Here we can see honest virologists being confronted by ones that might have been named in Fauci’s email.
The later amplify flawed preprints from Drosten.
The reads are not out yet (of course), so there is not much to say about this headline whore print.
The later amplify flawed preprints from Drosten.
The reads are not out yet (of course), so there is not much to say about this headline whore print.
https://twitter.com/jbloom_lab/status/1471573160654880769
Let’s have a look at the methods section of the paper.
2 rounds of 45 cycles of PCR?
This is what you do if you want to make polymerase slippage errors near GC rich triplet repeats.
People this library illiterate shouldn’t have the keys to $20K NovaSeq runs.
2 rounds of 45 cycles of PCR?
This is what you do if you want to make polymerase slippage errors near GC rich triplet repeats.
People this library illiterate shouldn’t have the keys to $20K NovaSeq runs.

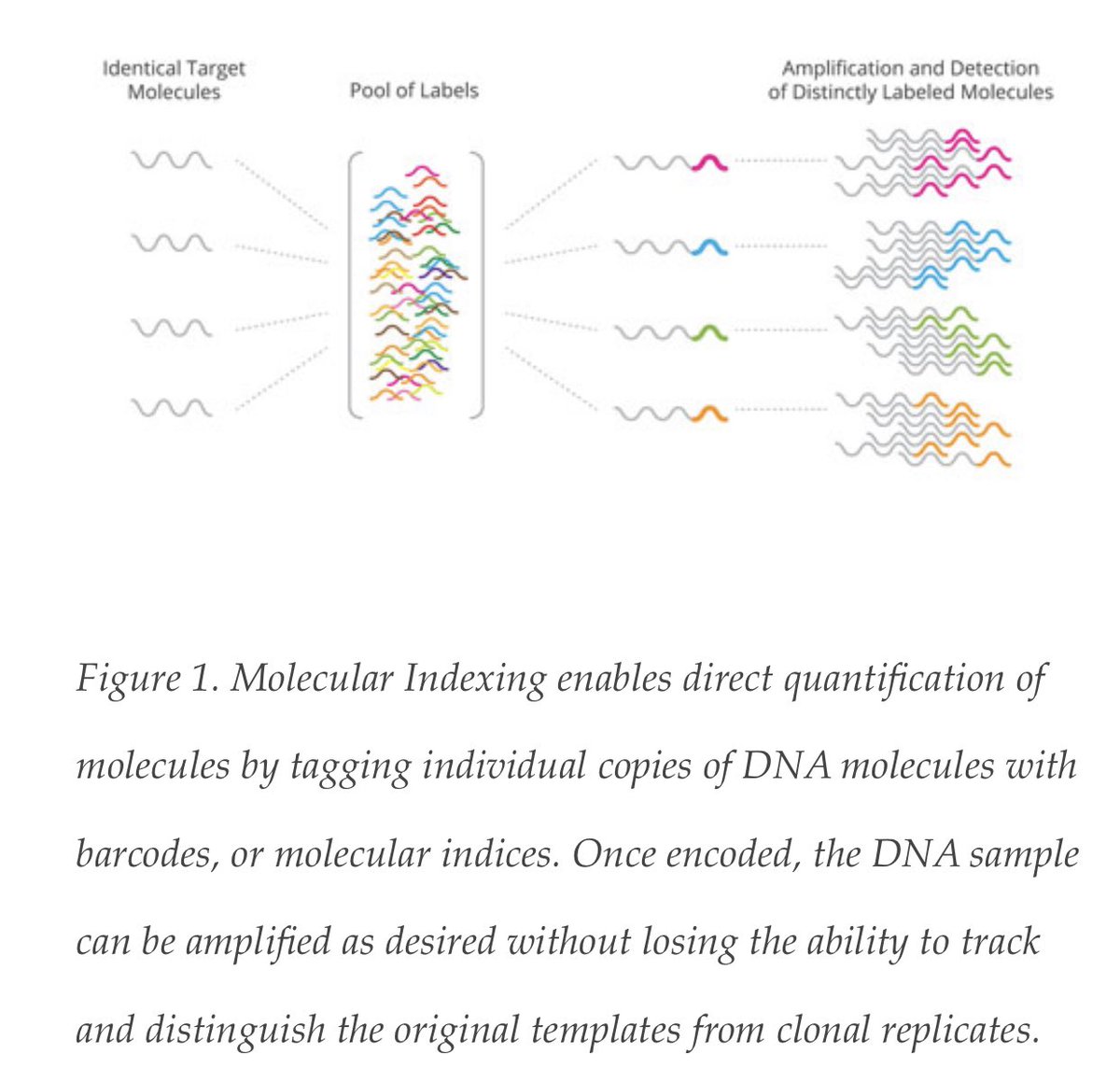
