BREAKING🔔 The 14th preprint from G2P-Japan🇯🇵 is out @biorxivpreprint. We illuminated the evolutionary rule underlying the convergent evolution of #Omicron and the virological characteristics of one of the latest lineages of concern, BQ.1.1. Please RT. 1/
biorxiv.org/content/10.110…
biorxiv.org/content/10.110…
In late 2022, the Omicron subvariants have highly diversified. However, some lineages have convergently acquired amino acid substitutions at five critical residues in the spike protein: R346, K444, L452, N460 and F486. 2/
https://twitter.com/dfocosi/status/1575836076089823232?s=20&t=P4COtUsJ2_LztpucW6DS5w
Omicron BQ.1.1 (aka #Cerberus), a descendant of BA.5, bears all five substitutions at the convergent sites: #R346T #K444T #L452R #N460K #F486V. 3/ 
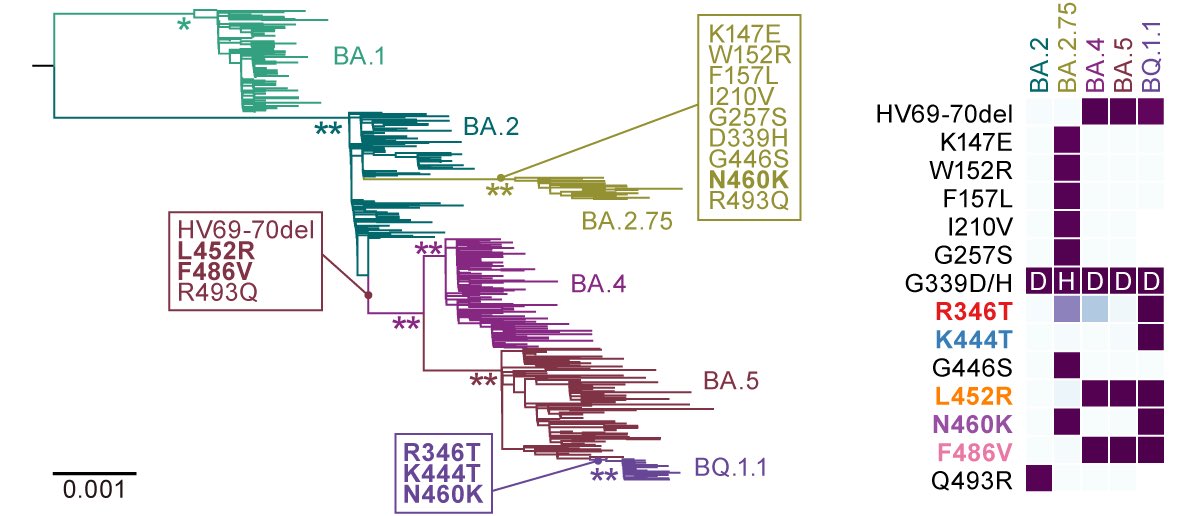
Here, we, particularly two shining young talents, @jampei2 & @SpyrosLytras, performed phylogenetic and epidemic dynamics analyses and showed that Omicron subvariants independently increased their viral fitness by acquiring the convergent substitutions. 4/ 

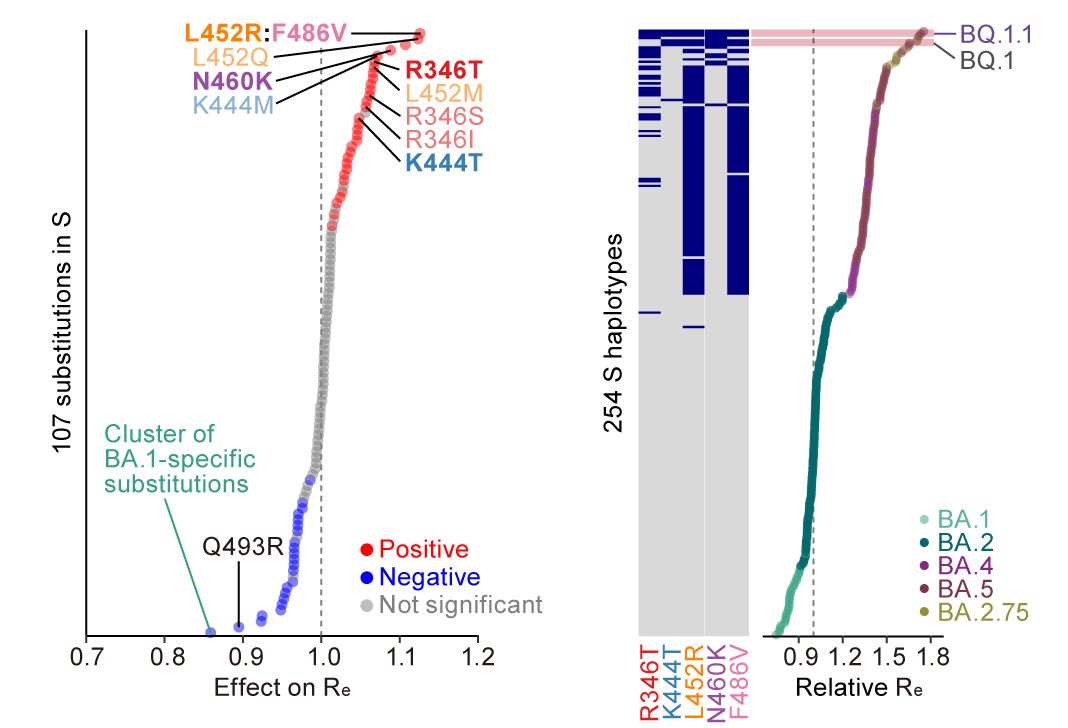
All five convergent substitutions (R346T, K444T, L452R, N460K & F486V) contributed to increasing relative basic reproduction number (Re) (left), and BQ.1.1, which harbors all five convergent substitutions, showed the highest fitness among the viruses investigated (right). 5/ 
