
Teacher
"Be ruthless with systems and be kind to people."
Michael Brooks, 1983-2020
How to get URL link on X (Twitter) App


 If you've missed the story about how BA.3.2 (a novel, divergent saltation variant) is hugely overrepresented in sequences from children, this was my original (very quick) analysis, which subsequent data extended & confirmed. 2/
If you've missed the story about how BA.3.2 (a novel, divergent saltation variant) is hugely overrepresented in sequences from children, this was my original (very quick) analysis, which subsequent data extended & confirmed. 2/https://x.com/LongDesertTrain/status/2036267271849382107

https://twitter.com/LongDesertTrain/status/2036291148193349739I've tried to make sense of BA.3.2's penchant for kids by considering its unique spike: more compact, more closed, & more antibody-evasive than any other variant.

https://twitter.com/SolidEvidence/status/2035872639118344466I suspect that the number of people continuously infected since 2020 or 2021 is much larger than we realize. It's impossible to prove, but there are case studies where a chronically infected person gets infected by a new variant, which drives out the original virus...
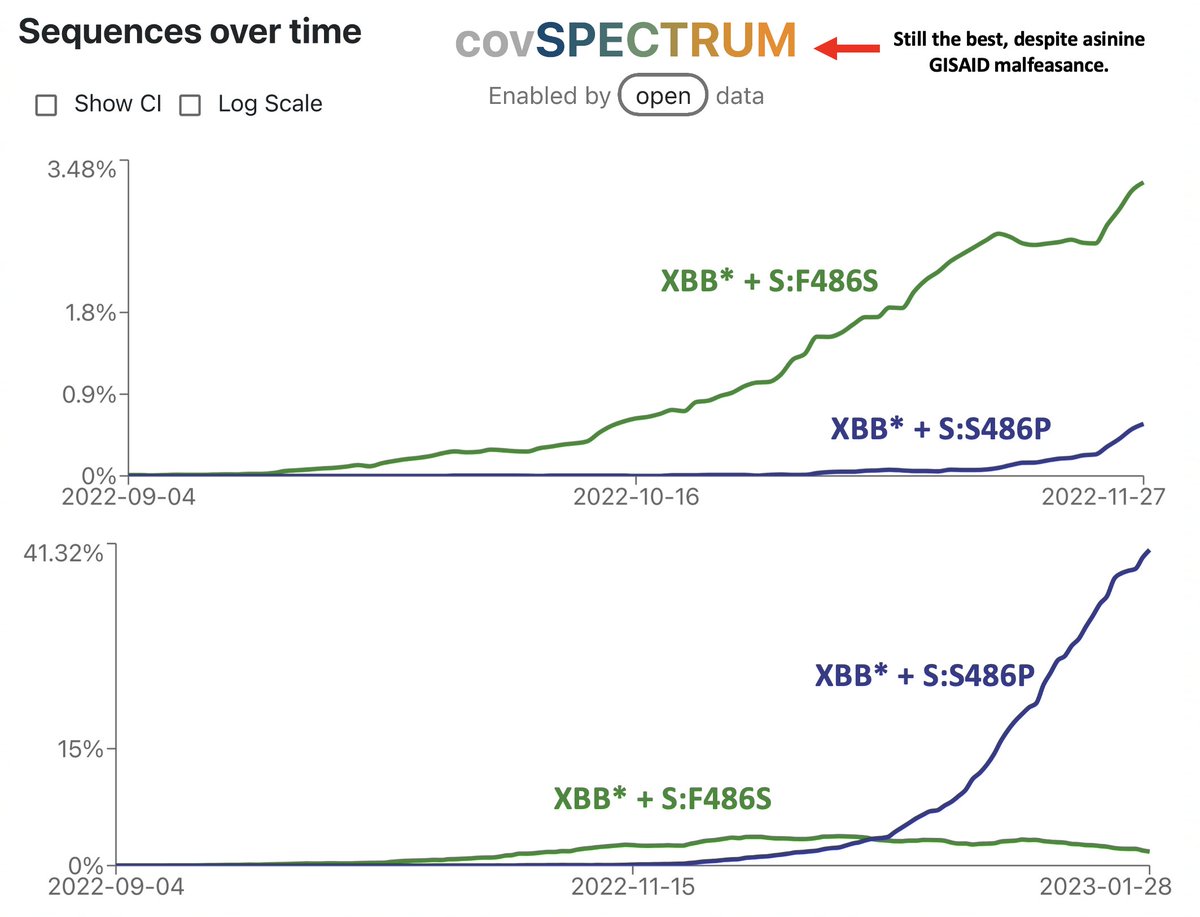
https://twitter.com/snpoehlm/status/2034920940514267424Until now, the broad pattern of SARS-2 evolution has been:
https://twitter.com/Tuliodna/status/2005638352381571577I would normally write a summary 🧵 of the BA.3.2 mutational analysis here, but as much of my contribution parallels my previous BA.3.2 threads I'll just link to those here, w/brief descriptions of each.
https://x.com/LongDesertTrain/status/1899647059872850369
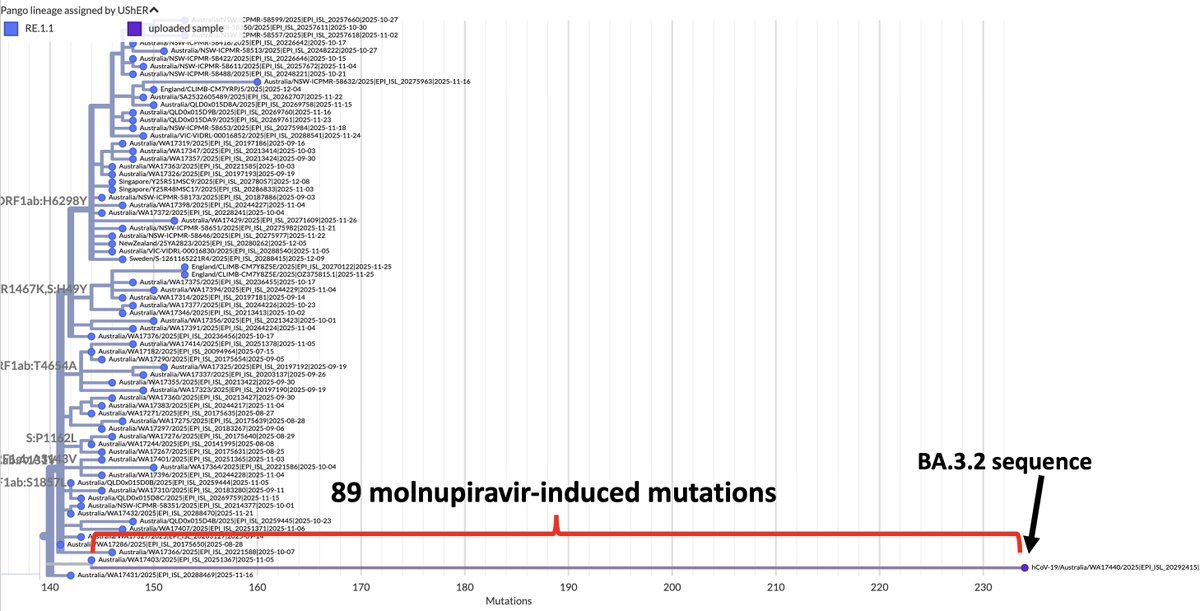
 Background on MOV: It's a mutagenic drug. Its purpose is to cause so many mutations that the virus becomes unviable & is cleared. But we've long known this often does not happen. Instead, the virus persists in highly mutated form & can be transmitted. 2/
Background on MOV: It's a mutagenic drug. Its purpose is to cause so many mutations that the virus becomes unviable & is cleared. But we've long known this often does not happen. Instead, the virus persists in highly mutated form & can be transmitted. 2/https://x.com/LongDesertTrain/status/1577256143281233921

