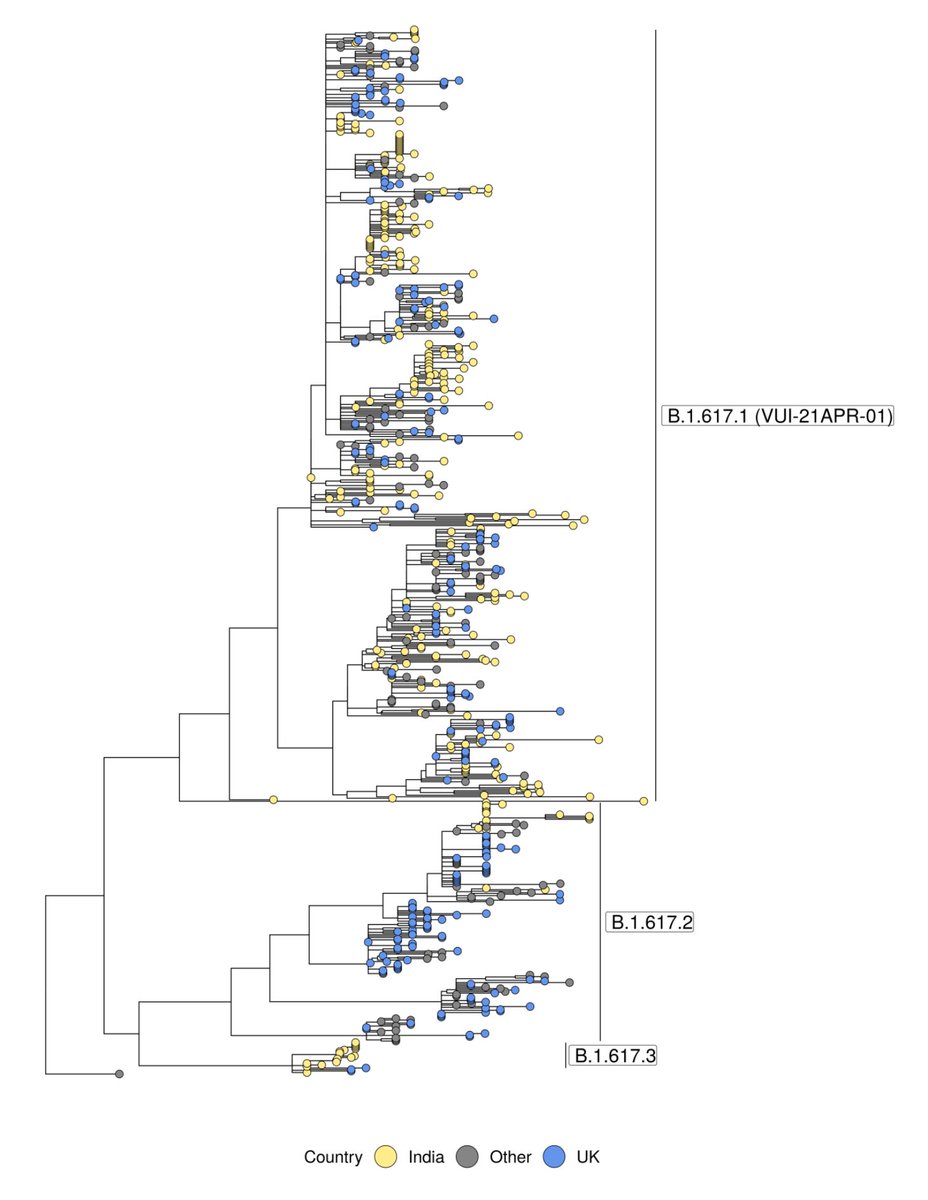
We've pushed the weekly numbers update (thanks @theosanderson!) at covid19.sanger.ac.uk. As expected, in the most recent 2 week window, B.1.617.2 is the most common lineage in England. Still driven by local concentrations of high case numbers (L cases, R B.1.617.2): 

