Machine learning has enabled scientific breakthroughs in several fields.
Biotechnology is one of the most fascinating, as researchers could perform mindblowing tasks with the new tools.
Here are my favorite problems that machine learning helps to solve!
🧵 👇🏽
Biotechnology is one of the most fascinating, as researchers could perform mindblowing tasks with the new tools.
Here are my favorite problems that machine learning helps to solve!
🧵 👇🏽
These are the topics we are going to talk about:
1. Predicting protein structure from amino acid sequences.
2. Accelerating high-throughput screening for drug discovery.
3. Mapping out the human cell atlas.
4. Precision medicine.
Let's dive in!
1. Predicting protein structure from amino acid sequences.
2. Accelerating high-throughput screening for drug discovery.
3. Mapping out the human cell atlas.
4. Precision medicine.
Let's dive in!
1. Predicting protein structure from amino acid sequences.
Proteins are the workhorses of biology. In our body, myriads of processes are controlled by proteins. They enable life. Yet compared to their importance, we know so little about them!
Proteins are the workhorses of biology. In our body, myriads of processes are controlled by proteins. They enable life. Yet compared to their importance, we know so little about them!
We know that they are composed of amino acids, but predicting their structure from the amino acid chain remained unsolved for a very long time.
After decades of work, a giant leap was made by AlphaFold that revolutionized the field.
deepmind.com/blog/article/a…
After decades of work, a giant leap was made by AlphaFold that revolutionized the field.
deepmind.com/blog/article/a…
AlphaFold was created by Google, and compared to its predecessors, its increase in predictive performance represented a lightyear jump.
2. Accelerating high-throughput screening for drug discovery.
Even after decades of research, sometimes the best way to discover drugs is to try out millions of candidate molecules. These experiments produce an insane amount of data. Machine learning helps make sense of it!
Even after decades of research, sometimes the best way to discover drugs is to try out millions of candidate molecules. These experiments produce an insane amount of data. Machine learning helps make sense of it!
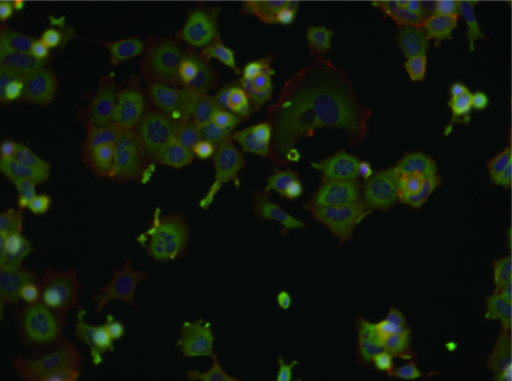
One common method is to treat cell cultures and use microscopy to detect and analyze every single cell, measuring the effect of the candidate molecules.
However, tens of thousands of images (like below) are not simple to analyze.
(Image source: bbbc.broadinstitute.org/BBBC021)
However, tens of thousands of images (like below) are not simple to analyze.
(Image source: bbbc.broadinstitute.org/BBBC021)
Deep learning enabled tools are much better at this task than their predecessors.
With these, we can perform precise analysis on every single cell at scale, accelerating the drug discovery process.
With these, we can perform precise analysis on every single cell at scale, accelerating the drug discovery process.
3. Mapping out the human cell atlas.
The Human Cell Atlas aims "To create comprehensive reference maps of all human cells—the fundamental units of life—as a basis for both understanding human health and diagnosing, monitoring, and treating disease."
humancellatlas.org
The Human Cell Atlas aims "To create comprehensive reference maps of all human cells—the fundamental units of life—as a basis for both understanding human health and diagnosing, monitoring, and treating disease."
humancellatlas.org
Cell types are characterized by their biochemical fingerprints. However, there are millions of variables and nonlinear relations. Both supervised and unsupervised machine learning methods can help with this task.
