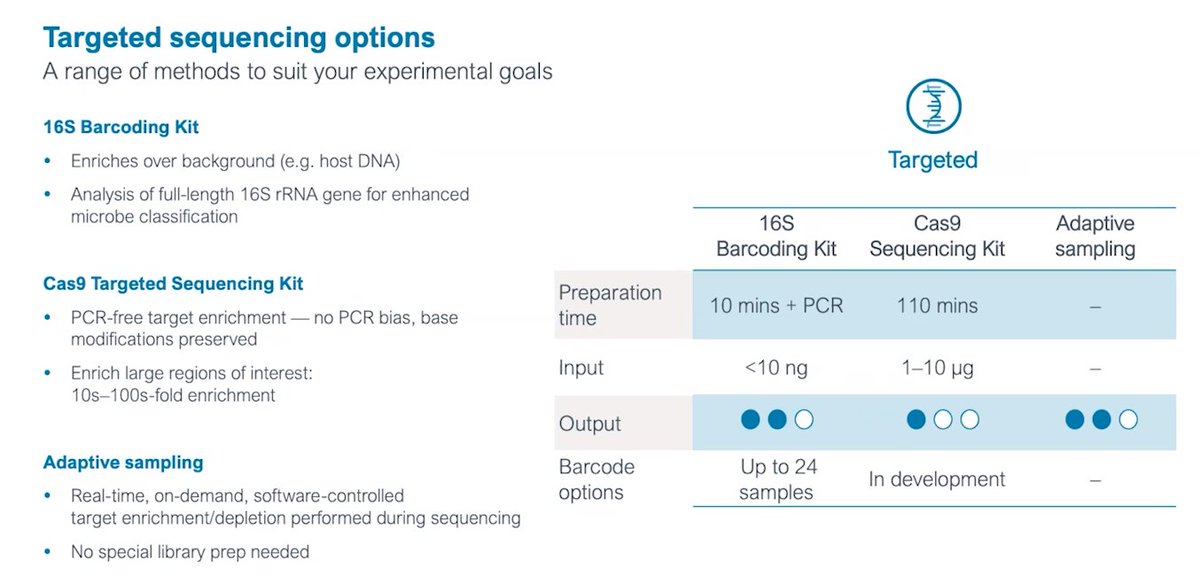
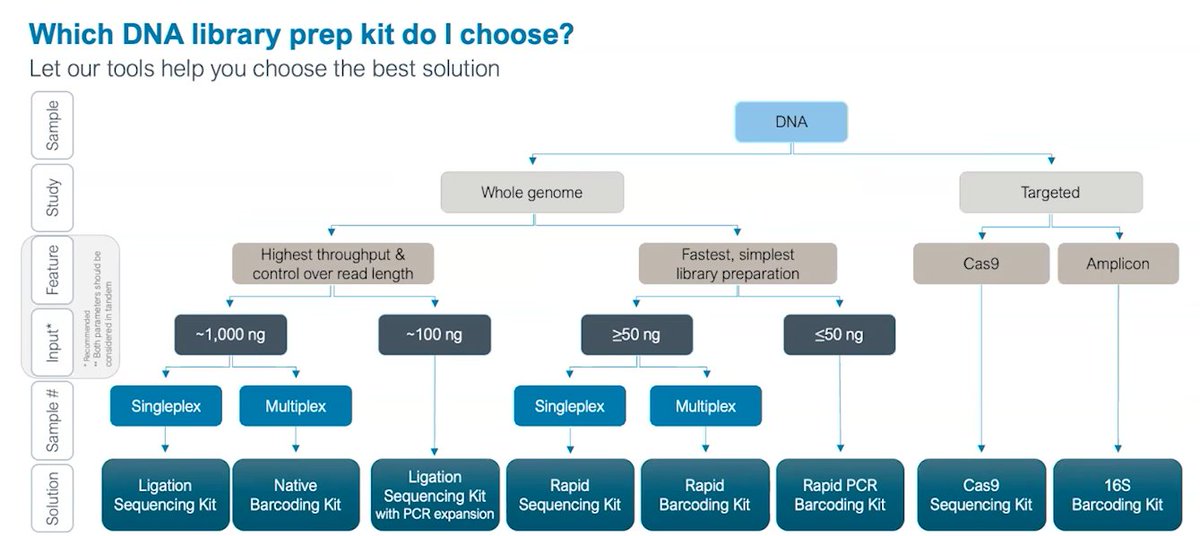
On library prep at #Nanoporeconf, a description for PCR-free methods showing the difference between ligation (max output) and rapid mode (10minutes, minimal lab equipment needed). Ultralong reads (ULR) also enabled, all Kit14. 

• • •
Missing some Tweet in this thread? You can try to
force a refresh

 Read on Twitter
Read on Twitter