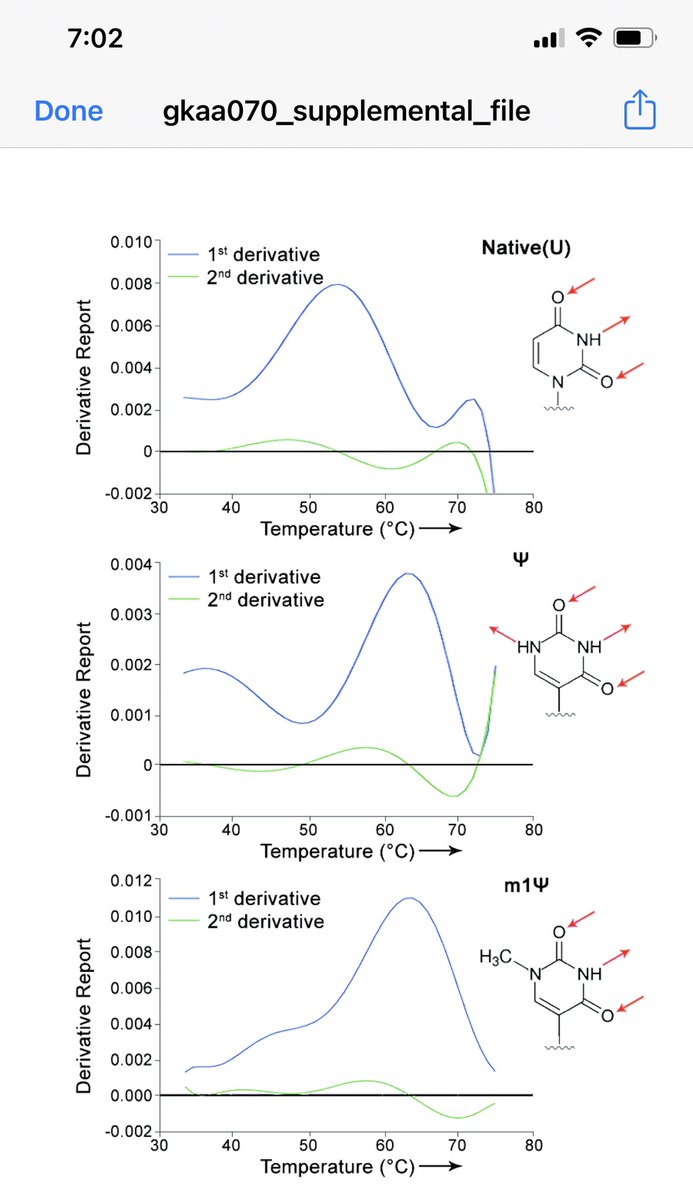
We have Cannabis Whole Genome Sequencing honed to the point where people are using it to untangle the history of the famous Skunk #1 line.
I wasnt around then so I cant speak with any authority on the oral history but I can help people better understand Kannapedia.net
I wasnt around then so I cant speak with any authority on the oral history but I can help people better understand Kannapedia.net
Phylo-Trees can be complicated so lets just take a look at the genetics of THCAS.
There are a few interesting mutations found in early cannabis lines that we will go over.
Ala250Asp
Pro333Arg
Pro542Leu
Ser355Asn
A recently sequenced Skunk line
kannapedia.net/strains/rsp124…
There are a few interesting mutations found in early cannabis lines that we will go over.
