
director of applied AI @TempusAI; prev: faculty @DanaFarber, group leader @Harvard & phd @eth
How to get URL link on X (Twitter) App


 Driven by @samwang36 & @RunziTan97745 & with @BoWang87, we benchmarked DeepSeek-R1-0528 on zero-shot scRNAseq cell type annotation against non-reasoning LLMs, classifiers & foundation models . What we found surprised us👇biorxiv.org/content/10.110…
Driven by @samwang36 & @RunziTan97745 & with @BoWang87, we benchmarked DeepSeek-R1-0528 on zero-shot scRNAseq cell type annotation against non-reasoning LLMs, classifiers & foundation models . What we found surprised us👇biorxiv.org/content/10.110…

 First, why is this model so important?
First, why is this model so important?https://twitter.com/simocristea/status/1879923278770241838


 1. New Cancer Cases and Deaths in 2025:
1. New Cancer Cases and Deaths in 2025:
 Long thread ahead, going deep into the molecular workings of breast cancer immunotherapy.
Long thread ahead, going deep into the molecular workings of breast cancer immunotherapy.
 1. Assembly theory (Sharma et al., Nature)
1. Assembly theory (Sharma et al., Nature)

 First things first:
First things first:

 1. Placenta 1/5
1. Placenta 1/5

 In this thread, we'll discuss:
In this thread, we'll discuss:
 It’s 1938, and Public Health Services are advising people that detecting and treating cancers early will save their lives.
It’s 1938, and Public Health Services are advising people that detecting and treating cancers early will save their lives. 
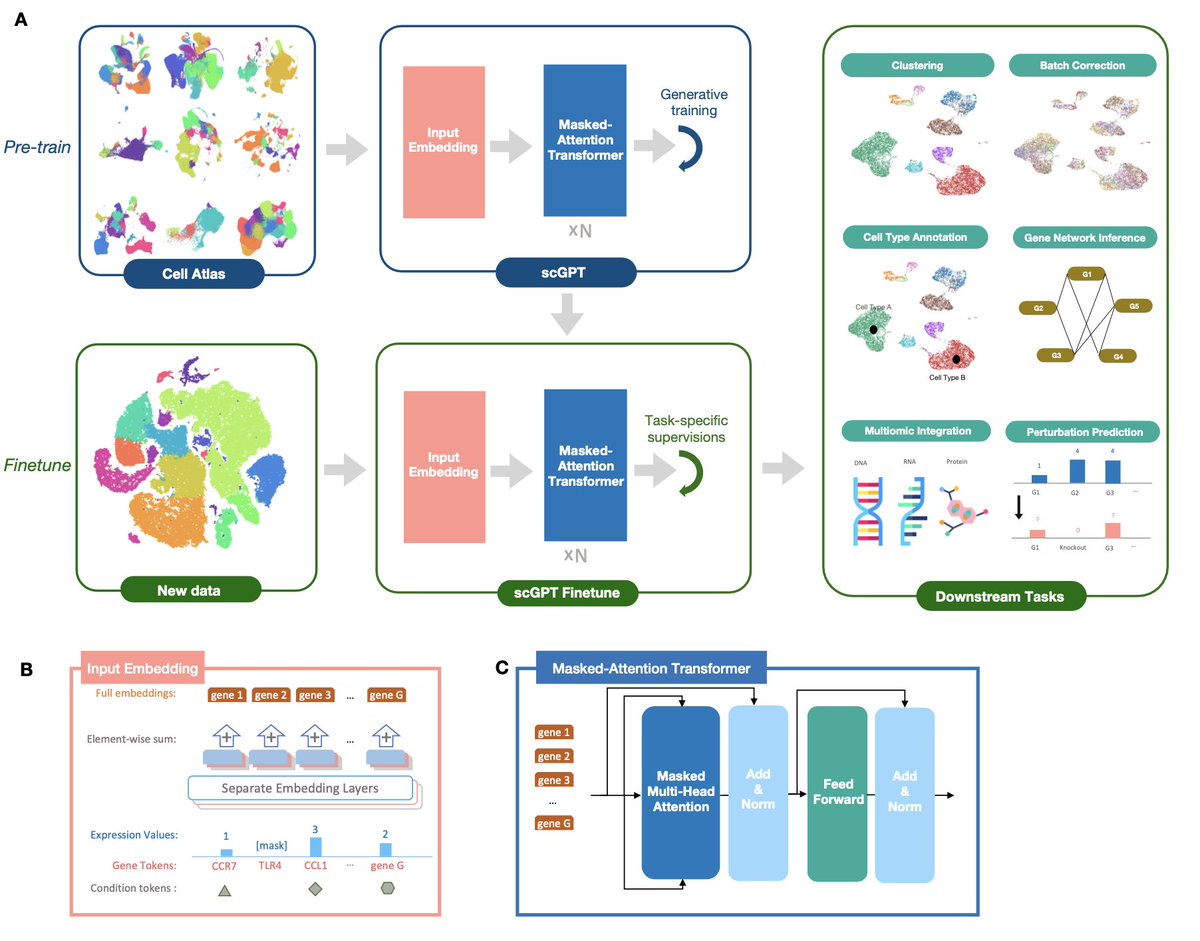
 The self-attention transformer (chatGPT) is so successful in natural language processing (NLP)
The self-attention transformer (chatGPT) is so successful in natural language processing (NLP)
 I'll start with a quick summary of the paper, such that we're all on the same page.
I'll start with a quick summary of the paper, such that we're all on the same page.
 This thread is organized as follows:
This thread is organized as follows:
 This thread is organized as follows:
This thread is organized as follows:
 First, some context.
First, some context.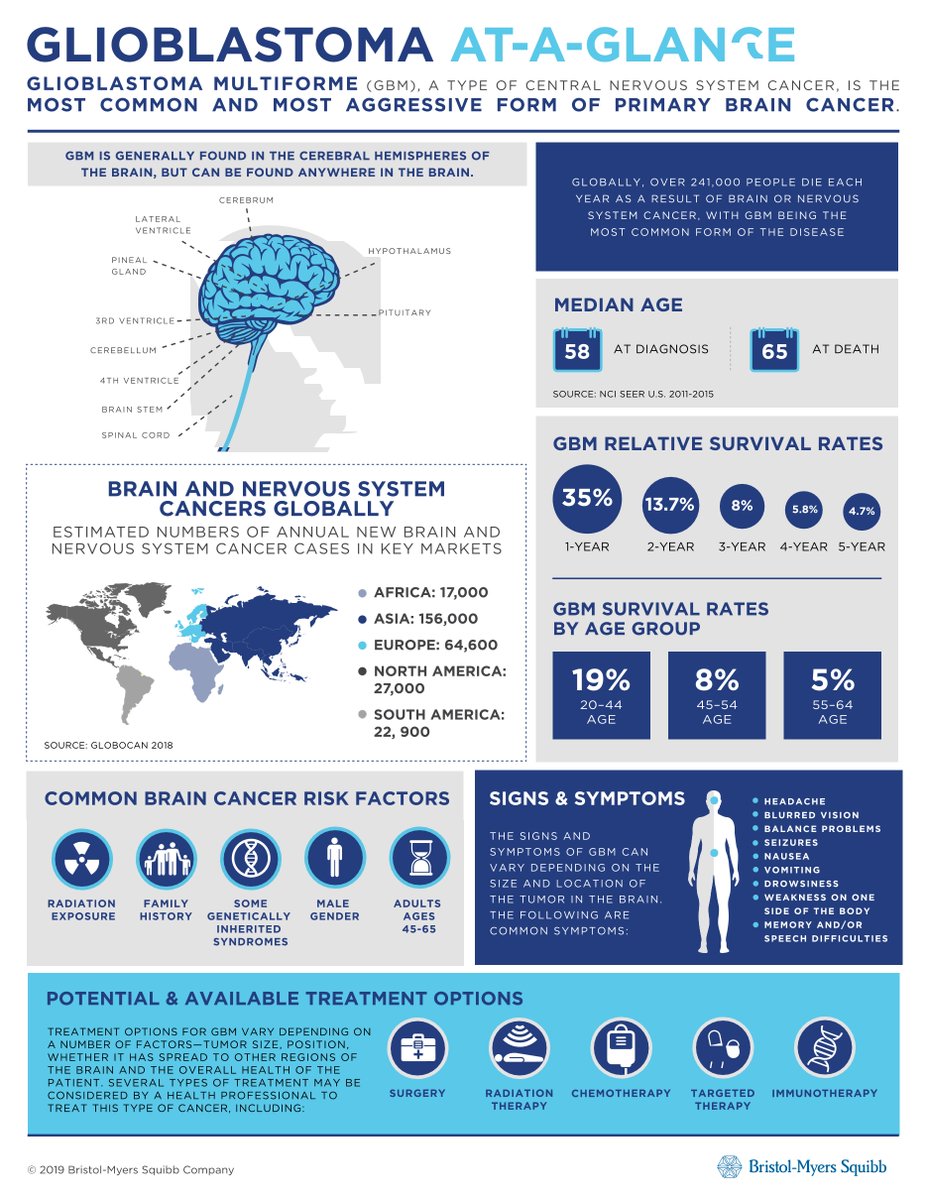

