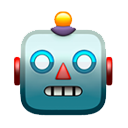
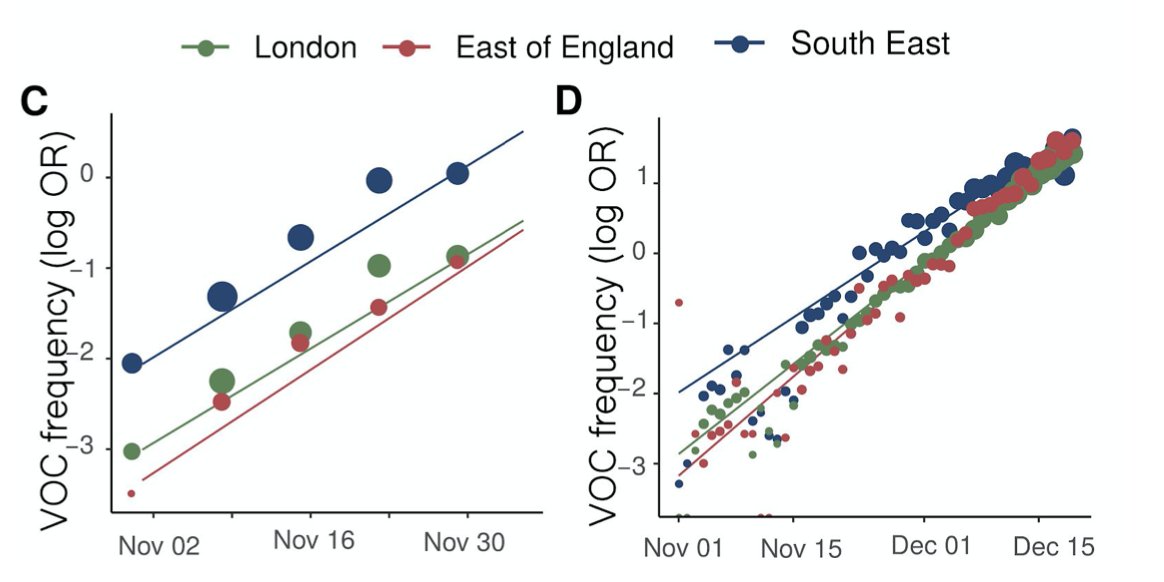
Tons of new info about B.1.617 in @PHE_uk's latest technical briefing on covid #variants (which are rather excellent if I do say so). A 🧵on some key points. assets.publishing.service.gov.uk/government/upl…
First off: what is the "India variant", well, it turns out that there are actually three clades of B.1.617, which have now been termed B.1.617.1, B.1.617.2 and B.1.617.3. All three clades appeared in India, likely descended from a common ancestor there some time ago. 