Having engineered very accurate sequencers, to address PCR polymerase error, you must utilize DNA barcodes that get attached to your PCR products in the first cycle of PCR. Any barcode that over amplifies through the PCR process is discounted.
For rare variants
This is the way
For rare variants
This is the way
The authors are trying to excuse the sloppy leaky virology labs all over the world by fabricating excuses that support zoonotic origins with very sloppy methods and the Bloom lab gave it to them.
90 cycles PCR
No molecular indexes (barcodes)
Squinting at unpaired reads
Bunk
90 cycles PCR
No molecular indexes (barcodes)
Squinting at unpaired reads
Bunk
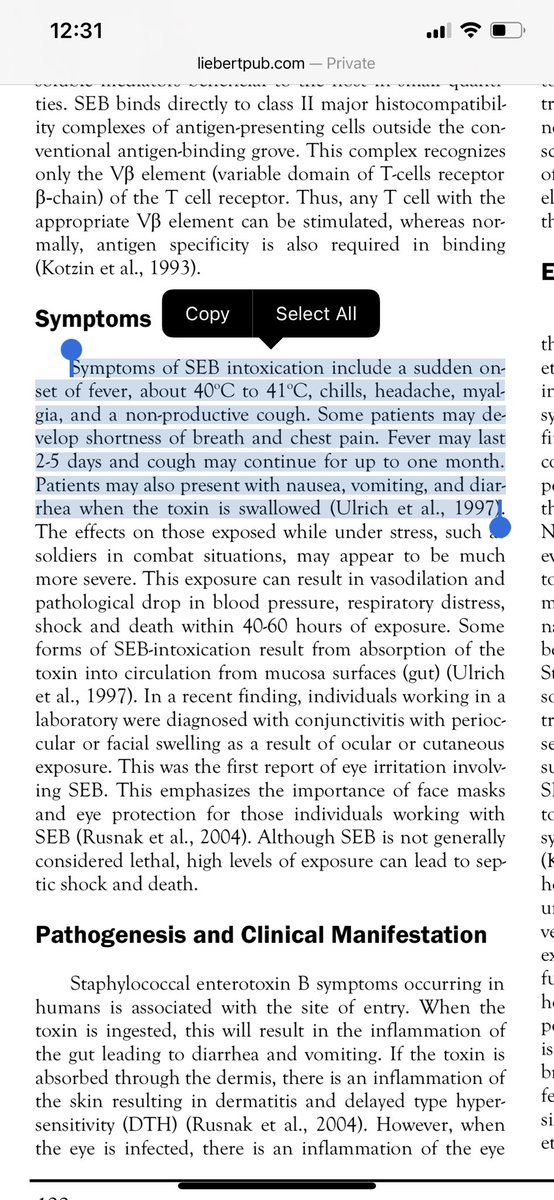
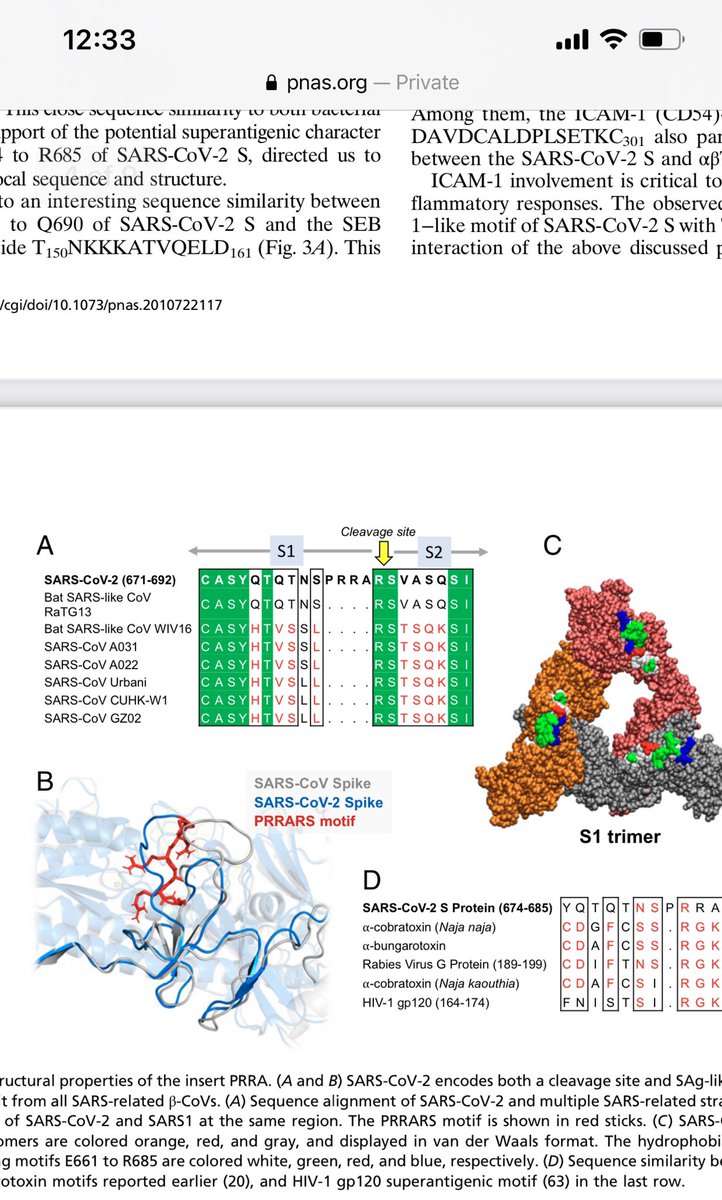
For those who like a deep dive on Illumina error rates, they tend to pile up on or 10bases past GGC motifs.
Probably because these regions for secondary structure on the Illumina arrays.
The FCS they are making claims about is GGCGGC window.
ncbi.nlm.nih.gov/pmc/articles/P…