https://twitter.com/JPWeiland/status/2002487175884255270I use CovSpectrum & Nextstrain every day—& I'm not the only one. Every Covid thread I've ever posted here has relied partly on CovSpectrum & Nextstrain for information & visuals. These vital tools have now been stolen from us by a world-class grifter. 2/ thinkglobalhealth.org/article/to-fin…

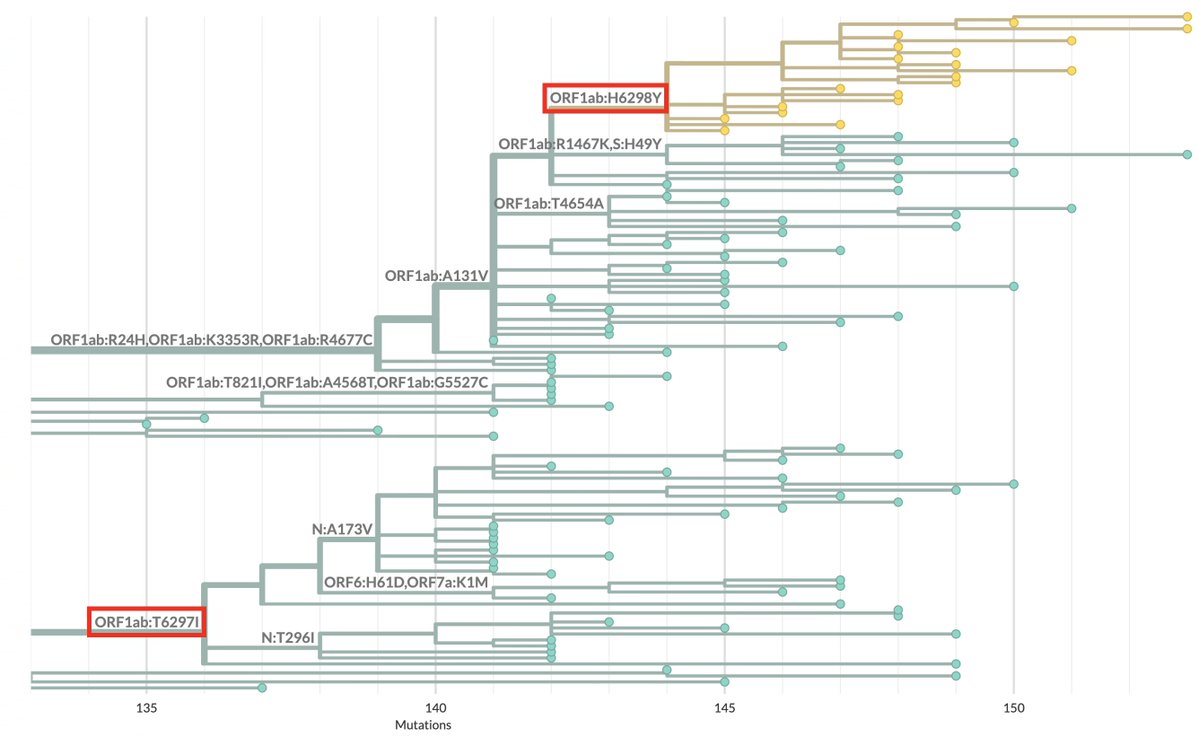
https://twitter.com/JosetteSchoenma/status/1998154156138545491One interesting (and possibly coincidental) aspect of the BA.3.2 tree: Two large branches have NSP14 mutations at adjacent AA residues—ORF1b:T1896I and ORF1b:H1897Y. 2/4

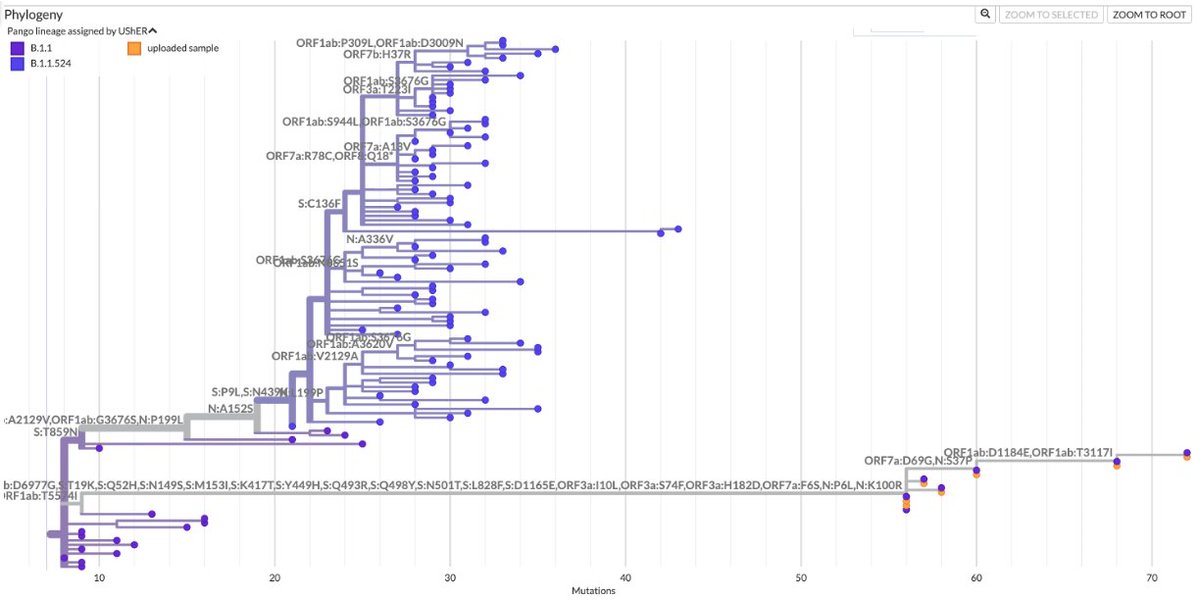
https://twitter.com/SolidEvidence/status/1991888519112093740The first instance involved a small cluster of sequences that hospitalized several people & resulted in the death of a young child in early 2022. More on this one later. 2/15

