Can AI rewrite our human genome? ⌨️🧬
Today, we announce the successful editing of DNA in human cells with gene editors fully designed with AI. Not only that, we've decided to freely release the molecules under the @ProfluentBio OpenCRISPR initiative.
Lots to unpack👇
Today, we announce the successful editing of DNA in human cells with gene editors fully designed with AI. Not only that, we've decided to freely release the molecules under the @ProfluentBio OpenCRISPR initiative.
Lots to unpack👇
AI has become increasingly pervasive in our daily lives from how we sift through information, produce content, and interact with the world. This marks a new chapter where AI is used to alter the fundamental blueprint of who we are - our DNA.
We were immediately drawn to gene editing due to the pressing societal needs, potential for one-and-done cures to disease, and the scientific challenge + complex biology involving protein, RNA, and DNA.
Our LLMs were trained on massive scale sequence and biological context to generate millions of diverse CRISPR-like proteins that do not occur in nature, thereby exponentially expanding virtually all known CRISPR families at-will. 
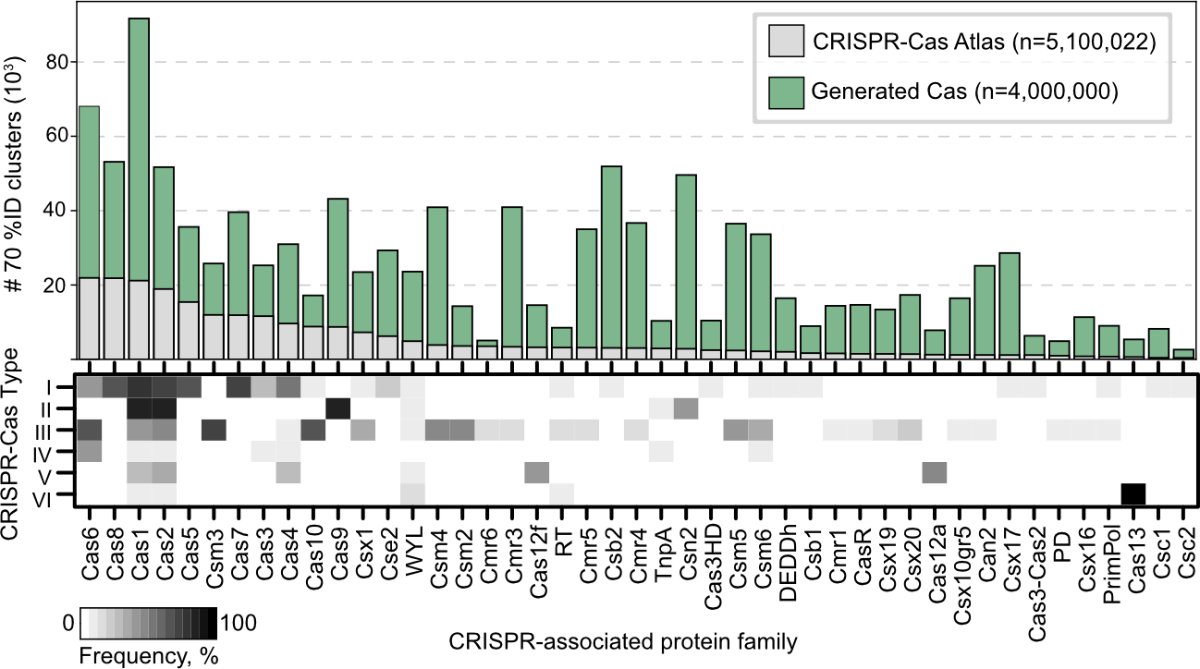
We then focus on type II effector complexes, generating cas9-like proteins and gRNAs. These proteins are hundreds of mutations away from anything in nature.