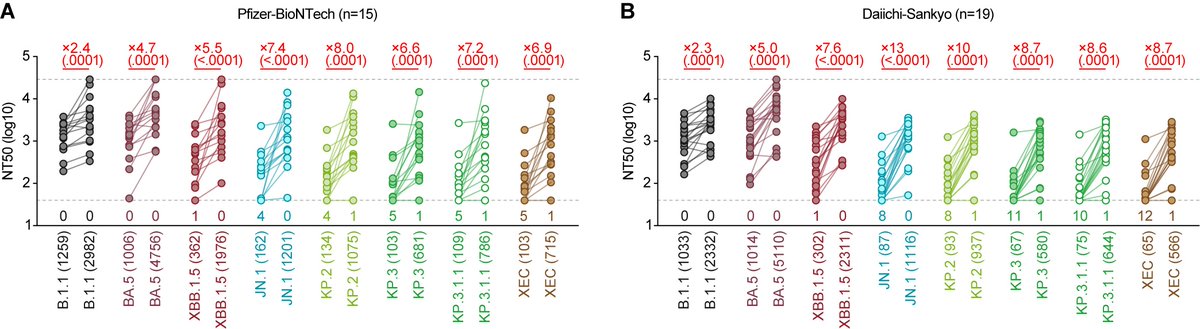
Neutralization assay showed that BQ.1.1 is more resistant to breakthrough BA.2/5 infection sera than BA.5. 6/ 

We have shown lines of data suggesting the close association of viral fusogenicity and intrinsic pathogenicity. For instance, both BA.5 (the link is shown below⬇️) and BA.2.75, descendants of BA.2, are more fusogenic and pathogenic than parental BA.2. 7/
https://twitter.com/SystemsVirology/status/1569995966954233856?s=20&t=YcTm4-TEfIdF9vaST71ihw
Notably, the BQ.1.1 spike exhibited greater fusogenicity than the BA.5 spike, suggesting that BQ.1.1 is more pathogenic than BA.5. 8/ 
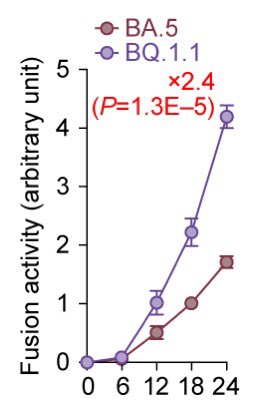
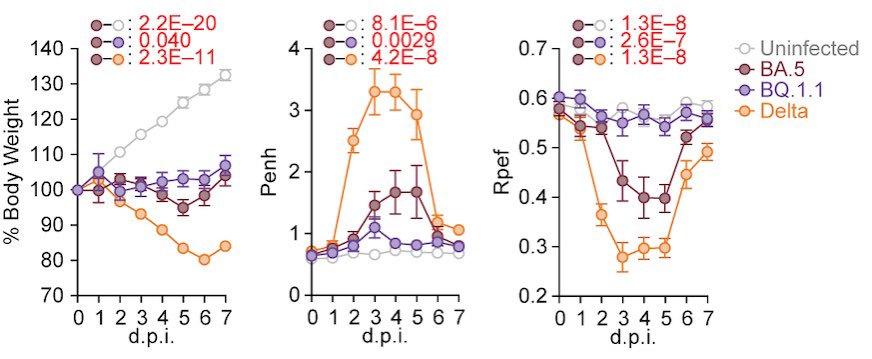
However, unexpectedly, experiments using hamsters showed that the pathogenicity of BQ.1.1 in hamsters is comparable to or even lower than that of BA.5.
Our multiscale investigations provide insights into the evolutionary trajectory of Omicron subvariants. 9/9
Our multiscale investigations provide insights into the evolutionary trajectory of Omicron subvariants. 9/9
• • •
Missing some Tweet in this thread? You can try to
force a refresh