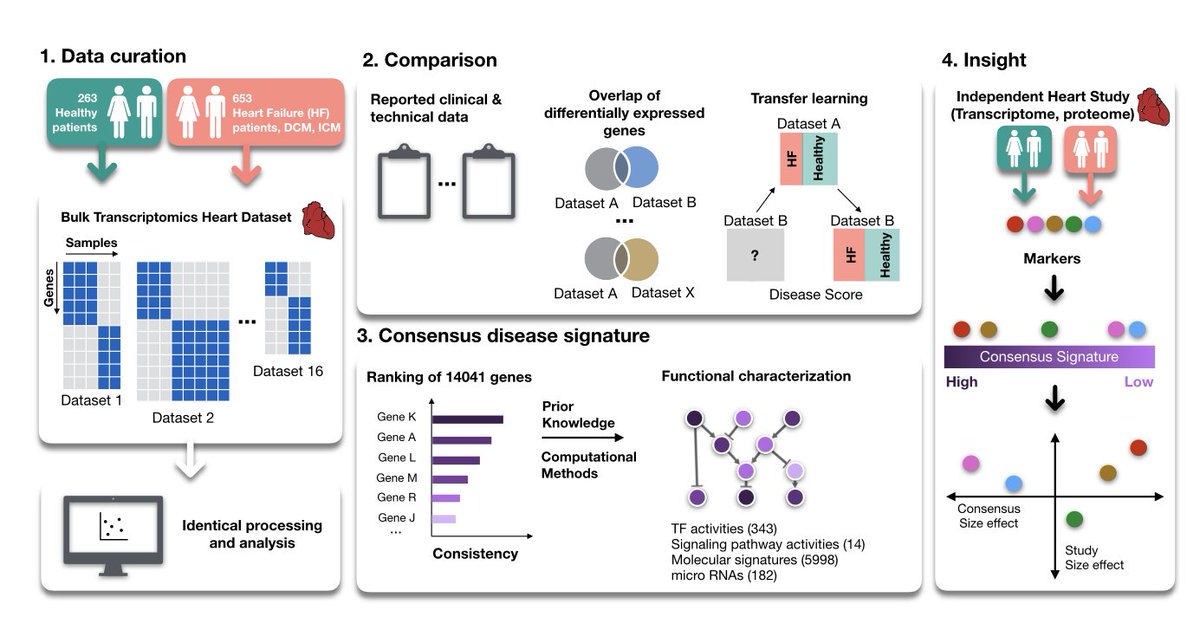
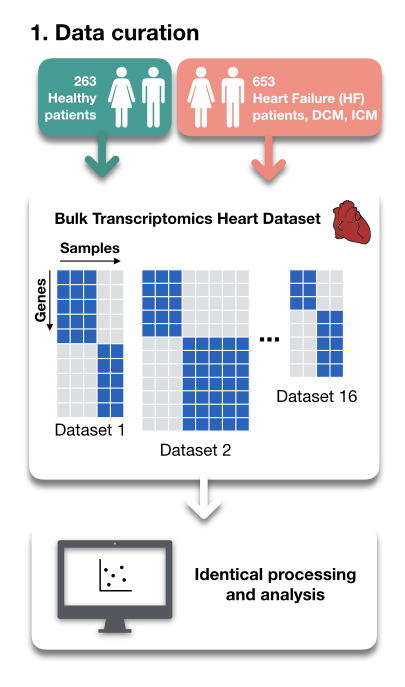
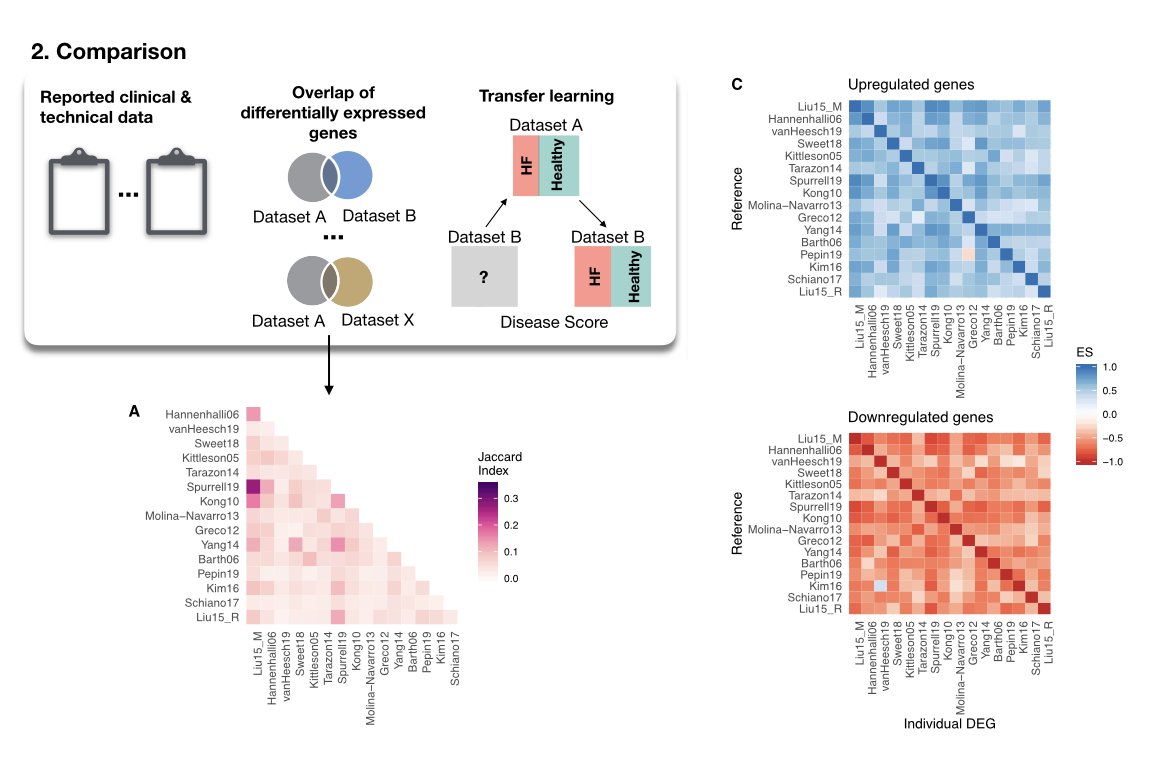
There are many tools and resources to estimate Cell-Cell Communication from scRNA data - do you wonder which one you should use? We do too! To find out, we compared 6 methods and 15 resources doi.org/10.1101/2021.0…. 

