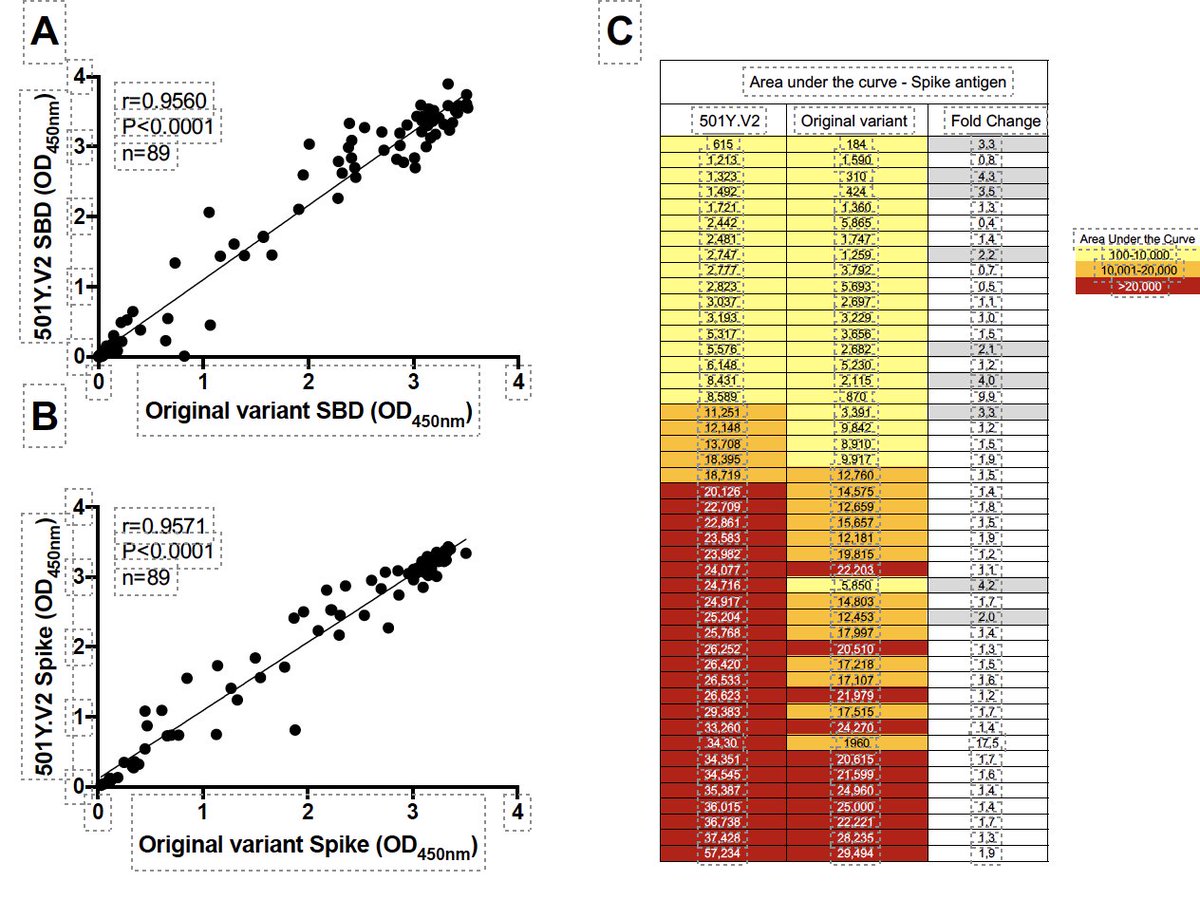
A truly global collaboration between groups in South Africa, U.K., Belgium, Sweeden, USA to understand convergent evolution of the variants of concern.
medrxiv.org/content/10.110…
medrxiv.org/content/10.110…
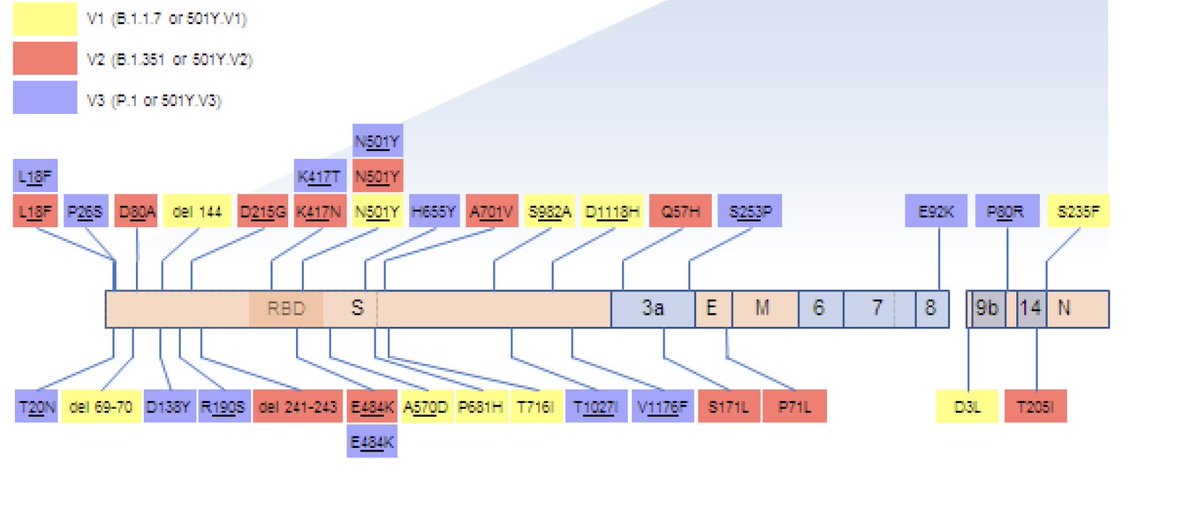
ARS-CoV-2 genome map indicating the locations and encoded amino acid changes of what we considered here to be signature mutations of 501Y.V1 (B.1.1.7), 501Y.V2 (B.1.251) and 501Y.V3 (p1) sequences. Amazing how different and similar the variants are... 
Signals of positive selection detected at 37 signature mutation sites in the VOCs between March 2020 and January 2021. In short, must of the sites in Spike are in convergent evolution, including 501, 484, 417, 18, etc. 

Figure 3 of this excellent paper, show the signature mutations and the directional selection amino acids. It seems that the variants are climbing the fitness peak of escape neutralisation and potentially transmission. 

Locations of amino acids encoded by codons that are evolving under positive selection and/or encode convergent amino acid changes between the lineages mapped to the 3D structure of Spike 

Fantastic work by @krisp_news @robertson_lab @LemeyLab @sergeilkp @UCTIDM @H3ABioNet @CovidGenomicsUK ! A must read pre-print
• • •
Missing some Tweet in this thread? You can try to
force a refresh