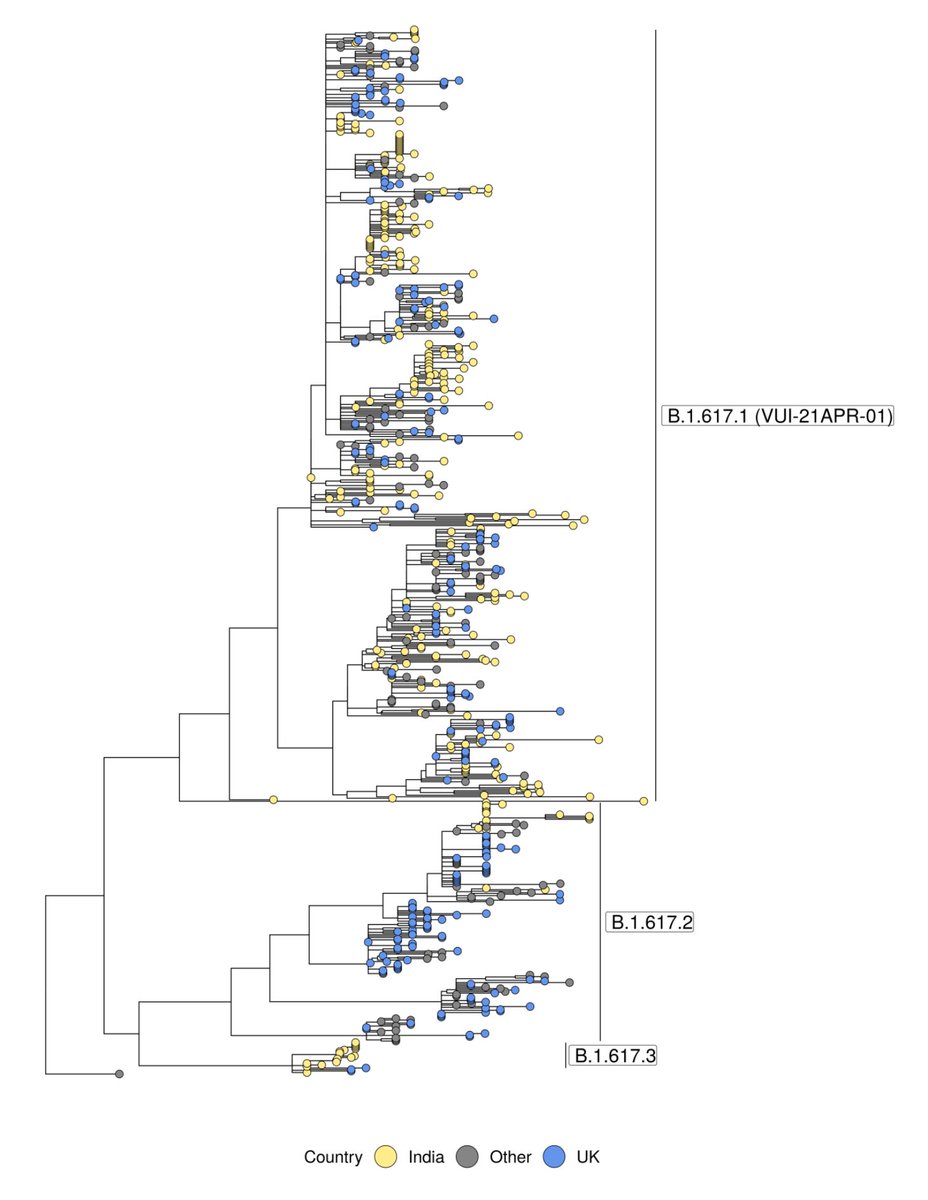
The "full" VUI that has garnered attention is B.1.617.1, but interestingly B.1.617.2, which does *not* have the E484Q mutation (apparent reversion) is enriched in the most recent sequences in the UK & elsewhere. So the already dumb "double mutant" label makes even less sense now.
You'll see the UK sequences (blue dots) are all over the tree above, and also all over the map below, strongly suggesting multiple introductions, rather than substantial community spread. 
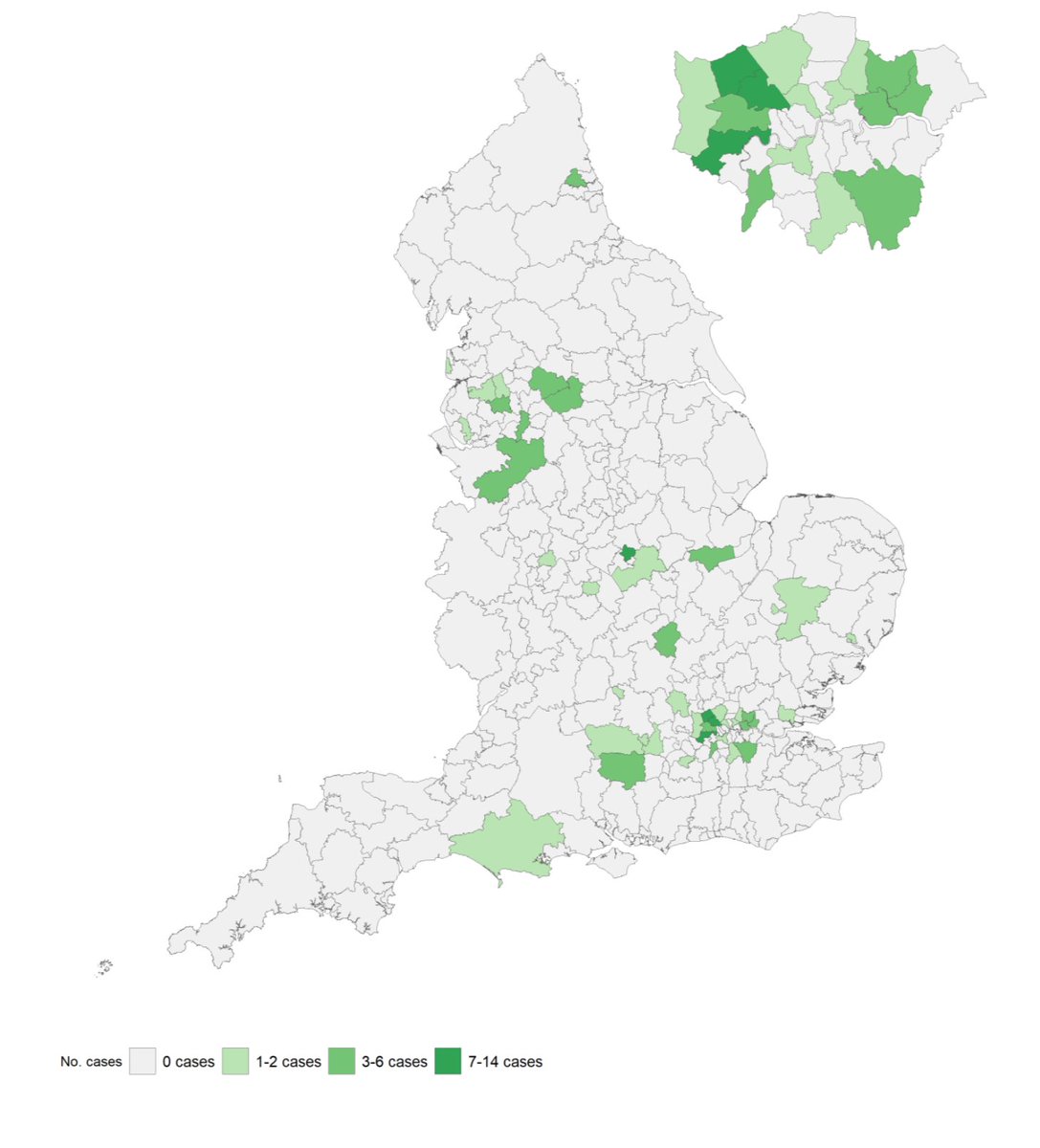
