Update on @my_helix SARS-CoV-2 surveillance program
Check public.tableau.com/profile/helix6…
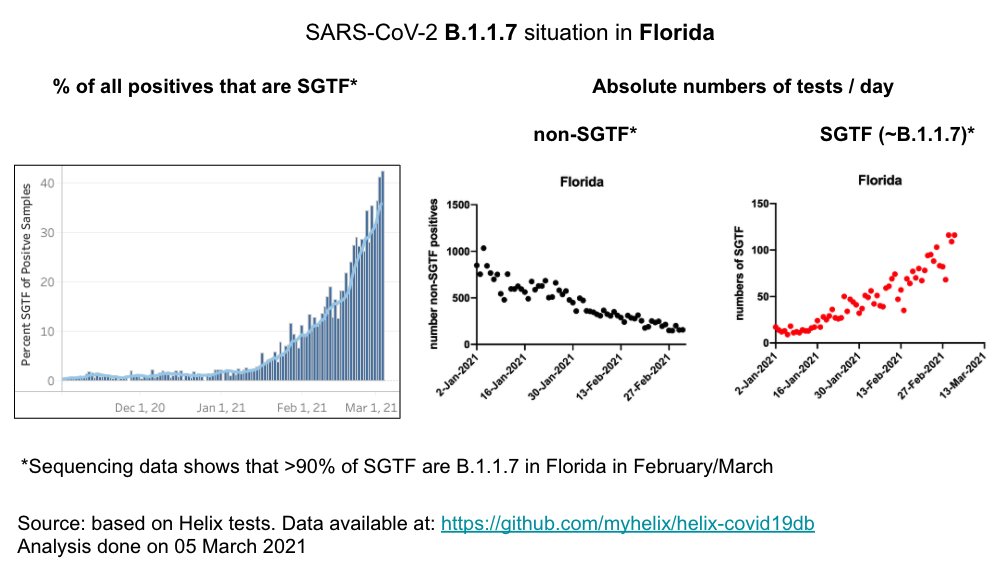
~50% of cases in Florida, Georgia & Texas are B.1.1.7.
in FL: Overall cases continue to 📉 but 52.8% of pos were SGTF (S-gene target failure) on March 7.
=> We will track closely evolution
Check public.tableau.com/profile/helix6…
~50% of cases in Florida, Georgia & Texas are B.1.1.7.
in FL: Overall cases continue to 📉 but 52.8% of pos were SGTF (S-gene target failure) on March 7.
=> We will track closely evolution

2/
In Georgia
54% of positives were SGTF on March 7.
While still ~90% of these are likely B.1.1.7, interesting to note that B.1.525 also sequenced several times in GA and also leads to SGTF
(note: B.1.525 is not a variant of concern)
In Georgia
54% of positives were SGTF on March 7.
While still ~90% of these are likely B.1.1.7, interesting to note that B.1.525 also sequenced several times in GA and also leads to SGTF
(note: B.1.525 is not a variant of concern)

3/
Texas
53% of positives were SGTF on March 7. Overall cases still going down. Again, we will keep testing, sequencing and tracking to see what happens as things reopen, and vaccinated population increases.
Texas
53% of positives were SGTF on March 7. Overall cases still going down. Again, we will keep testing, sequencing and tracking to see what happens as things reopen, and vaccinated population increases.

4/ Also
I am amazed how this real world experiment replicated what was observed & described in UK 3 months earlier.
Different setting, knowledge, people & still same results.
On Feb 07, we said FL would reach 50% B117 on March 8 ... medrxiv.org/content/10.110…
Go science.
I am amazed how this real world experiment replicated what was observed & described in UK 3 months earlier.
Different setting, knowledge, people & still same results.
On Feb 07, we said FL would reach 50% B117 on March 8 ... medrxiv.org/content/10.110…
Go science.

• • •
Missing some Tweet in this thread? You can try to
force a refresh