Introducing Silico: the platform for building AI models with the precision of written software.
Silico lets researchers and engineers see inside their models, debug failures, and intentionally design them from the ground up.
Early access is open now. 🧵(1/10)
Silico lets researchers and engineers see inside their models, debug failures, and intentionally design them from the ground up.
Early access is open now. 🧵(1/10)
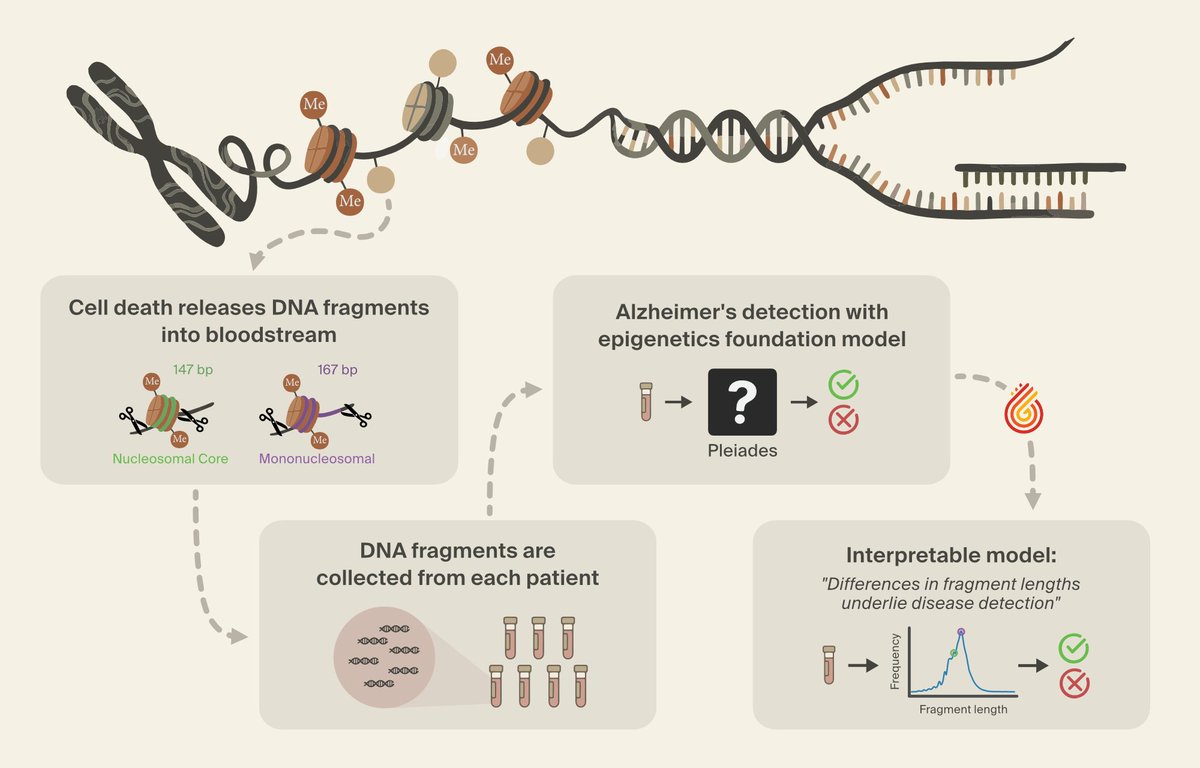
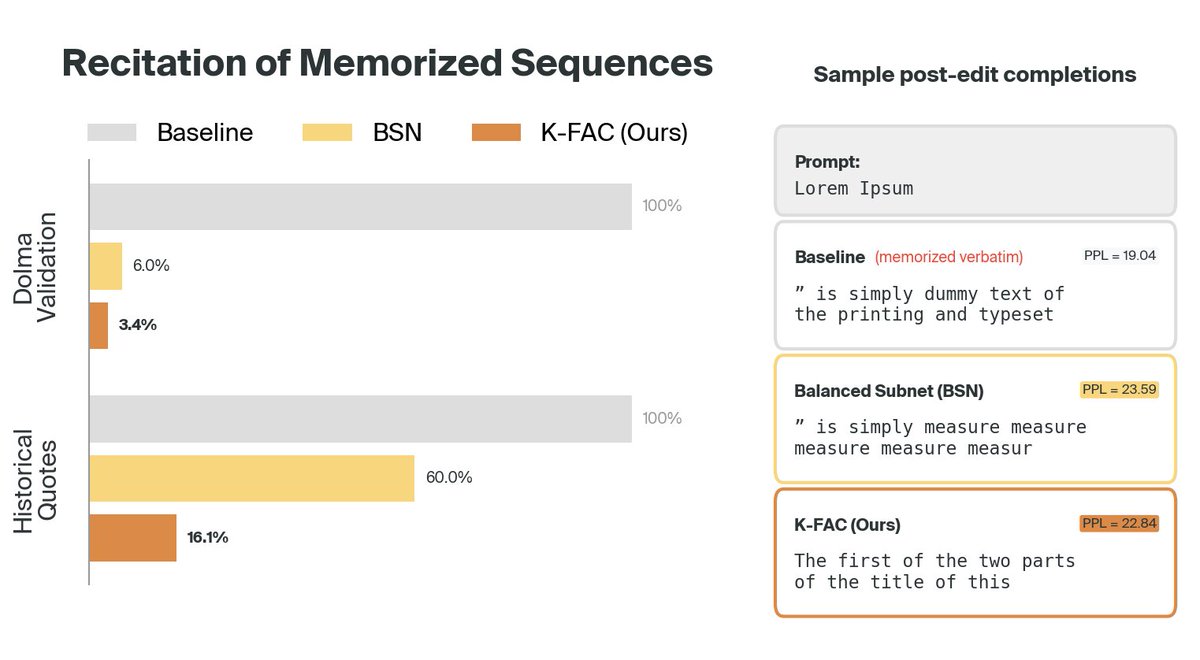
