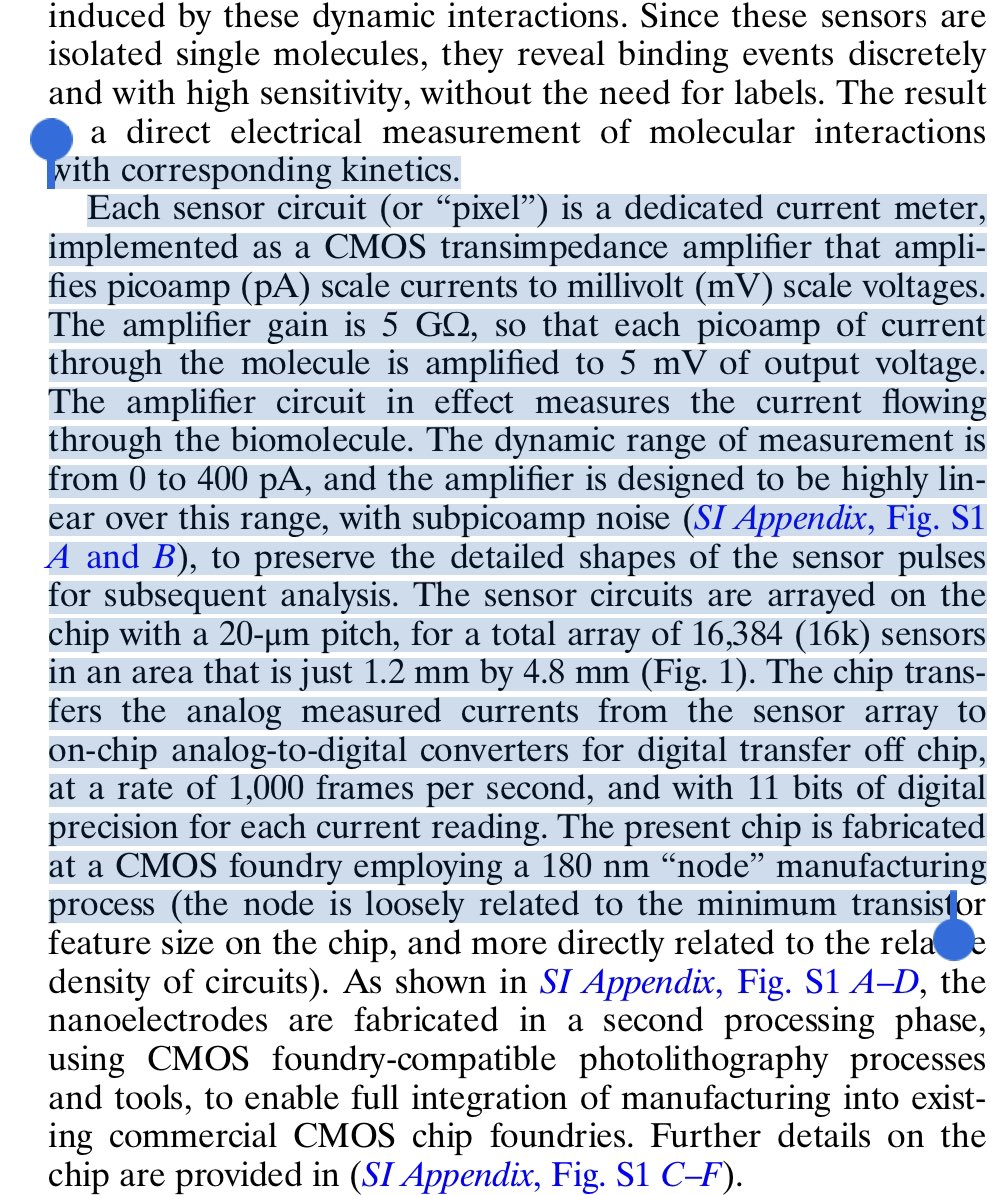
The Persistence length of dsRNA is an interesting thought.
4.2kb is about 1um in length. 43T of these is 43,730 kilometers.
The Circumference of the Earth is ~40,000km.
A 100ug injection is enough degradation resistant RNA in your body to circle the Earth.
4.2kb is about 1um in length. 43T of these is 43,730 kilometers.
The Circumference of the Earth is ~40,000km.
A 100ug injection is enough degradation resistant RNA in your body to circle the Earth.

Caveat... The shot is 99+% ssRNA (in theory) which has a shorter persistence length since it folds on itself but for illustrative purposes, dsRNA is shown. The 4.2Kb is the length of the ssRNA in the shot.
This tweet was inspired by @birb_k and @GirardotMarc
https://twitter.com/birb_k/status/1493003911338225665?s=20&t=aV3XButrVp-hBuwmjjwplw
RNA calculator is from NEB. nebiocalculator.neb.com/#!/ssrnaamt
• • •
Missing some Tweet in this thread? You can try to
force a refresh