Finally out! Emergence of a SARS-CoV-2 variant of concern with mutations in spike glycoprotein disq.us/t/3vewz4s
So many people to thank, thank you and thank you! @rjlessells @houzhou @Mittenavoig @MRCza @dsigovza
Great to see South African science advancing fast!
So many people to thank, thank you and thank you! @rjlessells @houzhou @Mittenavoig @MRCza @dsigovza
Great to see South African science advancing fast!
The paper starts by showing how the second wave in South Africa arise so quickly, which was completely unexpected as we were in the start of our summer. 

It show how a new and unusual cluster emerged among the dozens of different lineages already circulating in South Africa. 

This new variant, the 501Y.V2 (or B.1.351) went to dominate the great majority of infections in the country. 
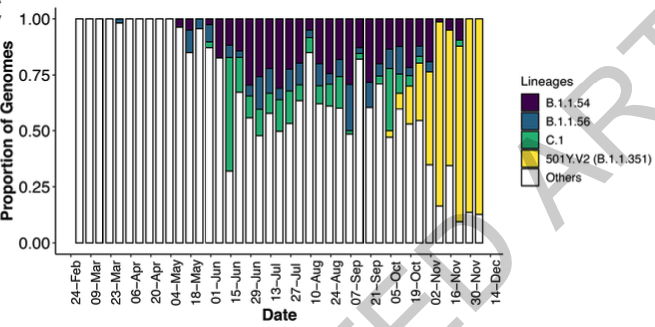
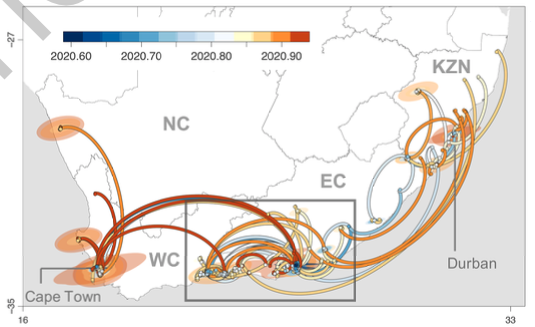
It also spread very fast across the coast in South Africa, moving very fast to south west (in direction of Cape Town) and north east (passing through Durban in the direction of Mozambique ). 
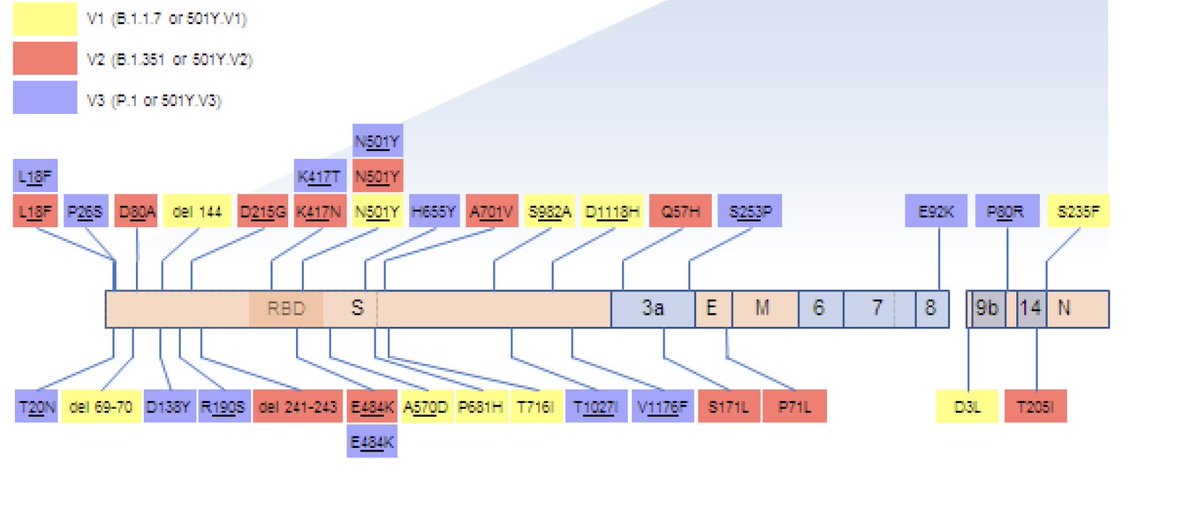
The 501Y.V2 variant has 23 mutations, which around 20 are amino acid changing mutation and 9 in the Spike region. 

As one map the mutations on the 3-D structure of the virus, it becomes clear that they are clustered in regions that are associated with ACE2 receptor binding (RBD) and antibody escape (RBD and NTD) 

A fantastic team effort of 5 Universities (UCT, Stellenbosch, Wits, UFS and KRISP/UKZN) and the large government labs (NHLS and NICD) in South Africa 

And of course to our national funders that believed that it is possible: Thanks @SAMRC, @dsigovza @tiaorgza !
Now we are expanding the network with national and international funders to further advance genomics surveillance in SA & help other African countries...
Now we are expanding the network with national and international funders to further advance genomics surveillance in SA & help other African countries...
• • •
Missing some Tweet in this thread? You can try to
force a refresh