https://twitter.com/snpoehlm/status/1985998506365231105@StuartTurville has pointed out that WA delayed Covid spread longer than elsewhere in Australia. China has a somewhat similar immune history (as do other SE Asian countries). Perhaps BA.3.2 will do well in China once it arrives there? 2/4
https://x.com/StuartTurville/status/1983308264407503168
https://twitter.com/snpoehlm/status/1980953067777368569
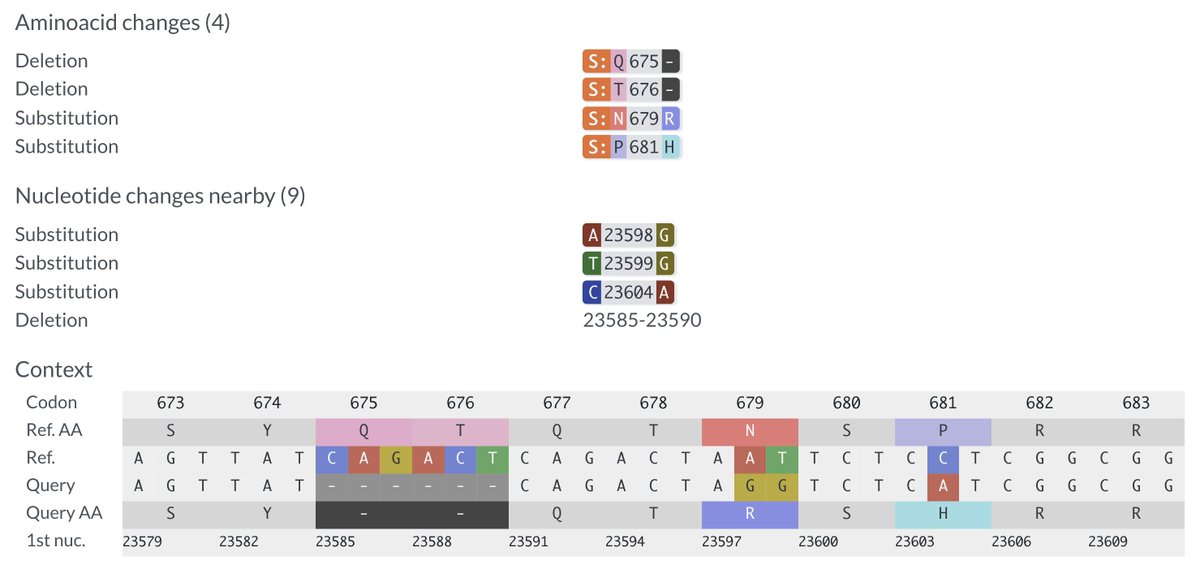
 Work by @TheMenacheryLab looked at a similar, more extensive, deletion. They deleted both QT repeats plus the next AA (∆QTQTN). In Vero cells (monkey kidney cells), it produced extra-large plaques & outcompeted WT virus—similar to furin cleavage site (FCS)-deletion mutants. 2/12
Work by @TheMenacheryLab looked at a similar, more extensive, deletion. They deleted both QT repeats plus the next AA (∆QTQTN). In Vero cells (monkey kidney cells), it produced extra-large plaques & outcompeted WT virus—similar to furin cleavage site (FCS)-deletion mutants. 2/12 
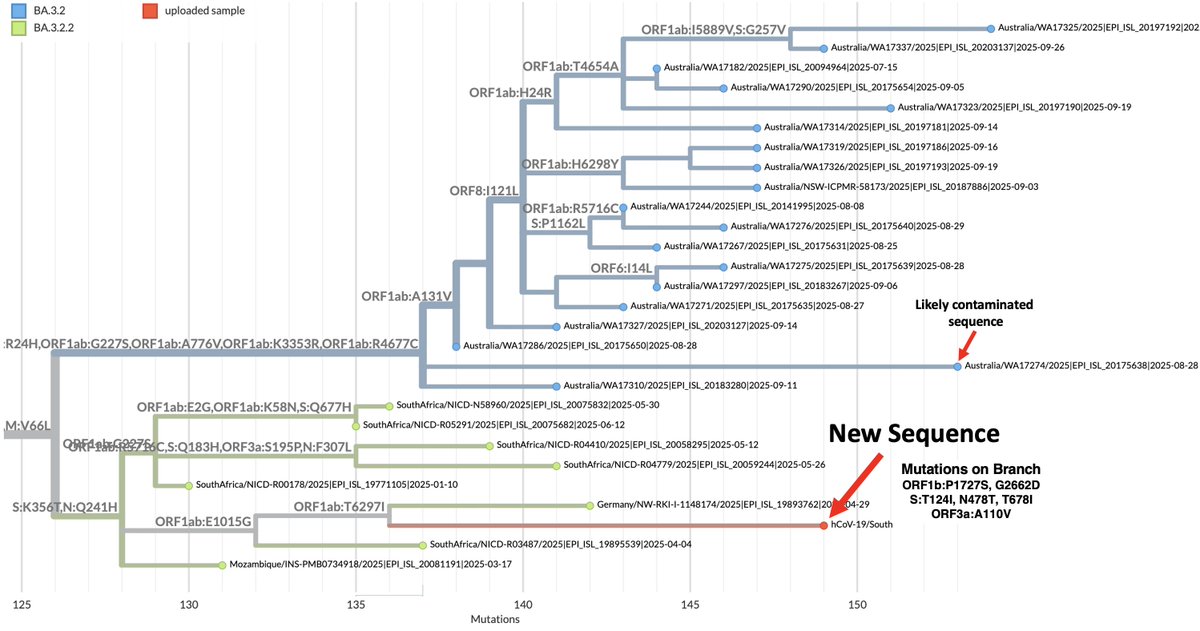
 For those not following closely, here's a 🧵 I made about BA.3.2 (not yet designated at the time) that I made some months ago, when it first burst upon the scene. 2/7
For those not following closely, here's a 🧵 I made about BA.3.2 (not yet designated at the time) that I made some months ago, when it first burst upon the scene. 2/7https://x.com/LongDesertTrain/status/1899647059872850369

