
Director of bioinformatics at AstraZeneca. YouTube at chatomics. On my way to helping 1 million people learn bioinformatics. Also talks about leadership.
How to get URL link on X (Twitter) App


 You ask for a plan from your CLI. It gets drafted in the cloud.
You ask for a plan from your CLI. It gets drafted in the cloud.
 2. Introduction to Data Science by the almighty Rafa!rafalab.dfci.harvard.edu/dsbook/
2. Introduction to Data Science by the almighty Rafa!rafalab.dfci.harvard.edu/dsbook/
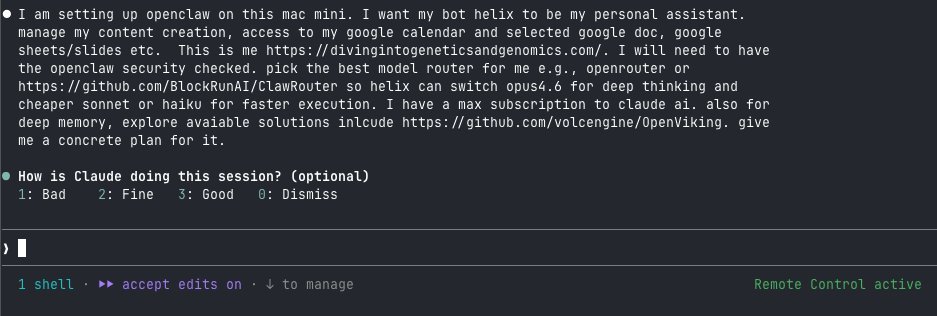
 The setup: OpenClaw as the agent framework, ClawRouter for model routing (Gemini Flash for simple tasks, Claude Sonnet for complex ones),
The setup: OpenClaw as the agent framework, ClawRouter for model routing (Gemini Flash for simple tasks, Claude Sonnet for complex ones),
 After Anthropic accidentally leaked 512K lines of Claude Code source through a bad npm package,
After Anthropic accidentally leaked 512K lines of Claude Code source through a bad npm package,
 A security researcher found a source map in the npm package pointing to the full TypeScript source.
A security researcher found a source map in the npm package pointing to the full TypeScript source.
 2/ The jellybean experiment.
2/ The jellybean experiment.
 2/ There are now hundreds of sequencing methods. Lior Pachter maintains a list:
2/ There are now hundreds of sequencing methods. Lior Pachter maintains a list:
 The thing I got wrong early: treating skills as markdown prompt files.
The thing I got wrong early: treating skills as markdown prompt files.