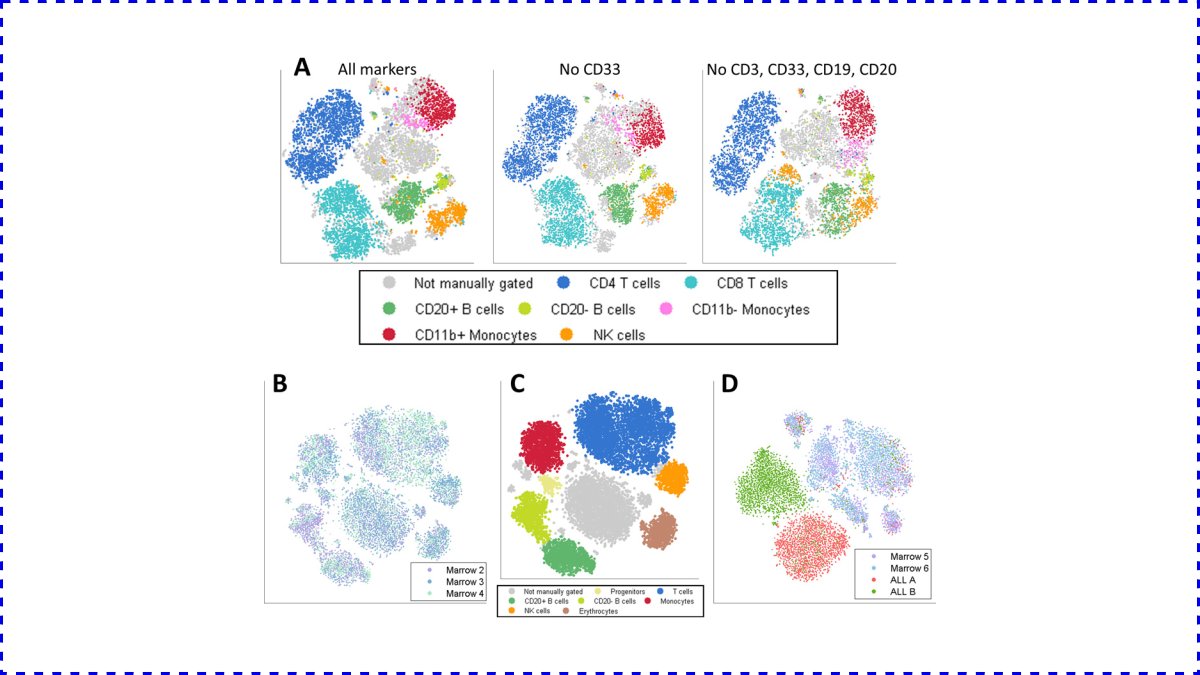
For example, dimensionality reduction tools such as PCA, t-SNE, UMAP, or even learned neural network embeddings are extremely useful when combined with high-throughput wet lab tools and expert insight.
(Image source: pubmed.ncbi.nlm.nih.gov/23685480/)
(Image source: pubmed.ncbi.nlm.nih.gov/23685480/)
4. Precision medicine.
Every one of us is different on a genetic level. Because certain conditions, like cancer, are so specific, individual therapies are the best.
However, as mentioned above, the human body is incredibly complex.
Every one of us is different on a genetic level. Because certain conditions, like cancer, are so specific, individual therapies are the best.
However, as mentioned above, the human body is incredibly complex.
Machine learning helps on several fronts:
• diagnostics,
• designing personalized therapies,
• interpreting massive genomic datasets,
• integrating multimodal measurements,
and many more.
• diagnostics,
• designing personalized therapies,
• interpreting massive genomic datasets,
• integrating multimodal measurements,
and many more.
So, the recent discoveries in life sciences finally enabled researchers to gather more and better data.
Some breakthroughs are already happening, some are on their way.
Nowadays, it is extremely exciting to work at the intersection of biology and machine learning.
Some breakthroughs are already happening, some are on their way.
Nowadays, it is extremely exciting to work at the intersection of biology and machine learning.
If you enjoyed this thread, consider following me and hitting a like/retweet on the first tweet of the post!
I regularly post simple explanations of seemingly complicated concepts in machine learning, make sure you don't miss out on the next one!
I regularly post simple explanations of seemingly complicated concepts in machine learning, make sure you don't miss out on the next one!
https://twitter.com/TivadarDanka/status/1386671410899206148
• • •
Missing some Tweet in this thread? You can try to
force a refresh












