Carl Fuller et al are at it again.
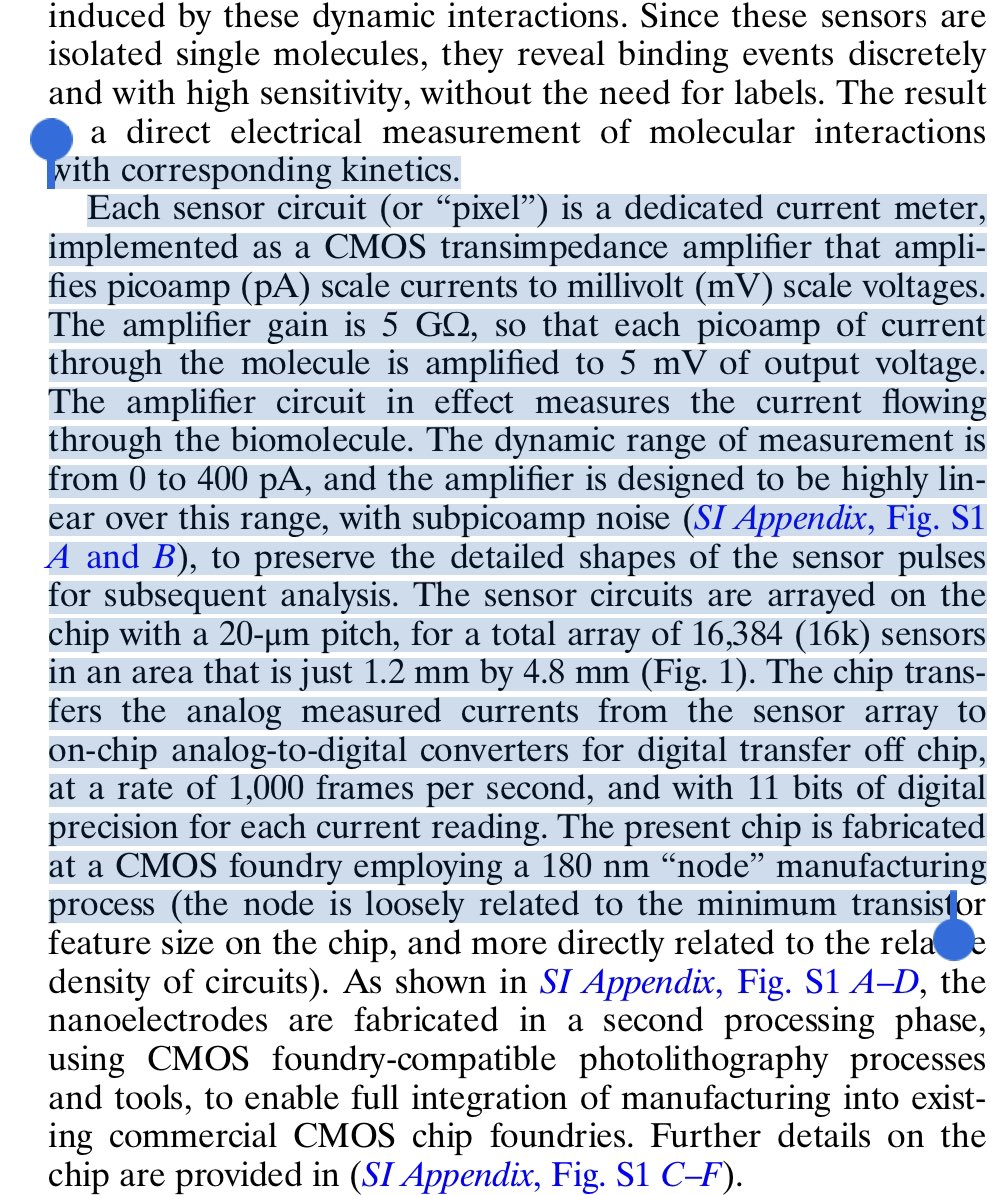
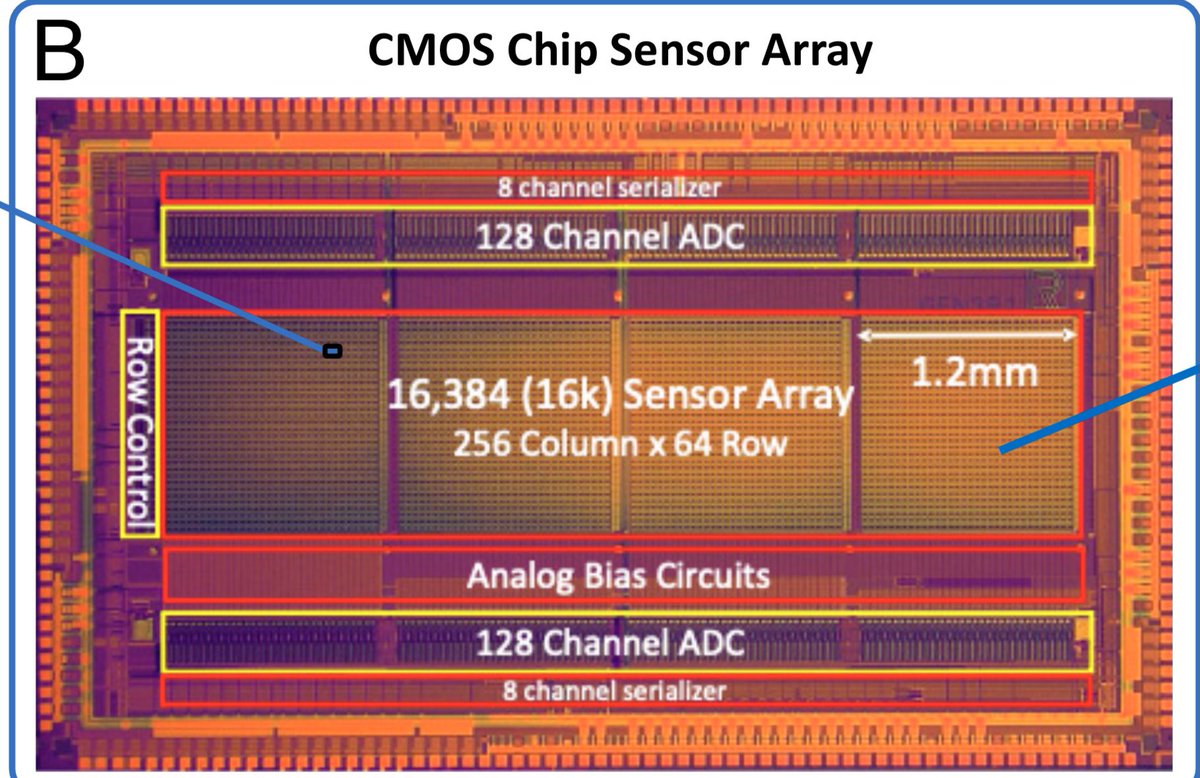
