Do you know Daisy Roulland-Dussoix? She is one of the discoverers of restriction enzymes, who’s findings paved the way for the development of recombinant DNA and cloning technologies. Accordingly, the finding was rewarded with a #NobelPrize. But the prize didn’t go to her... 🧵👇 

Daisy Roulland-Dussoix worked with Werner Arber to study the mechanism for the observed host-specificity of λ Phages. It was known from an important 1953 paper (Bertani & Weigle) that phages, that had replicated in a certain E. coli strain, could only re-infect the same strain. 

Roulland-Dussoix & Arber showed that host-specificity is linked with the phage’s DNA. Using phages carrying radiolabeled DNA, they showed that progeny with 2 parental DNA strands retained specificity, while progeny with newly synthesized daughter strands could adapt to new hosts. 



Back-to-back with this first paper, Roulland-Dussoix & Arber then add that the phage DNA is degraded immediately after injection into the E. coli cell,& that the DNA can be ‘rescued’ by the infection of additional DNA, suggesting competition between the DNA for recognition sites. 

It therefore seemed that enzymes, that competitively bind the DNA are involved in this process. These host enzymes are degrading the invasive phage DNA, and are thus ‘restricting’ the phage’s ability to infect the host cell: restriction enzymes.
In 1968 Meselson & Yuan identified the first restriction enzyme, the type I endonuclease EcoK from E. coli, followed by Smith & Wilcox, who identified the type II enzyme HindII in 1970, & Mertz & Davis who found EcoRI in 1972,& also showed that this enzyme produces ‘sticky-ends’. 

The lab of Daniel Nathans then proceeded to use these restriction enzymes to cut up the simian virus 40 genome into 11 defined pieces, followed by gel-separation, demonstrating that endonuclease analysis could be used for DNA mapping of genomes. 

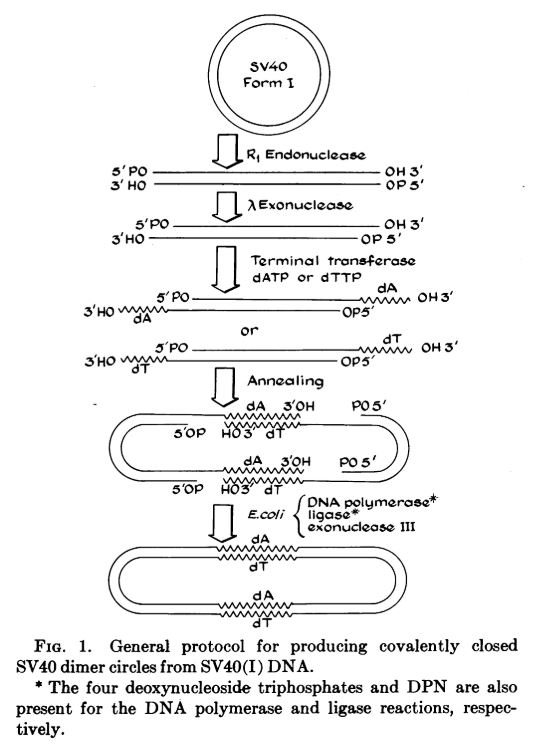
Then, in 1972/1973, restriction enzymes were used to splice gene fragments into plasmids and shuttled into bacterial cells, where they were replicated and expressed. Molecular cloning was developed,& with it started the Molecular Biology Revolution, changing the world forever. 


Accordingly, Nobel Prizes were in order! And the 1978 Prize in Physiology and Medicine "for the discovery of restriction enzymes and their application to problems of molecular genetics" went to Werner Arber, Daniel Nathans & Hamilton O. Smith. Daisy Roulland-Dussoix was ignored. 

Thankfully, Daisy Roulland-Dussoix went on to have a long and successful career in science despite this, initially continuing work on restriction enzymes & cloning with Herbert Boyer & eventually becoming a leading expert on mycobacterium/mycoplasmas.
Daisy Roulland-Dussoix died January 5, 2014. I would have liked to end this with a nice obituary for her, but unfortunately I could not find one...
• • •
Missing some Tweet in this thread? You can try to
force a refresh

















