
Happy to announce that our (@zhao7900, @YinyingWang) #deeplearning method INSCT for integration of #scrnaseq is out @NatMachIntell go.nature.com/2Uq73If! We extended triplet neural nets by defining 'batch-aware' triplet sampling, which overcomes batch effects in #scrnaseq data.
Instead of sampling triplets the traditional way, we added constraint to sample anchor-positives from different & anchor-negatives from the same batch. (We expect this approach to be also useful outside of #scrnaseq data - happy to discuss w anyone interested).
The semi-supervised mode allows robust transfer of cell labels across batches. Use case example: transfer labels from reference to new data set. 
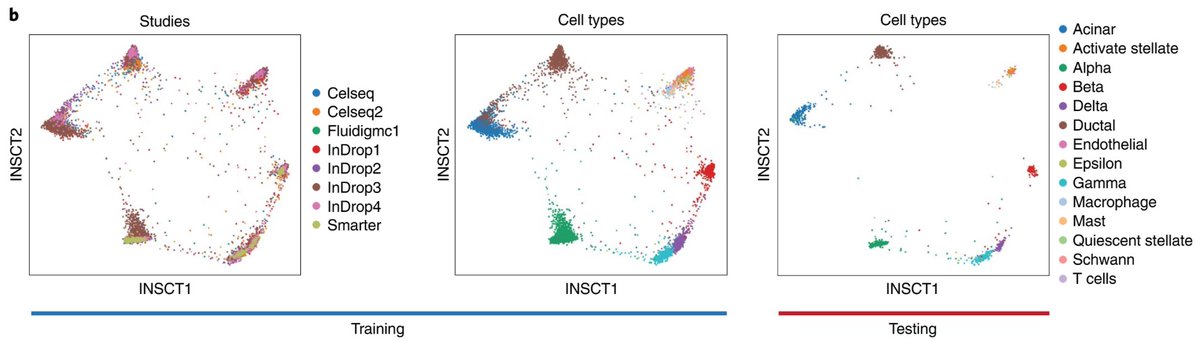
We demonstrated computational efficiency of INSCT by integrating millions of data points in <2hrs in standard web browser via @GoogleColab. Code freely avail: github.com/lkmklsmn/insct. Pls feel free to reach out!
• • •
Missing some Tweet in this thread? You can try to
force a refresh