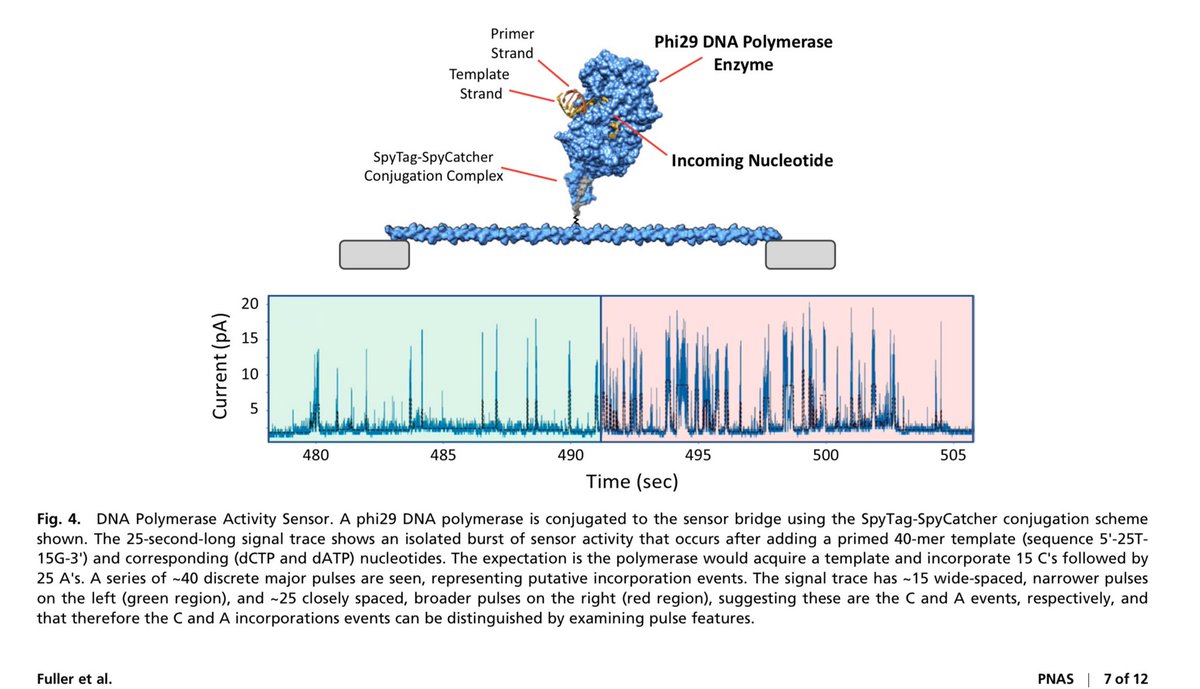
This is wild. 16,384 CMOS chip capable of single molecule movies at each pixel.