Probably because these regions for secondary structure on the Illumina arrays.
The FCS they are making claims about is GGCGGC window.
ncbi.nlm.nih.gov/pmc/articles/P…


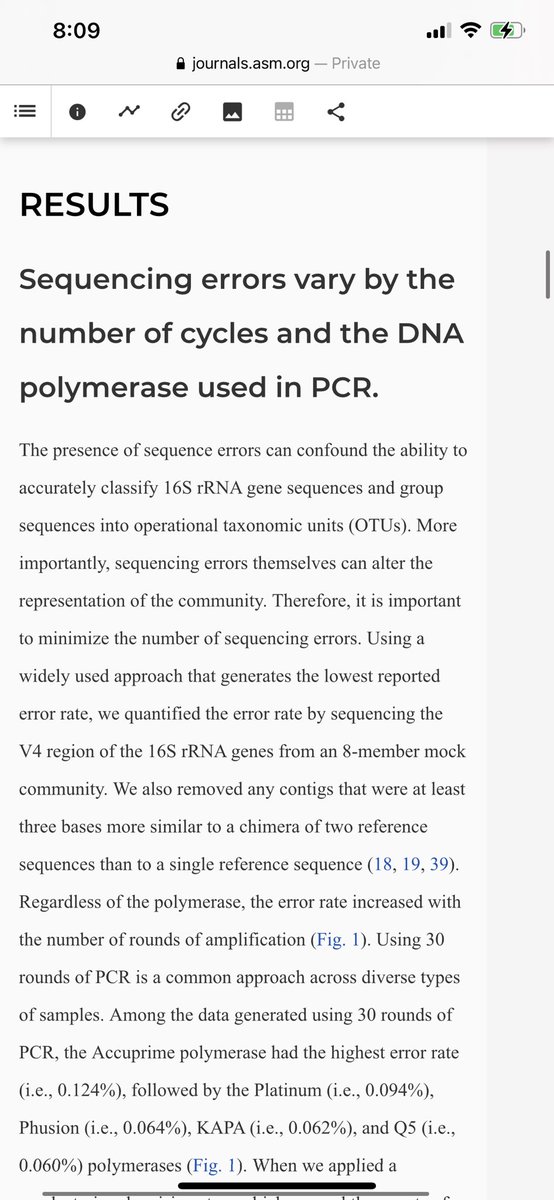
These authors don’t even bother to measure the PCR error after 30 cycles. Notice it grows exponentially.
journals.asm.org/doi/10.1128/mS…

journals.asm.org/doi/10.1128/mS…


• • •
Missing some Tweet in this thread? You can try to
force a refresh