And indeed, 97% of cases with completed investigations have either recently traveled to the UK, or are contacts of someone who has: 
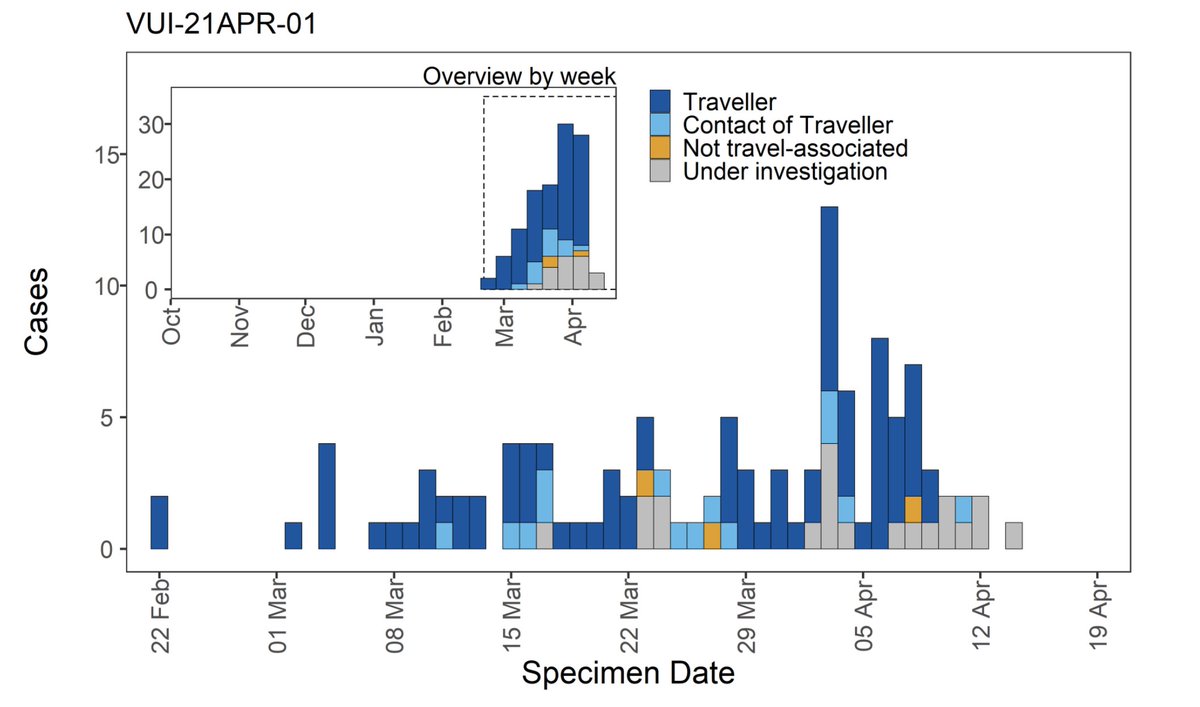
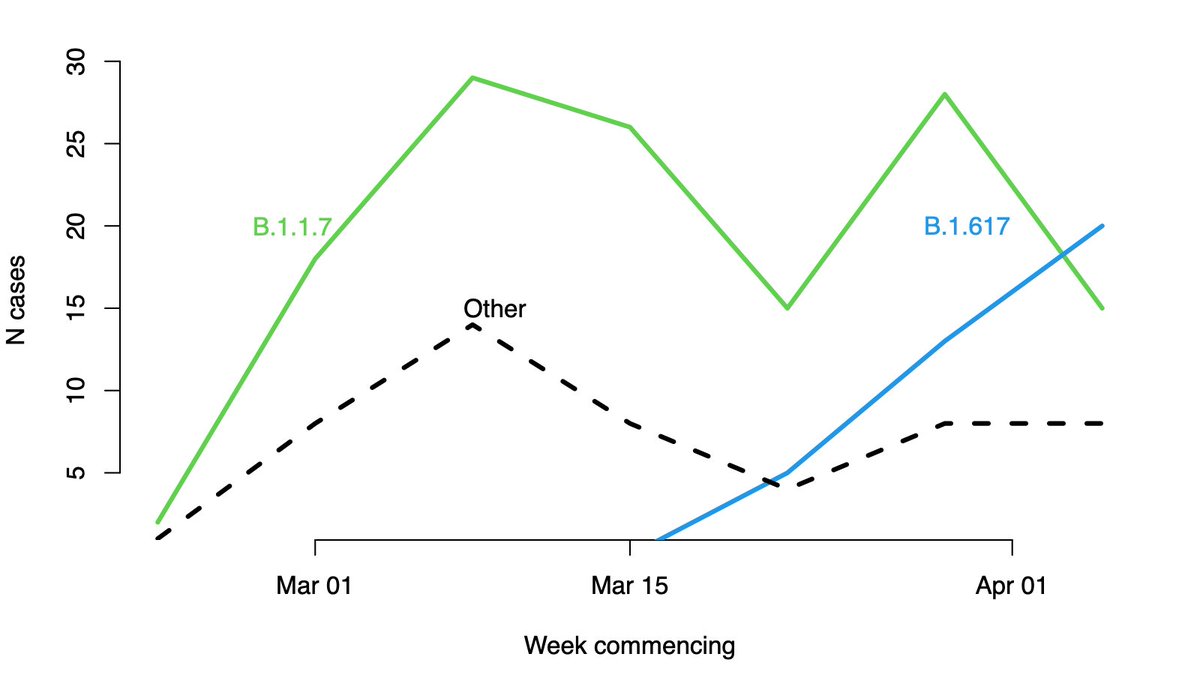
What if the question is flipped around: what variants of the virus are found in people who have recently arrived from India? Unsurprisingly there is re-import of B.1.1.7 (though that was covered less breathlessly by the media), and as expected we do see an increase in B.1.617 
So, in the space of a week we've learned that B.1.617 isn't what people assumed it was, and that the surveillance system in the UK has detected it at the border and (so far) kept it from community transmission. Big congrats to all the PHE team who did this work so fast.
Finally, while I hope these analyses help understand the virus, which variants are infecting people doesn't change hte tragedy in India. What matters now is supplying oxygen, sharing vaccines, and preventing transmission. Best wishes to those in the front lines of this fight.
• • •
Missing some Tweet in this thread? You can try to
force a refresh