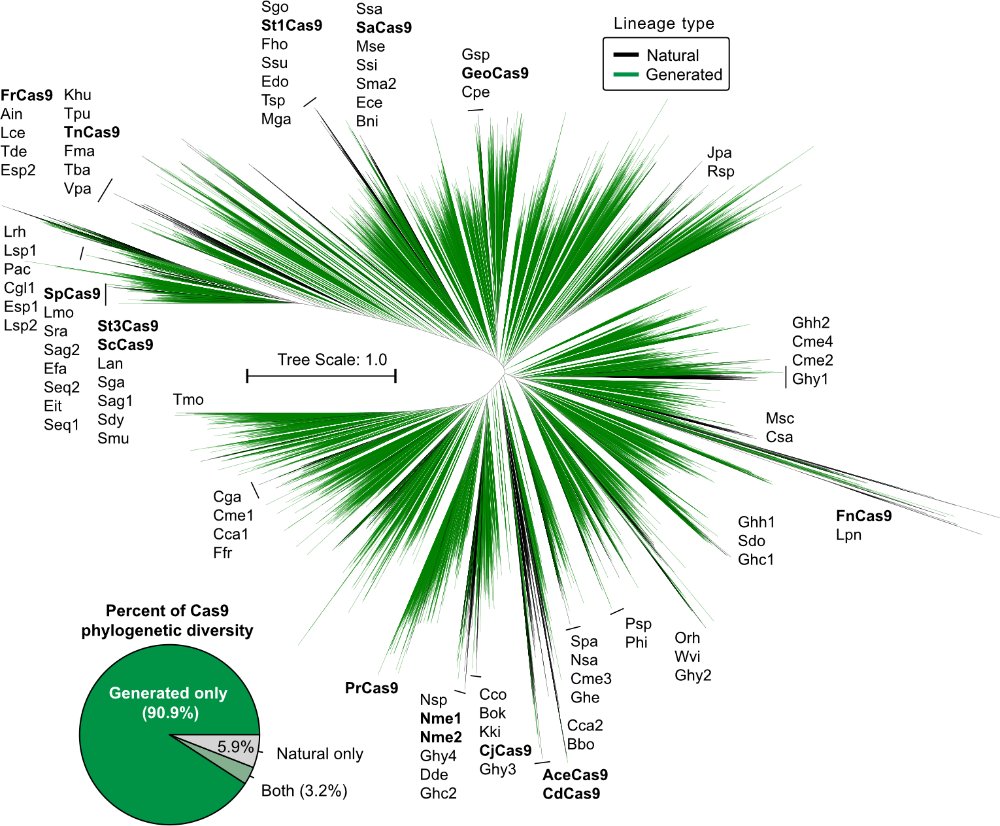
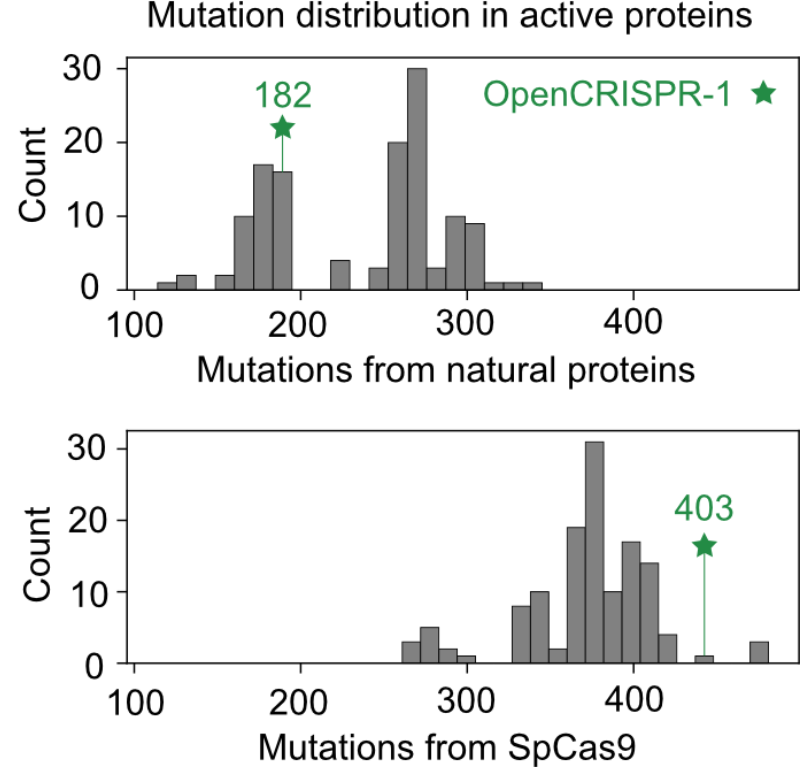
We then characterized our generations in the wet lab and found that the AI-designed gene editors show comparable or improved activity and specificity relative to SpCas9, the prototypical gene editing effector. More characterization is underway but we're already impressed.