Ala250Asp
Pro333Arg
Pro542Leu
Ser355Asn
A recently sequenced Skunk line
kannapedia.net/strains/rsp124…
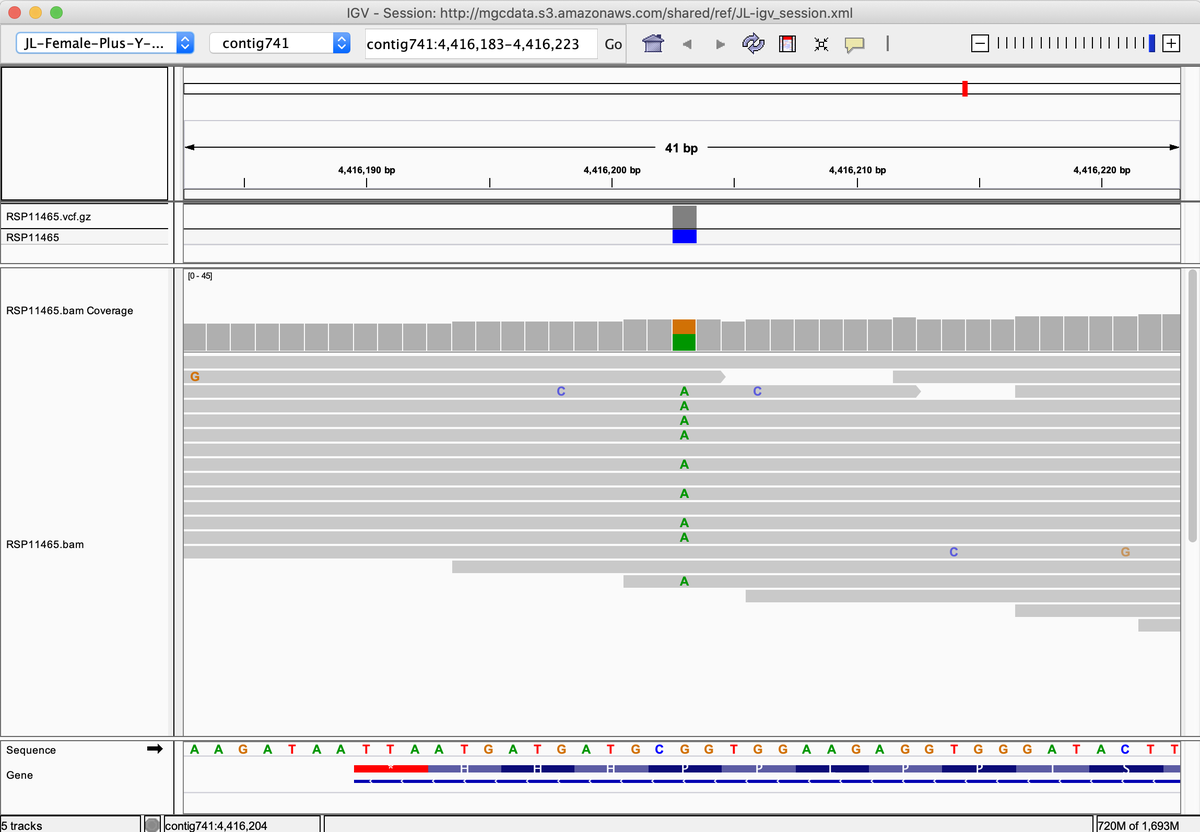
A250D and Pro333Arg are some of the most common mutations.
A250D is found in 12.7% of the NGS data. The C90 data finds this more frequently but less samples have been run through that pipeline.
P333R is found in 18.2% of the NGS data.
Click on the blue %Number


A250D is found in 12.7% of the NGS data. The C90 data finds this more frequently but less samples have been run through that pipeline.
P333R is found in 18.2% of the NGS data.
Click on the blue %Number
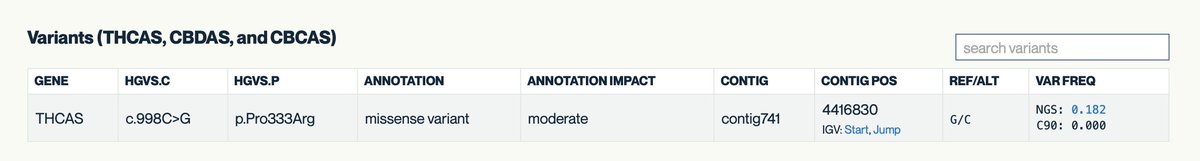


Kannapedia.net will bring up all of the strains (83) that share that THCAS mutation (P333R). Very helpful for sorting out the ancestry of just THCAS.
One can repeat this for A250D and get a list of 58 strains.

One can repeat this for A250D and get a list of 58 strains.
