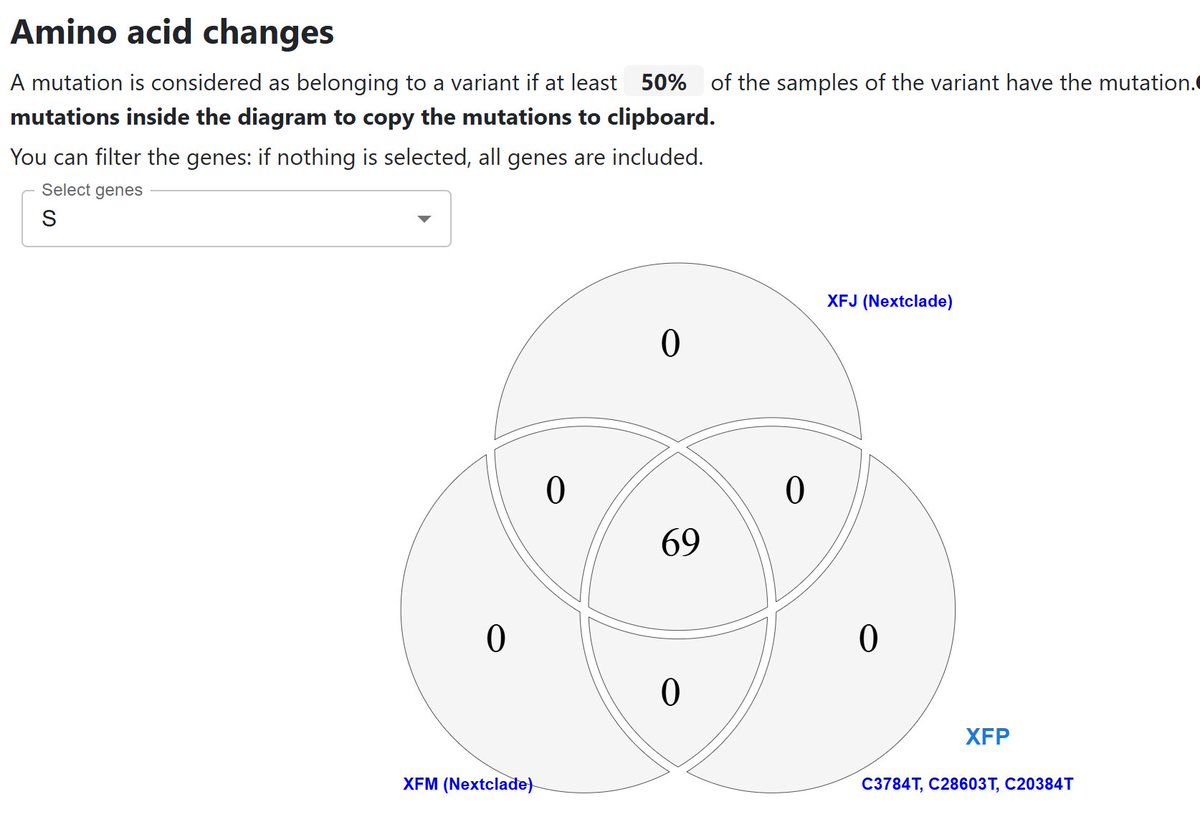

There are ~1200 positions in the spike protein of SARS-CoV-2, each with ~20 possible letters, so ~24,000 possible combinations.
🧵
🧵
Interactive genomic sequencing dataviz, code, acknowledgements and more info here:
🧵 endsgithub.com/Mike-Honey/cov…
🧵 endsgithub.com/Mike-Honey/cov…
• • •
Missing some Tweet in this thread? You can try to
force a refresh