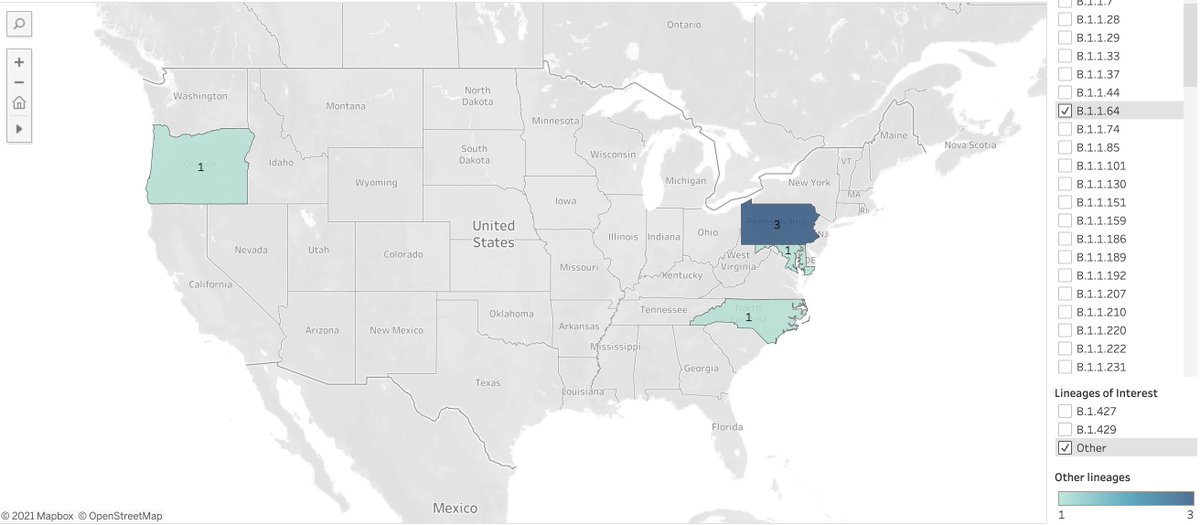
Data drop from @my_helix SARS-CoV-2 surveillance
Check any state public.tableau.com/profile/helix6…
Or download from github.com/myhelix/helix-…
To see if large vaccination effort can keep overall cases from📈again -> important to monitor FL
See @ScottGottliebMD for perspective
Check any state public.tableau.com/profile/helix6…
Or download from github.com/myhelix/helix-…
To see if large vaccination effort can keep overall cases from📈again -> important to monitor FL
See @ScottGottliebMD for perspective

2/
With overall number of cases 📉 quickly => important to look at % of positives that are B1.1.7
AND also the evolution of the absolute number of B1.1.7
Can do this based on Helix numbers,
or by multiplying % from Helix by overall number of cases reported by states and CDC
With overall number of cases 📉 quickly => important to look at % of positives that are B1.1.7
AND also the evolution of the absolute number of B1.1.7
Can do this based on Helix numbers,
or by multiplying % from Helix by overall number of cases reported by states and CDC
3/
CA: % of positives that are B117 now ~15-17%
Increase in absolute numbers of B117 is slower (compared to FL).
Still N of B117 is not decreasing, unlike the non-SGTF SARS-CoV-2 variants including B.1.429 & B.1.427 who are decreasing fast.
Note: Y-axis truncated in right 👇
CA: % of positives that are B117 now ~15-17%
Increase in absolute numbers of B117 is slower (compared to FL).
Still N of B117 is not decreasing, unlike the non-SGTF SARS-CoV-2 variants including B.1.429 & B.1.427 who are decreasing fast.
Note: Y-axis truncated in right 👇

4/
If you are curious about B.1.427 and B.1.429, the 2 variants of interest (NOT variants of concern), aka the 'CA' variant:
% of random (non-SGTF) sequences that are B.1.427 or B.1.429 has been relatively stable (60-70%) since mid-Jan in CA.
& absolute numbers 📉
If you are curious about B.1.427 and B.1.429, the 2 variants of interest (NOT variants of concern), aka the 'CA' variant:
% of random (non-SGTF) sequences that are B.1.427 or B.1.429 has been relatively stable (60-70%) since mid-Jan in CA.
& absolute numbers 📉

5/
A few other states now have more than 10% of the positives that are B.1.1.7*
* based on combination of 1) % of positives that are SGTF (S-gene target failure) AND 2) % of SGTF sequenced that are B.1.1.7
Georgia is one example. Now ~20% of positives are B117.
A few other states now have more than 10% of the positives that are B.1.1.7*
* based on combination of 1) % of positives that are SGTF (S-gene target failure) AND 2) % of SGTF sequenced that are B.1.1.7
Georgia is one example. Now ~20% of positives are B117.

6/
As always this is a big many teams effort incl.
@genesareclever @ETCirulli @kellyschiabor William Lee & many more from @my_helix
@illumina team
@dmaccannell and @CDCgov team
@K_G_Andersen @gkay92 and team
A lot more coming as we continue to surveille.
As always this is a big many teams effort incl.
@genesareclever @ETCirulli @kellyschiabor William Lee & many more from @my_helix
@illumina team
@dmaccannell and @CDCgov team
@K_G_Andersen @gkay92 and team
A lot more coming as we continue to surveille.
• • •
Missing some Tweet in this thread? You can try to
force a refresh