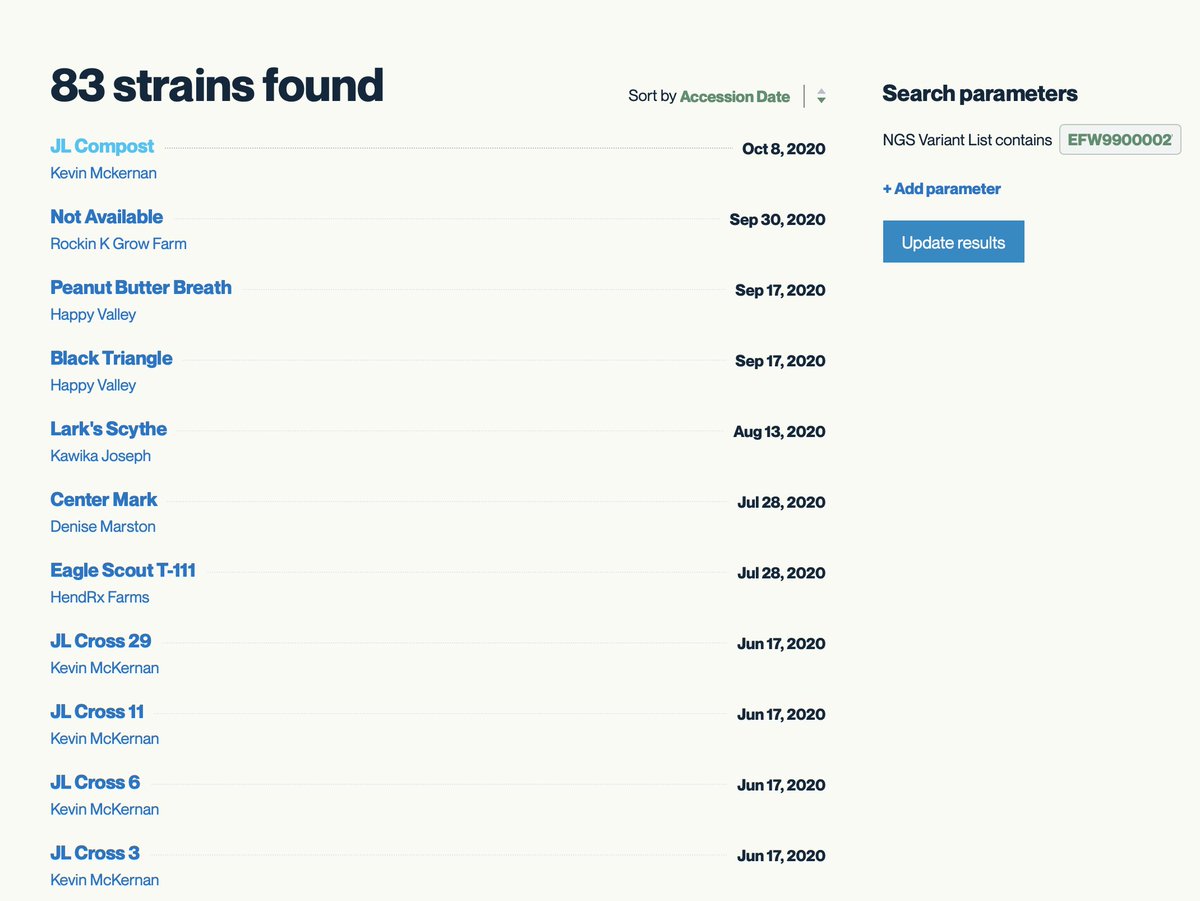
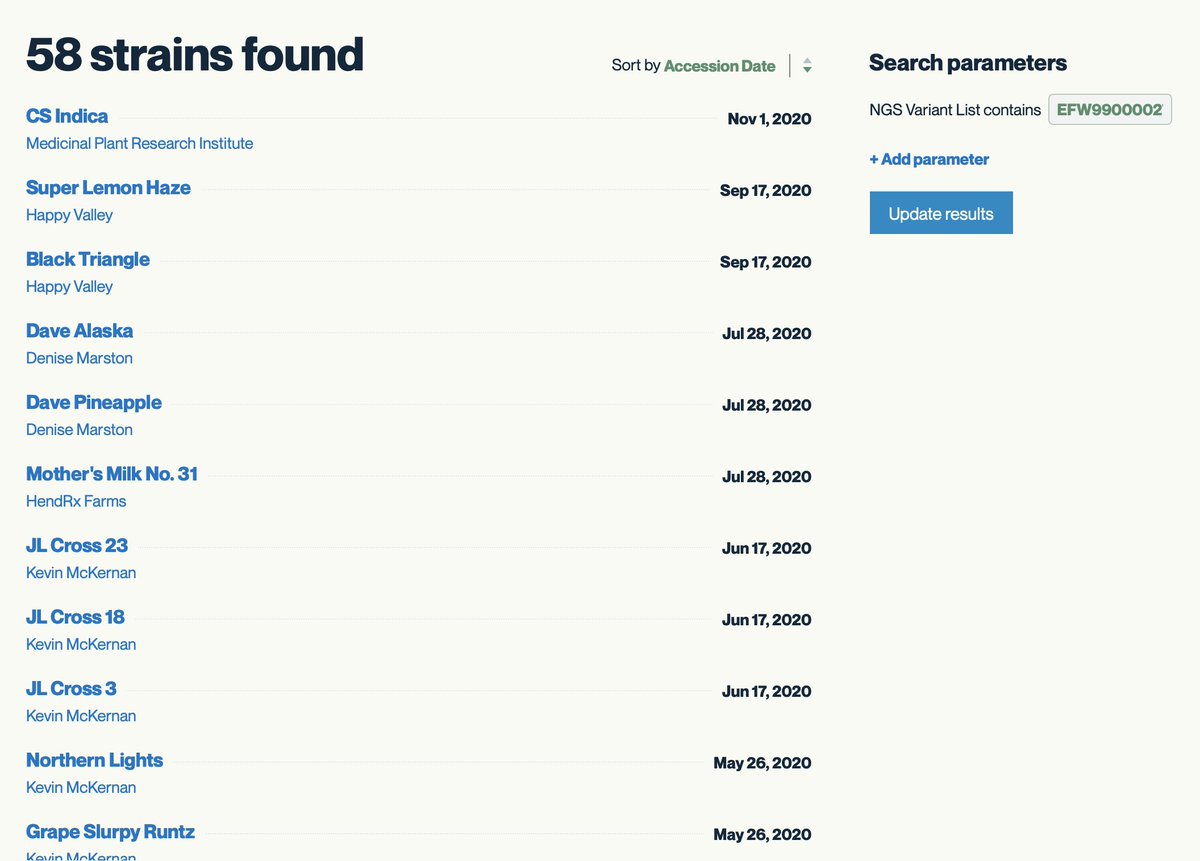
Kannapedia.net is a bit light on strains with Skunk in the name so don't take this as gospel trying to build a distributed consensus.
But of interest...
There is a Big Skunk from the Chinese Landrace study. Same THCAS as the JL reference
kannapedia.net/strains/srr147…
But of interest...
There is a Big Skunk from the Chinese Landrace study. Same THCAS as the JL reference
kannapedia.net/strains/srr147…
The most recent Skunk added to Kannapedia has different THCAS.