 In South America, this may have already happened. Recent sequences are scarce, but they nearly all have some sort of FCS-weakening mutation, mostly S:S680P in XFG.3.4.1, but with several others (S680F, S680Y, R683Q, R683W) contributing as well. 2/4
In South America, this may have already happened. Recent sequences are scarce, but they nearly all have some sort of FCS-weakening mutation, mostly S:S680P in XFG.3.4.1, but with several others (S680F, S680Y, R683Q, R683W) contributing as well. 2/4 

https://twitter.com/LongDesertTrain/status/1962980575054332049The ever-fertile mind of @Nucleocapsoid proffers the possibility that exosomes could be responsible for viral spread in some tissue reservoirs. I don't know much about this topic and so don't have much to say at the moment, but I'm trying to l learn. 2/
https://x.com/Nucleocapsoid/status/1962978769184153687

 First, a brief summary of the relevant parts of the preprint. They examined 30 people (from NIH RECOVER cohort) for 6 months after they had Covid, taking detailed blood immunological markers at 3 time points. 20 had Long Covid (PASC), 10 did not (CONV). 2/ biorxiv.org/content/10.110…
First, a brief summary of the relevant parts of the preprint. They examined 30 people (from NIH RECOVER cohort) for 6 months after they had Covid, taking detailed blood immunological markers at 3 time points. 20 had Long Covid (PASC), 10 did not (CONV). 2/ biorxiv.org/content/10.110…

https://twitter.com/snpoehlm/status/1950476928760037506
 It was collected July 15, & is most closely related to the recent S African seqs from May & June.
It was collected July 15, & is most closely related to the recent S African seqs from May & June.

https://twitter.com/LongDesertTrain/status/1940384545741926911
 Bottom line, in my view: BA.3.2 has spread internationally & is likely growing, but very slowly. If nothing changes, its advantage vs circulating lineages, which seem stuck in an evolutionary rut, will likely gradually grow as immunity to dominant variants solidifies... 2/9
Bottom line, in my view: BA.3.2 has spread internationally & is likely growing, but very slowly. If nothing changes, its advantage vs circulating lineages, which seem stuck in an evolutionary rut, will likely gradually grow as immunity to dominant variants solidifies... 2/9

https://twitter.com/LongDesertTrain/status/18996470598728503692 interesting aspects of the new BA.3.2:


 This was a BA.3.2.1, the branch with S:H681R + S:P1162R (not S:K356T + S:A575S).
This was a BA.3.2.1, the branch with S:H681R + S:P1162R (not S:K356T + S:A575S).


 When the B.1.595 was collected this infection was >1 yr old, w/no sign of Omicron. BA.1 ceased circulating ~1 year prior.
When the B.1.595 was collected this infection was >1 yr old, w/no sign of Omicron. BA.1 ceased circulating ~1 year prior. 