With machine learning and network science, we can start to recover the source code of the global virome. We wrote an instruction manual - out now in @NatureMicrobiol - and along the way, we've tried to solve what it would mean to really "predict and prevent the next pandemic" 🧵
https://twitter.com/viralemergence/status/1463538947078995969
Practically every big question about viral ecology, evolution, and emergence - from "why do bats have so many deadly viruses?" to "can we spot a pandemic flu before the first human case?" - is a variation on a fundamental scientific challenge: predicting the host-virus network. 

Over three years of research, we've compiled these kinds of studies into a unified framework, allowing us to put our finger on a new convergence science - "the science of the host-virus network" - that uses computational inference to understand viral biology across scales. 



Every year, this set of tools has become more actionable for protecting human and wildlife health. We can use these tools to spot "Disease X" before the first human case, to sample wildlife reservoirs for new viruses more efficiently, and even to predict climate change impacts.
So why have decades of efforts to "predict and prevent the next pandemic" fallen flat? Two answers: (1) We keep missing our chances to actually apply these tools for actionable prediction. (2) Prevention actually falls on health systems, not scientists. 
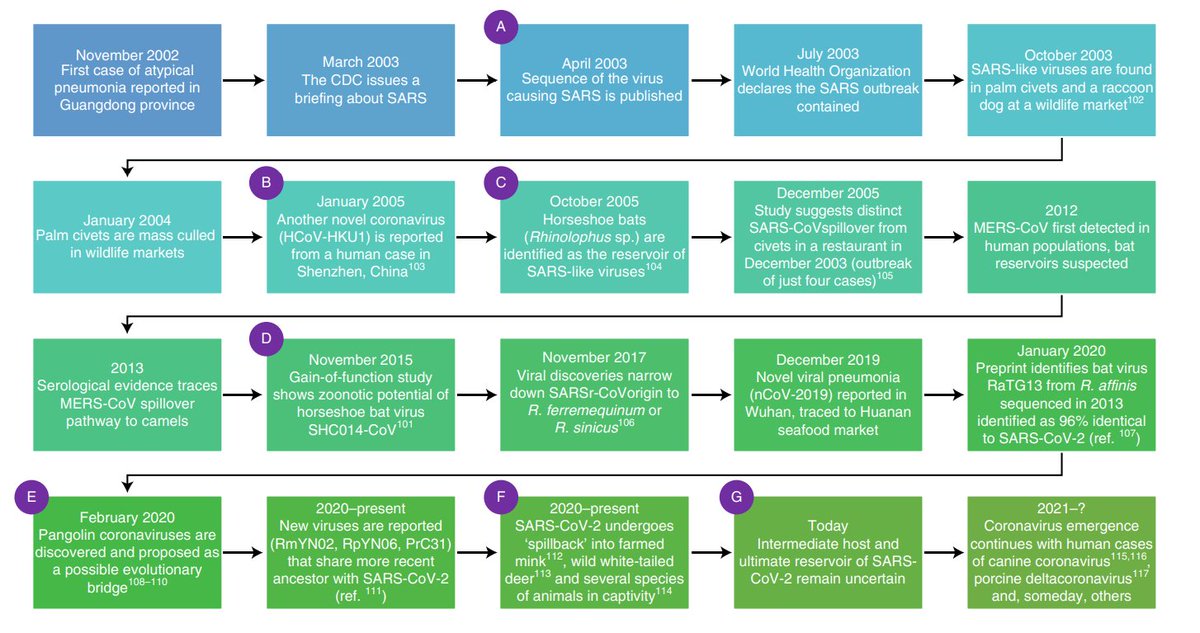
But we're at the tipping point right now where we can start to use these tools to do a lot of good. That's why @viralemergence exists in the first place - we want to build and share new tools, and the datasets we need to power them, to close this gap and save lives.
If that sounds good, you can see more of what we do at viralemergence.org, and we'll be posting about different aspects of our study and the work we do on the host-virus network all day 🎉
• • •
Missing some Tweet in this thread? You can try to
force a refresh