P542L. Only 1.8% of the sample have this mutation.
kannapedia.net/strains/rsp124…
P542L. Only 1.8% of the sample have this mutation.
kannapedia.net/strains/rsp124…
8 other strains share this rare mutation. It is close to the end of the gene and its not clear what the function of this mutation is but in general Proline changes should be monitored.
kannapedia.net/strains?NGS=EF…
kannapedia.net/strains?NGS=EF…
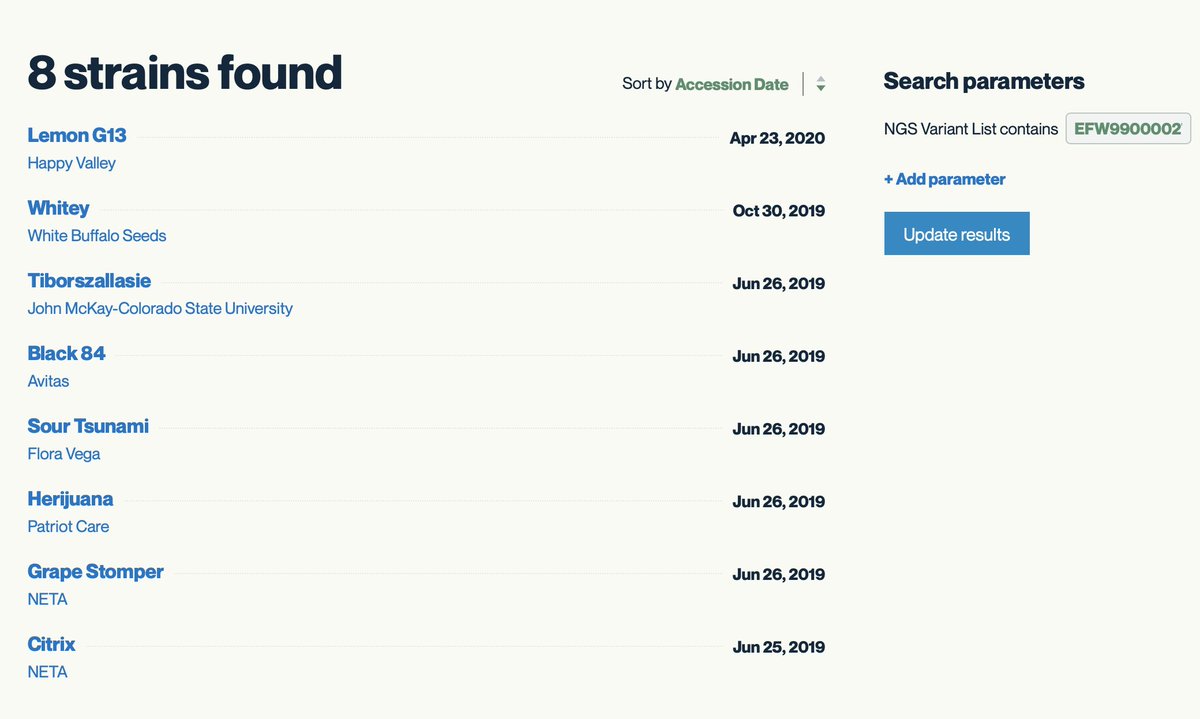
So other than using these variants as a mechanism to track ancestry, do any of the mutations play a role in THCA production?