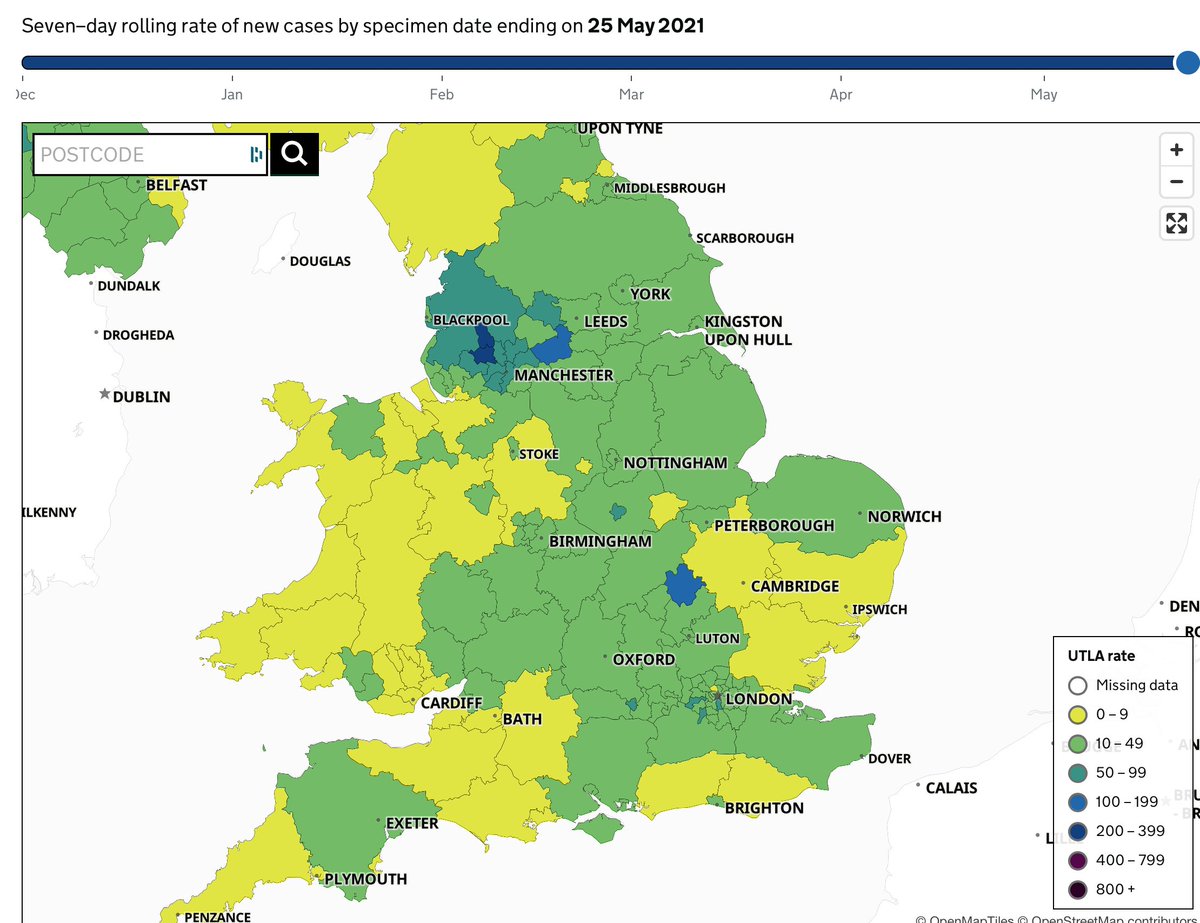

A couple of notes on the site. We have updated the text describing ascertainment: we are no longer excluding surge tests, which make up a large fraction of all tests in key areas in recent weeks. I think this provides least biased frequencies now. Feedback welcome.
