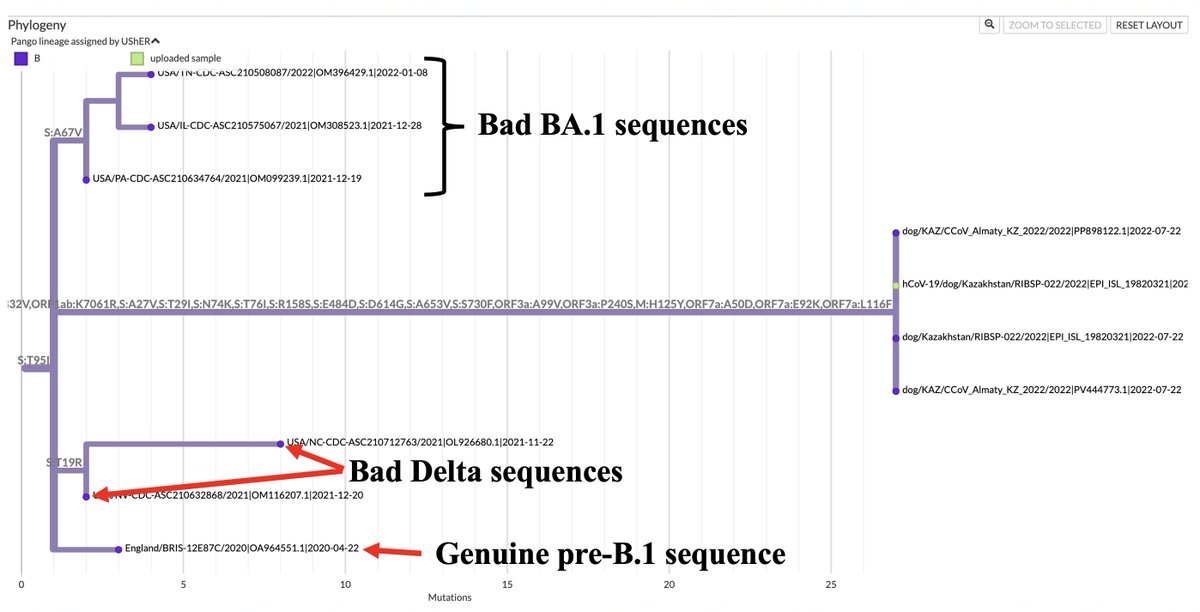
Interesting recombinant showed up today from Texas. It's a mixture of B.1.595, BA.1, and some flavor of JN.1. Most of the genome is from B.1.595. The ancestry of this one is clear: it directly descends from a B.1.595 sequence collected in January 2023, also in Texas. 1/11 

When the B.1.595 was collected this infection was >1 yr old, w/no sign of Omicron. BA.1 ceased circulating ~1 year prior.
Now a BA.1 spike appears w/just 5 changes from baseline BA.1, none in the RBD—S12F, T76I, Q271K, R765H, S939F.
This is a zombie BA.1 spike. 2/
Now a BA.1 spike appears w/just 5 changes from baseline BA.1, none in the RBD—S12F, T76I, Q271K, R765H, S939F.
This is a zombie BA.1 spike. 2/

There are only a few signs of JN.1, & they're scattered. In ORF1a, we see JN.1's V3593F, P3395H, & R3821K, but the NSP6 deletion btwn these—universal in Omicron—is absent. In
M has JN.1's D3H + T30A & E19Q (in JN.1 & BA.1), yet A63T—also in both BA.1 & JN.1 is absent. 3/11
M has JN.1's D3H + T30A & E19Q (in JN.1 & BA.1), yet A63T—also in both BA.1 & JN.1 is absent. 3/11

Oddly, it has the 3-nuc, loss-of-function ORF6:D61L—from JN.1 since BA.1 lacks it.
It also took N:P13L & ∆31-33 from BA.1/JN.1 but lacks N:R203K/G204R (& therefor N*).
Finally, it has the standard ∆27934-29759 s2m deletion, which all BA.2 descendants have. 4/11
It also took N:P13L & ∆31-33 from BA.1/JN.1 but lacks N:R203K/G204R (& therefor N*).
Finally, it has the standard ∆27934-29759 s2m deletion, which all BA.2 descendants have. 4/11