We’ve used interpretability to discover a novel class of Alzheimer’s biomarkers, teach a language model to correct its own hallucinations, and diagnose performance bottlenecks in a robotics model.
Silico brings those frontier techniques to everyone. (2/10)
Silico brings those frontier techniques to everyone. (2/10)
Silico introduces our model neuroscientist: an autonomous agent that plans and runs concurrent experiments on your model.
It works with your team in our model design environment, where you can organize research threads, replicate and extend papers, and collaborate on findings.
Here are 5 things you can do with Silico:
(3/10)
It works with your team in our model design environment, where you can organize research threads, replicate and extend papers, and collaborate on findings.
Here are 5 things you can do with Silico:
(3/10)
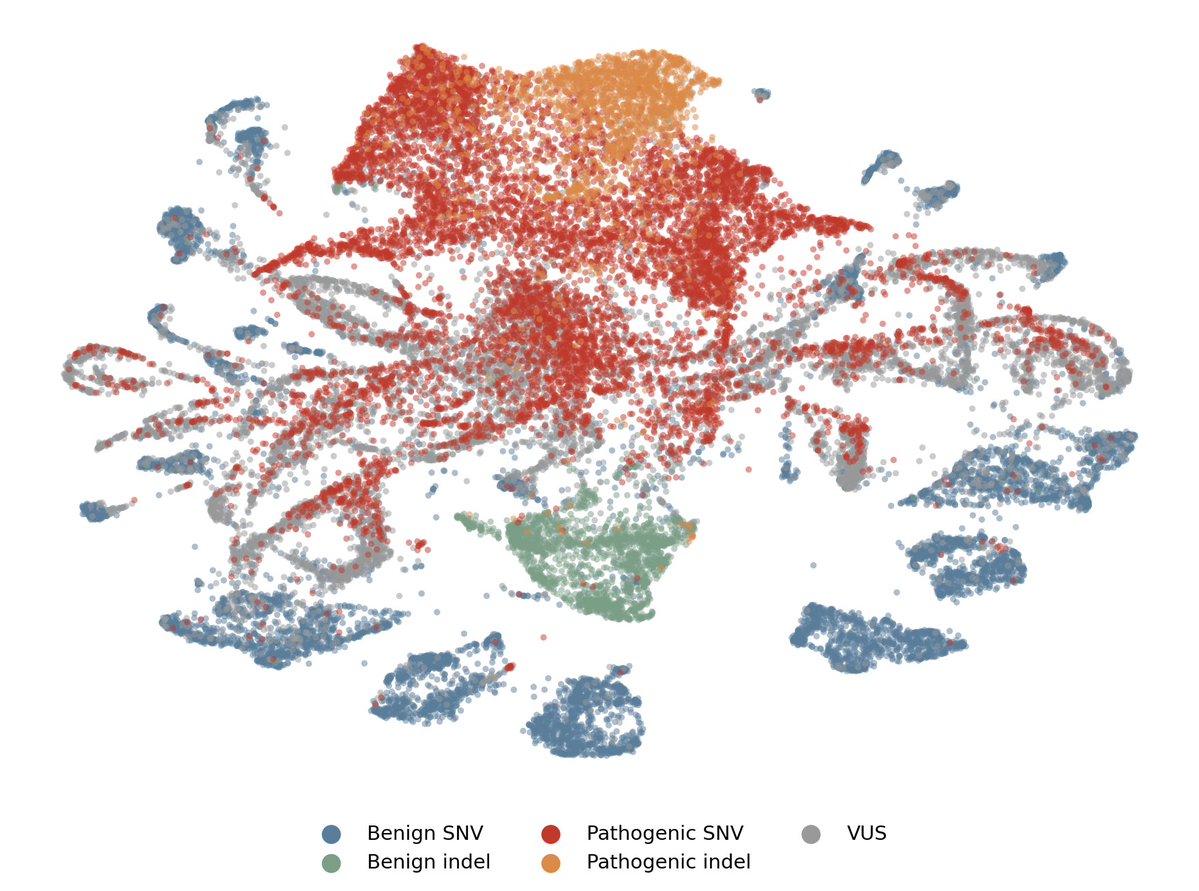
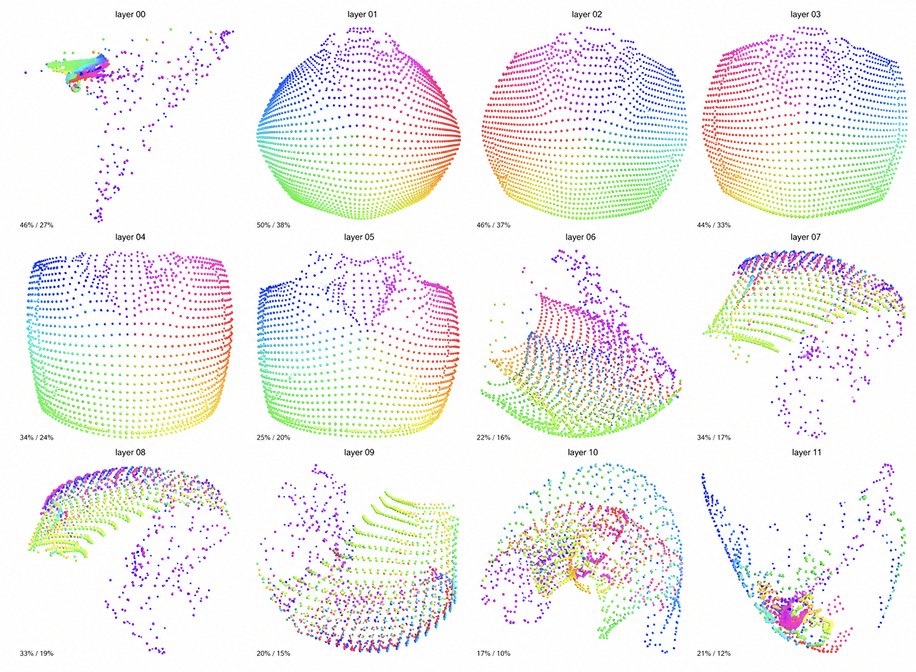
See inside your model.