I've mentioned this before, but please be cautious interpreting proportions where we have few genomes. For example the "100%" in Dover doesn't mean much because it based on 3 genomes in the most recent 2 weeks of data: 
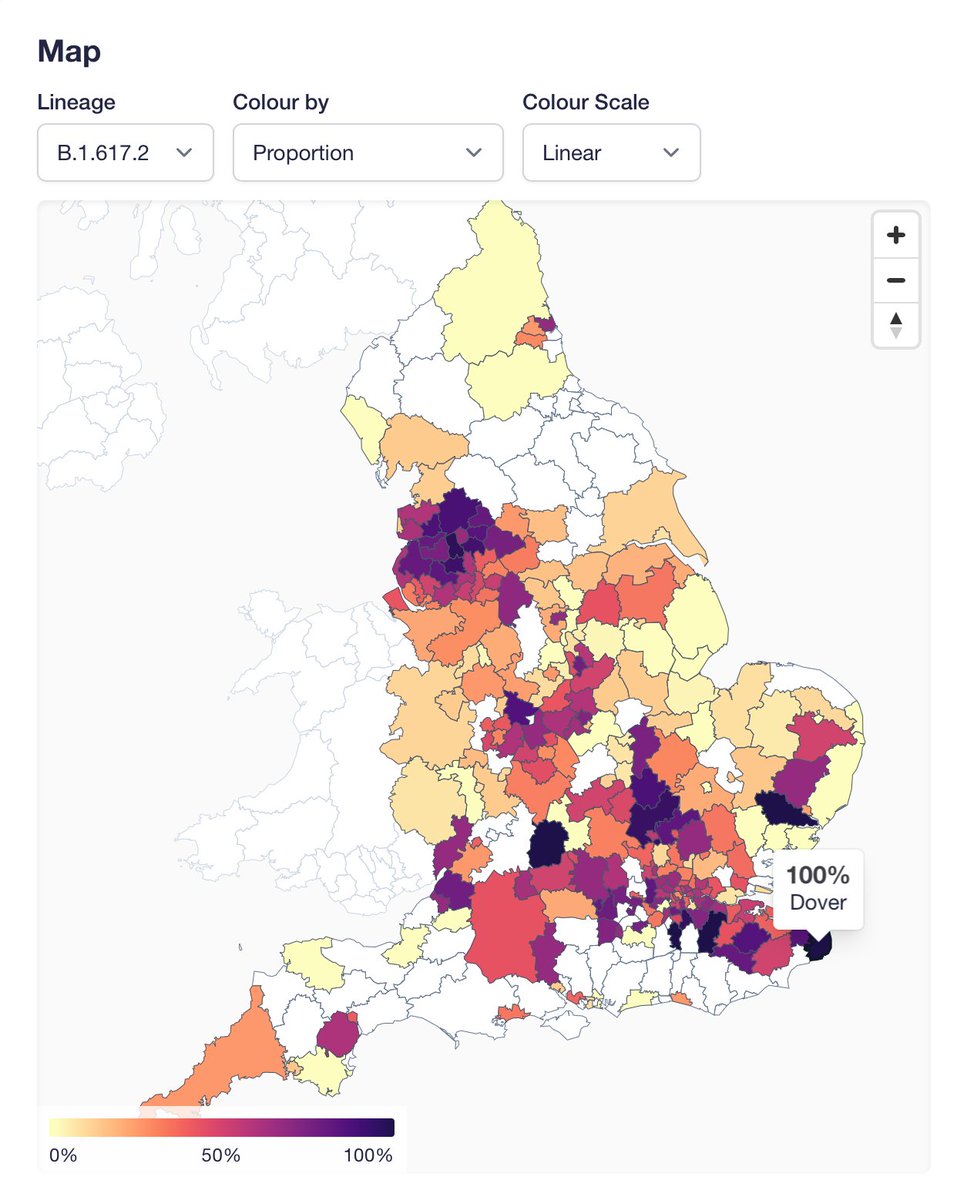
Finally, one thing the proportions map is good for is actually to spot our coverage holes (white areas). Some will be filled soon (e.g. we are now sequencing samples from the Plymouth lab covering some of the Southwest).
• • •
Missing some Tweet in this thread? You can try to
force a refresh