Also odd: None of the usual convergent chronic-infection muts are here. The Jan 2023 B.1.595 spike got 1 non-RBD spike mut—P1162S—in ~15 months & in ~2.5 years, the BA.1 spike has added just 4-5 subs, none in the RBD.
There's no sign of combat with an immune system. 5/11
There's no sign of combat with an immune system. 5/11
It's nearly certain all these sequences are from the same person. When the B.1.595 sequence was collected in 2023, they were also infected with BA.1, but no sign of it showed up in the sequence.
Where was it hiding out? In the deep lung? In the GI tract? 6/11
Where was it hiding out? In the deep lung? In the GI tract? 6/11
But viruses that have spent long periods of time in the GI tract have distinct mutational signatures, as @SolidEvidence has documented. I've also discovered a distinct mutational pattern from bronchoalveolar lavage sequences (deep lung).
This sequence has none of them. 7/11
This sequence has none of them. 7/11
There's no chance this virus spreads in the current environment. But cases like this might have relevance in the long term. As immunity to successive Omicron variants grows, old variants may become newly fit.
But they long ago vanished.... or have they? 8/11
But they long ago vanished.... or have they? 8/11
Apparently not. Usually, when old variants reappear after years, their spikes have transformed.
But this a 4.5-year-long infection that's never had a single RBD mutation.
In 5-10 yrs, something close to ancestral spike could be newly fit, esp. in children. 9/11
But this a 4.5-year-long infection that's never had a single RBD mutation.
In 5-10 yrs, something close to ancestral spike could be newly fit, esp. in children. 9/11
Could an ancient spike reappear at that time & take over? Something like this happens with norovirus. Norovirus variants that circulated years/decades ago sometimes reappear, relatively unchanged, & proceed to sweep the world. See paper by Chris Ruis. 10/ academic.oup.com/ve/article/6/2…
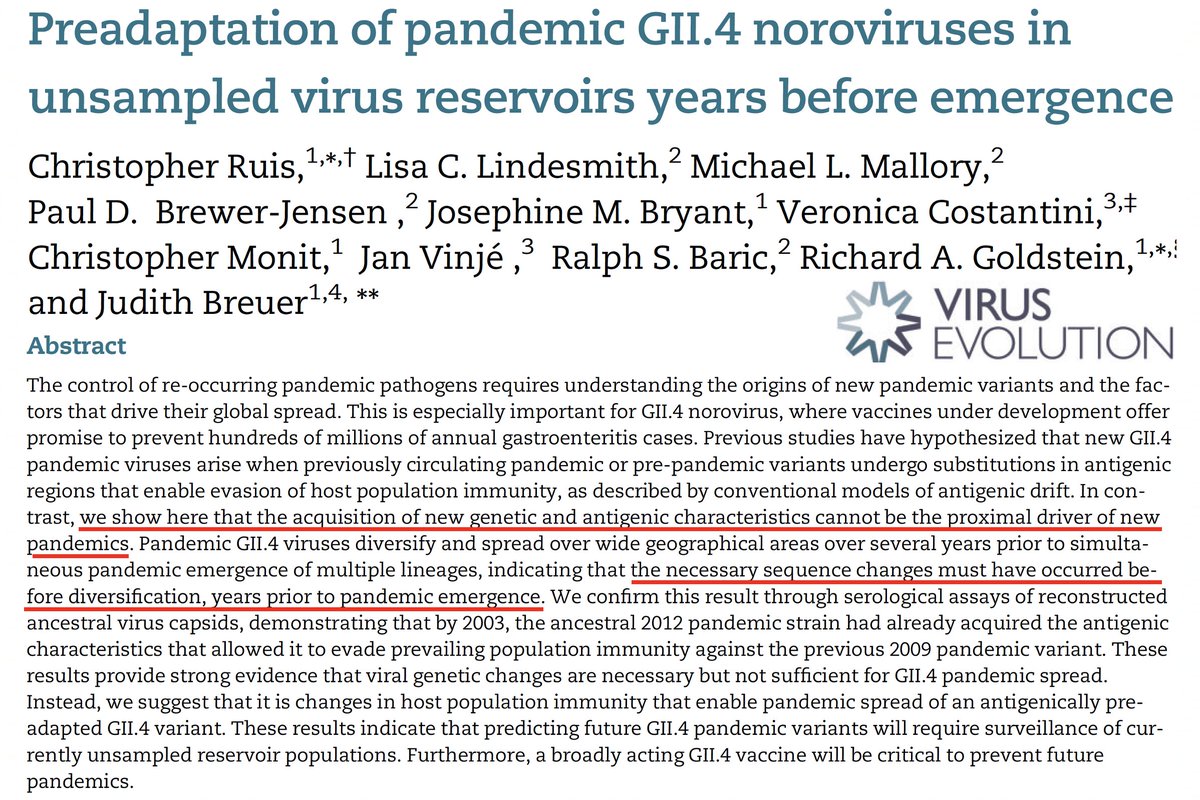

We've not seen this. When a new SARS-2 variant sweeps, it's honed its Ab-evasion skills in years of combat w/an immune system too weak to clear it but strong enough to challenge it. But in the long term, who knows? Maybe we'll see a zombie-spike sweep someday. 🧟♂️ 11/end
• • •
Missing some Tweet in this thread? You can try to
force a refresh