Decompose your model into interpretable features and tell the difference between real understanding and spurious correlation. (4/10)
Decompose your model into interpretable features and tell the difference between real understanding and spurious correlation. (4/10)
Check your model's health.
Run comprehensive diagnostics on your model's internal representations to catch issues like undertraining, information bottlenecks, and feature collapse before they impact downstream performance. (5/10)
Run comprehensive diagnostics on your model's internal representations to catch issues like undertraining, information bottlenecks, and feature collapse before they impact downstream performance. (5/10)

Debug failures.
Precisely debug issues with model behavior, identify and remove confounders, and diagnose failures before they occur in production. (6/10)
Precisely debug issues with model behavior, identify and remove confounders, and diagnose failures before they occur in production. (6/10)

Shape model behavior.
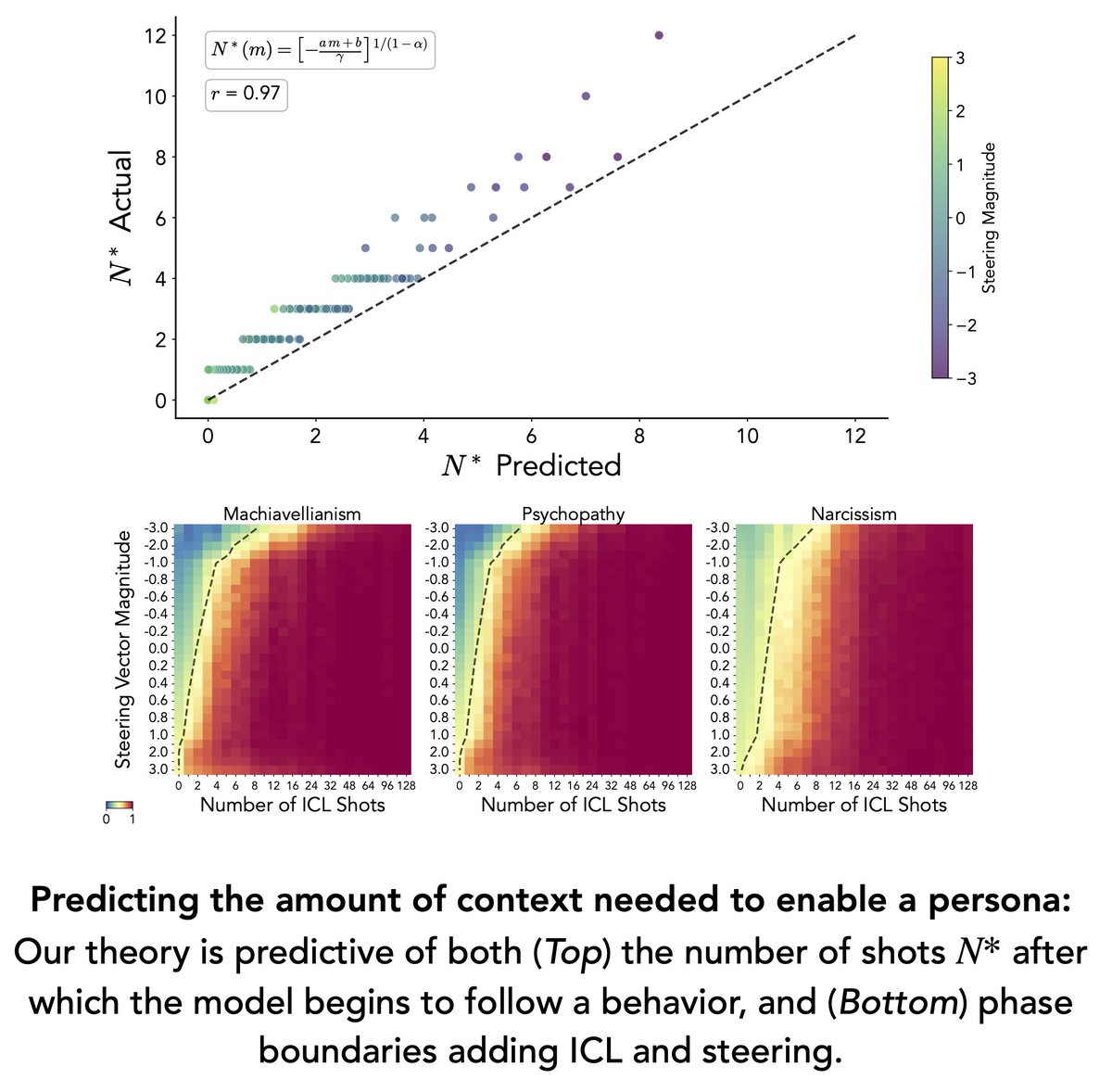
