BREAKING🔔 The 53th paper from G2P-Japan🇯🇵 is out at Lancet Infectious Diseases @TheLancetInfDis. We elucidated the virological characteristics of SARS-CoV-2 NB.1.8.1. Please repost🔥 1/
thelancet.com/journals/lanin…
thelancet.com/journals/lanin…
NB.1.8.1 is a descendant strain of XDV (XDV is a recombinant of XDE, FL.13.4 and JN.1) and is currently spreading in Asian countries. 2/ 

NB.1.8.1 has 7 mutations in spike and 23 mutations in the non-spike region compared to JN.1. Compared to XEC spike, NB.1.8.1 spike has 4 substitutions, G184S, K478I, A435S and L1104V. 3/ 

Transmissibility (Re, relative effective reproduction number):
The Re of NB.1.8.1 was ~1.1-1.2-fold higher than that of LP.8.1, a major variant recently. This suggests that NB.1.8.1 has the potential to become the major variant in the near future. 4/
The Re of NB.1.8.1 was ~1.1-1.2-fold higher than that of LP.8.1, a major variant recently. This suggests that NB.1.8.1 has the potential to become the major variant in the near future. 4/

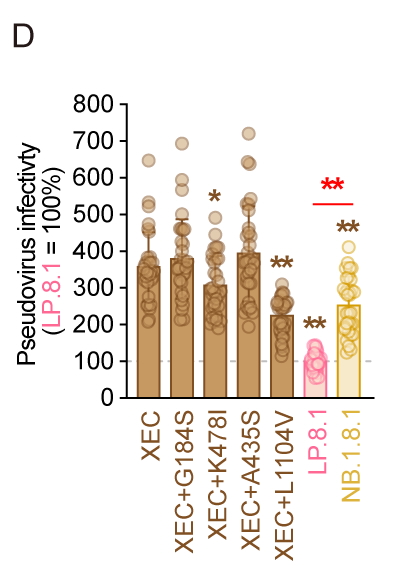
Pseudovirus infectivity:
The infectivity of NB.1.8.1 was higher than that of LP.8.1 but lower than that of XEC. Compared to XEC, K478I and L1104V were the substitutions contributing to lower infectivity. 5/
The infectivity of NB.1.8.1 was higher than that of LP.8.1 but lower than that of XEC. Compared to XEC, K478I and L1104V were the substitutions contributing to lower infectivity. 5/
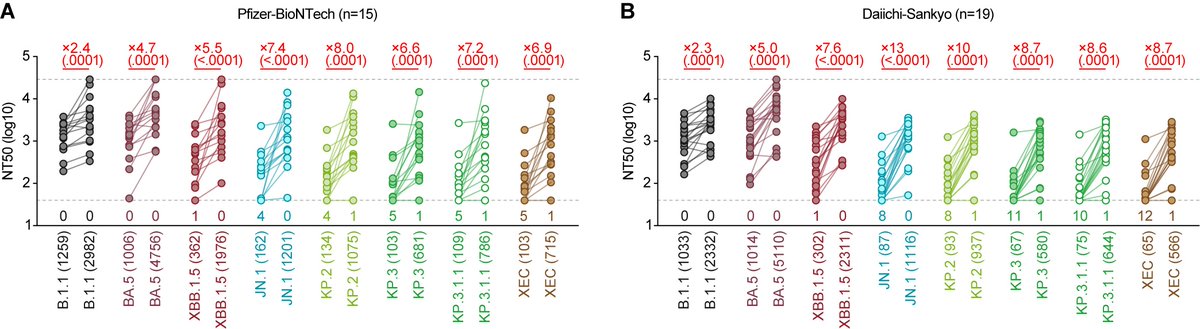
Immune resistance:
50% neutralizing titer of NB.1.8.1 in both XEC convalescent sera and JN.1 vaccine sera was similar to that of XEC and LP.8.1🤔 6/
50% neutralizing titer of NB.1.8.1 in both XEC convalescent sera and JN.1 vaccine sera was similar to that of XEC and LP.8.1🤔 6/

In sum, the advantage of NB.1.8.1 over LP.8.1 may be attributed to its higher infectivity. However,
1, Some data are inconsistent with another paper by @yunlong_cao
2, There are many mutations other than spike.
Therefore, more detailed study will be required. 7/
1, Some data are inconsistent with another paper by @yunlong_cao
2, There are many mutations other than spike.
Therefore, more detailed study will be required. 7/
• • •
Missing some Tweet in this thread? You can try to
force a refresh