 Last year, together with @LabPolyak @harvardmed, we published a study in which we did something totally awesome: we experimentally showed how a TGFBR1 inhibitor drug 💊 prevents breast tumor initiation in two different rat models!
Last year, together with @LabPolyak @harvardmed, we published a study in which we did something totally awesome: we experimentally showed how a TGFBR1 inhibitor drug 💊 prevents breast tumor initiation in two different rat models!https://twitter.com/simocristea/status/1600578512578035733

 First, some background.
First, some background.

 This method comes from the @MetaAI FAIR protein folks: @BrianHie, @salcandido, @ebetica, @OriKabeli, @proteinrosh, @nikismetanin, @TomSercu, @alexrives and is available as a preprint.
This method comes from the @MetaAI FAIR protein folks: @BrianHie, @salcandido, @ebetica, @OriKabeli, @proteinrosh, @nikismetanin, @TomSercu, @alexrives and is available as a preprint.
 The very nice paper discussed in this thread comes from a team led by @nikhil_ai at Salesforce @SFResearch 👏
The very nice paper discussed in this thread comes from a team led by @nikhil_ai at Salesforce @SFResearch 👏
 This summary is based on @PetarV_93’s recent paper with introductory theoretical notions on Graph Neural Networks.
This summary is based on @PetarV_93’s recent paper with introductory theoretical notions on Graph Neural Networks.