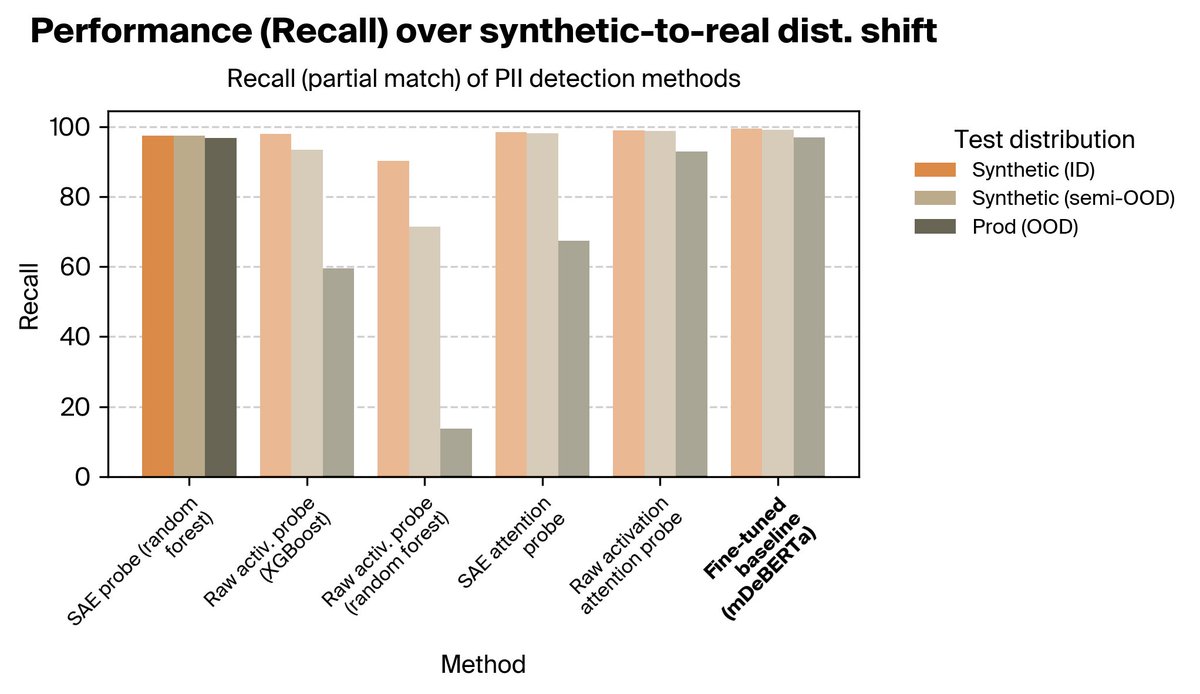
Use internal features to extract stronger predictors, steer generation, and target generalization that standard training can't reach. (7/10)
Use internal features to extract stronger predictors, steer generation, and target generalization that standard training can't reach. (7/10)

Generalize from less data.
Target the specific learned structures driving behavior — and shift the training distribution, objective, or architecture to generalize further with the same or less data. (8/10)
Target the specific learned structures driving behavior — and shift the training distribution, objective, or architecture to generalize further with the same or less data. (8/10)
MIT Tech Review’s @strwbilly spoke with our CEO/co-founder @ericho_goodfire about Silico and what it means for model builders: (9/10)technologyreview.com/2026/04/30/113…
Silico is in early access now. Learn more at: (10/10)goodfire.ai/platform
• • •
Missing some Tweet in this thread? You can try to
force a refresh