Each pixel has a protein nanowire bridging it. These are assembled with electrophoresis.
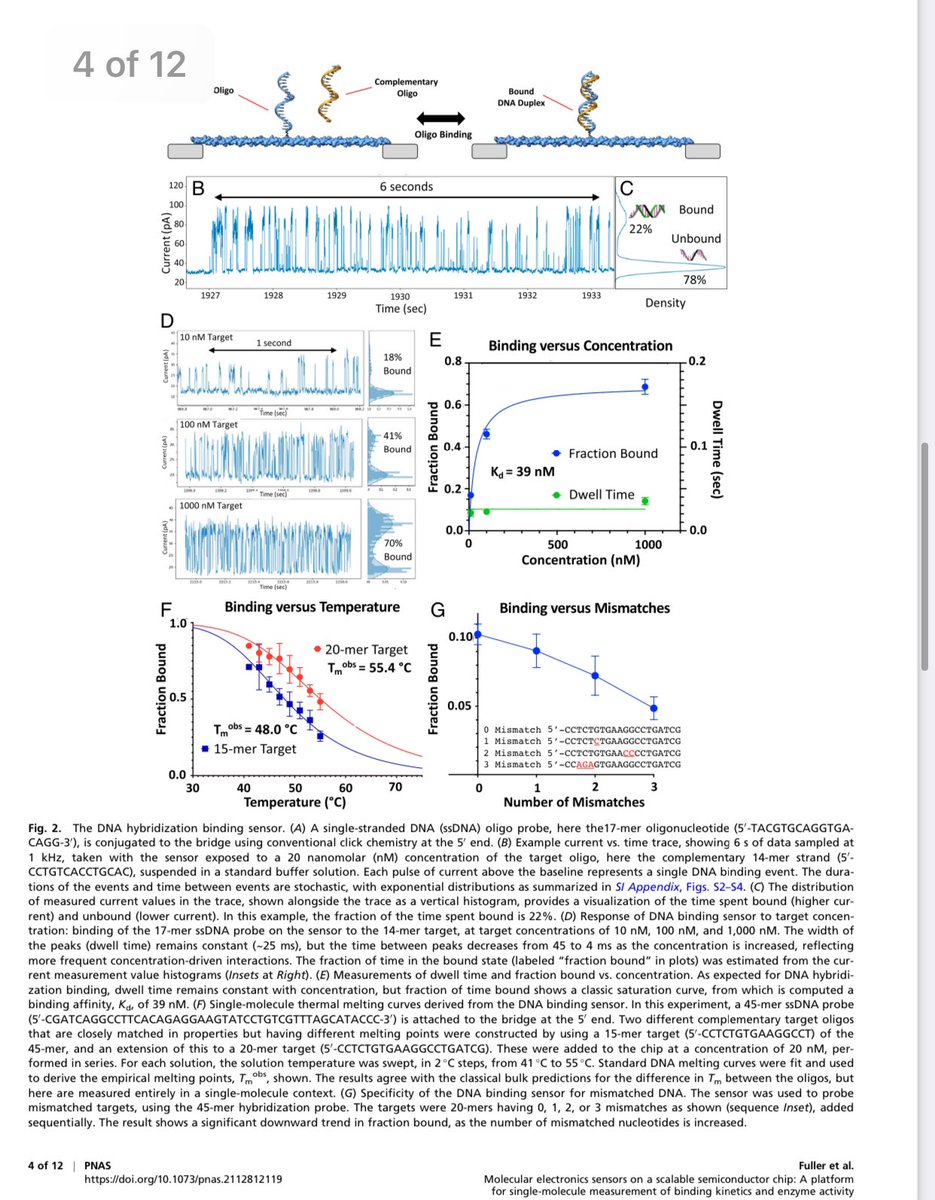
Once wired, click chemistry binds one molecule.
pnas.org/content/pnas/1…
This is wild. 16,384 CMOS chip capable of single molecule movies at each pixel.
Each pixel has a protein nanowire bridging it. These are assembled with electrophoresis.
Once wired, click chemistry binds one molecule.
pnas.org/content/pnas/1…
• • •
Missing some Tweet in this thread? You can try to
force a refresh