For this, we built a framework that decouples Cell-Cell communication tools from their inbuilt resources. The framework is open access and available at github.com/saezlab/ligrec…. 

We analyzed the ‘lineage’ of the different resources that shows how they are all interconnected, but individual resources had varying proportions of the collective CCC prior knowledge. 

We saw that both the resource and the method can have a considerable impact on the results. It is however difficult to know what works best given the lack of ground truths to benchmark (see Supp. Note 4). Do you have any benchmark suggestions? 🙂
We think these disagreements are a word of caution, as the choice of method can largely affect the interpretation of the data. 

We thank all the scientists for the development of these great cell-cell communication tools and resources incl. @msbr89 @mirjana_e @roserventotormo, @teichlab @suoqin_jin @rui_hou_ @JARamilowski @Al__Forrest, @g_palla1, @fabian_theis, @eagut, @hmbaghdassarian, @LewisLabUCSD
And we thank all authors involved @DanielBDimitrov @deeeenes @charlotte_boys @shin_nagai @roramirezf94 @ben_szalai @vanvanka_123 @DugourdAjf @alvaldeolivas @saezlab @JulioSaezRod
We welcome feedback and if this interests you, join us as we extend our framework and explore potential ground truth and simulated datasets.
• • •
Missing some Tweet in this thread? You can try to
force a refresh