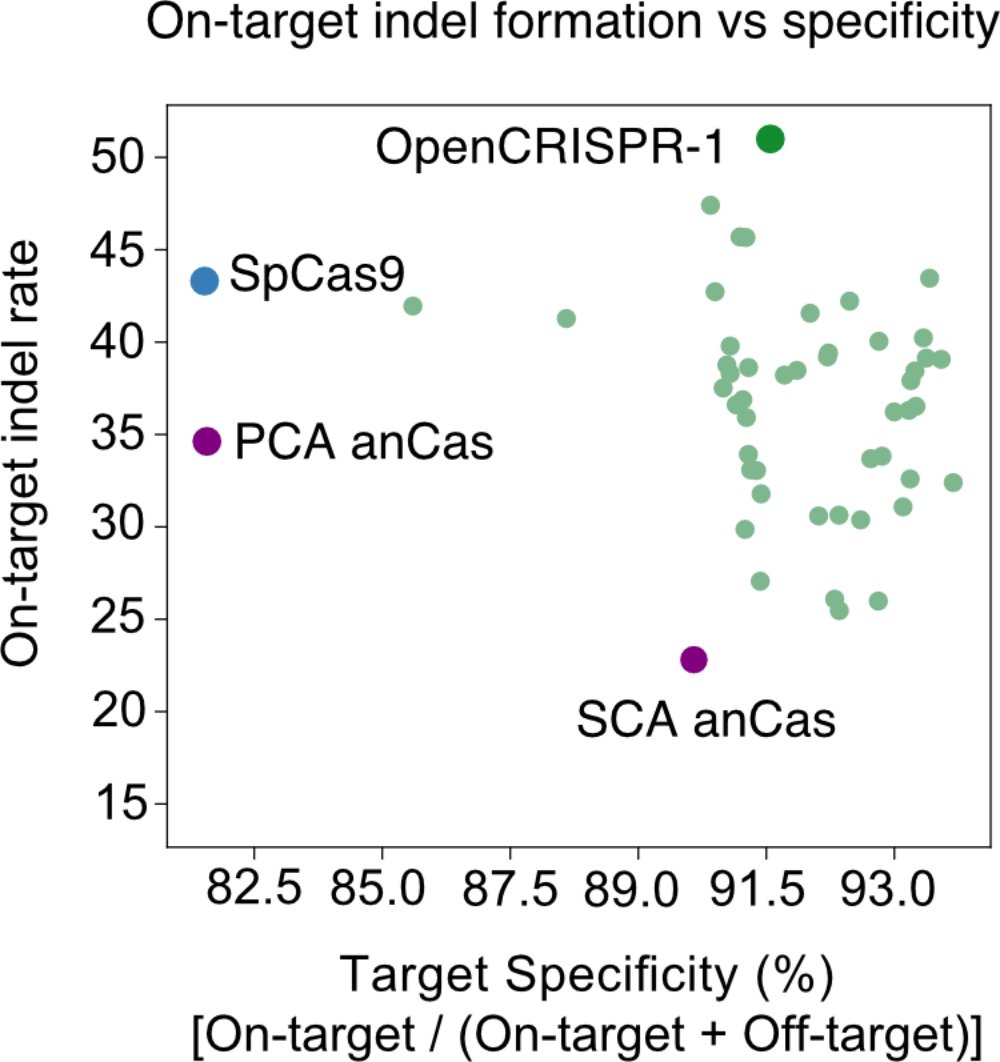
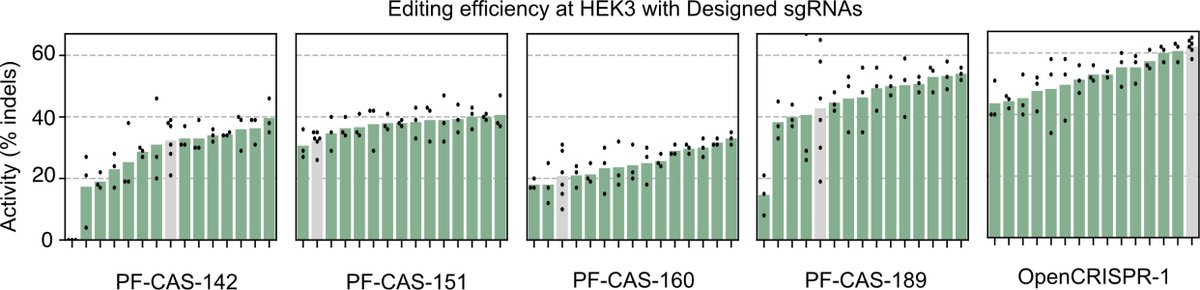
We also created an AI-designed base editor which exhibited really exciting performance in precise A->G edits. 
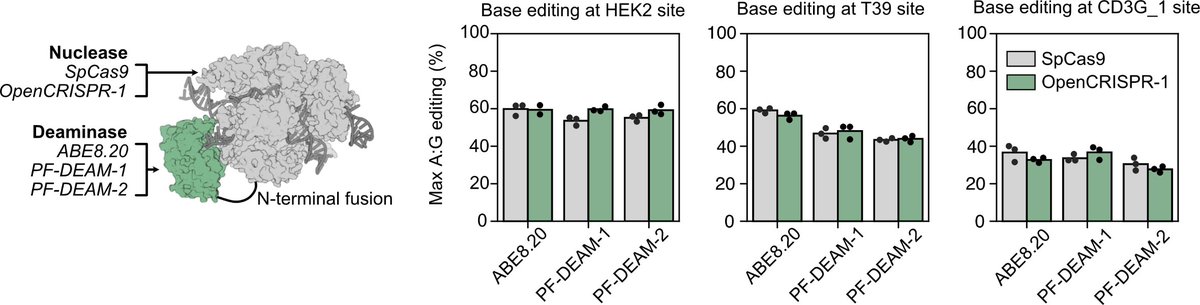
The results point to a future where AI precisely designs what is needed to create a range of bespoke cures for disease. There is still much to build to achieve this vision. To spur innovation and democratization, we are freely releasing OpenCRISPR-1. Try it out! 

This was truly a team effort across all disciplines of the company. @jeffruffolo SNayfach JGallagher @AadyotB JBeazer RHussain JRuss JYip EHill @MartinPacesa @alexjmeeske PCameron and the broader Profluent team. If you want to build with us, join. We’re hiring.
Paper:
Blog:
NYTimes:
Press Release:
Access OpenCRISPR-1: biorxiv.org/content/10.110…
profluent.bio/blog/editing-t…
nytimes.com/2024/04/22/tec…
businesswire.com/news/home/2024…
forms.gle/zzPFYuesZXZGT5…
Blog:
NYTimes:
Press Release:
Access OpenCRISPR-1: biorxiv.org/content/10.110…
profluent.bio/blog/editing-t…
nytimes.com/2024/04/22/tec…
businesswire.com/news/home/2024…
forms.gle/zzPFYuesZXZGT5…
• • •
Missing some Tweet in this thread? You can try to
force a refresh