It's not clear that individual mutations are having an impact. You must recall, there are at least 2 THCAS in every genome. P542L is heterozygous
It's not clear that individual mutations are having an impact. You must recall, there are at least 2 THCAS in every genome. P542L is heterozygous
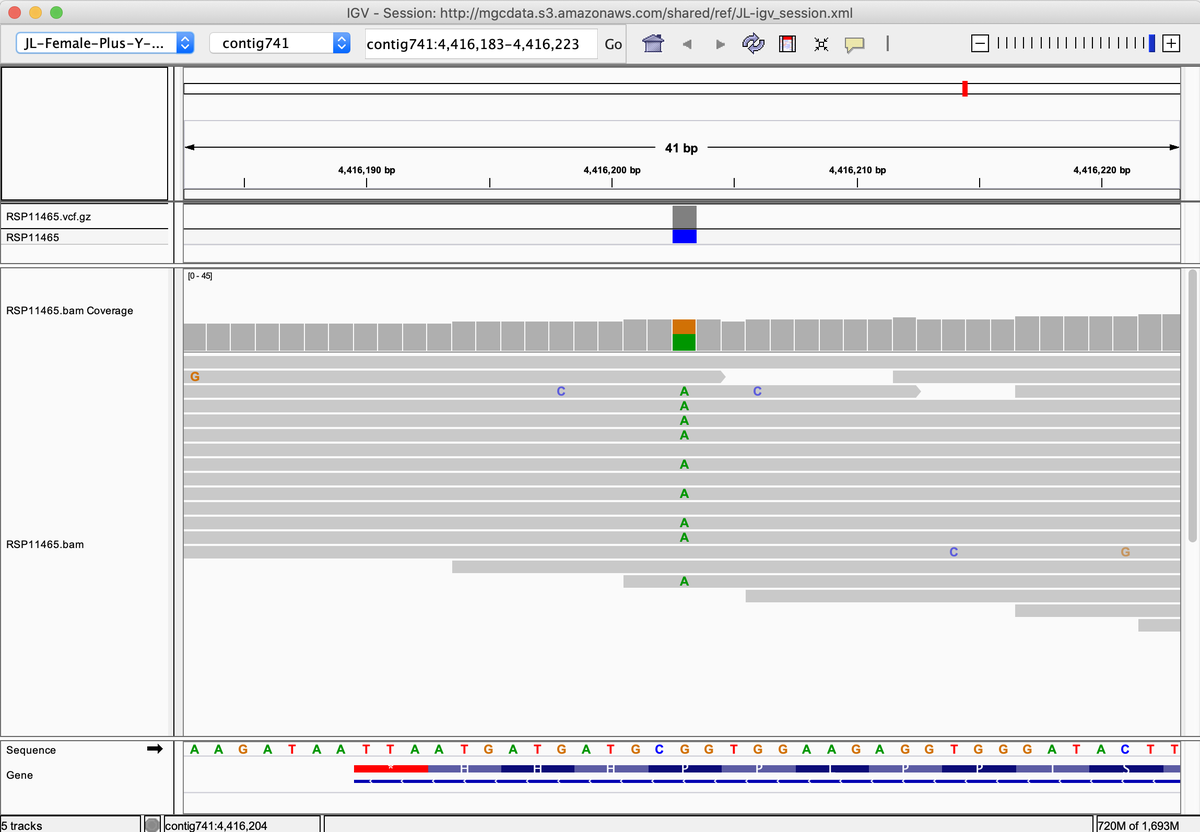
A few samples in Kannapedia.net have this variant in a homozygous state. One is an old Hemp line (Tiborszallasie).
kannapedia.net/strains/rsp112…
kannapedia.net/strains/rsp112…
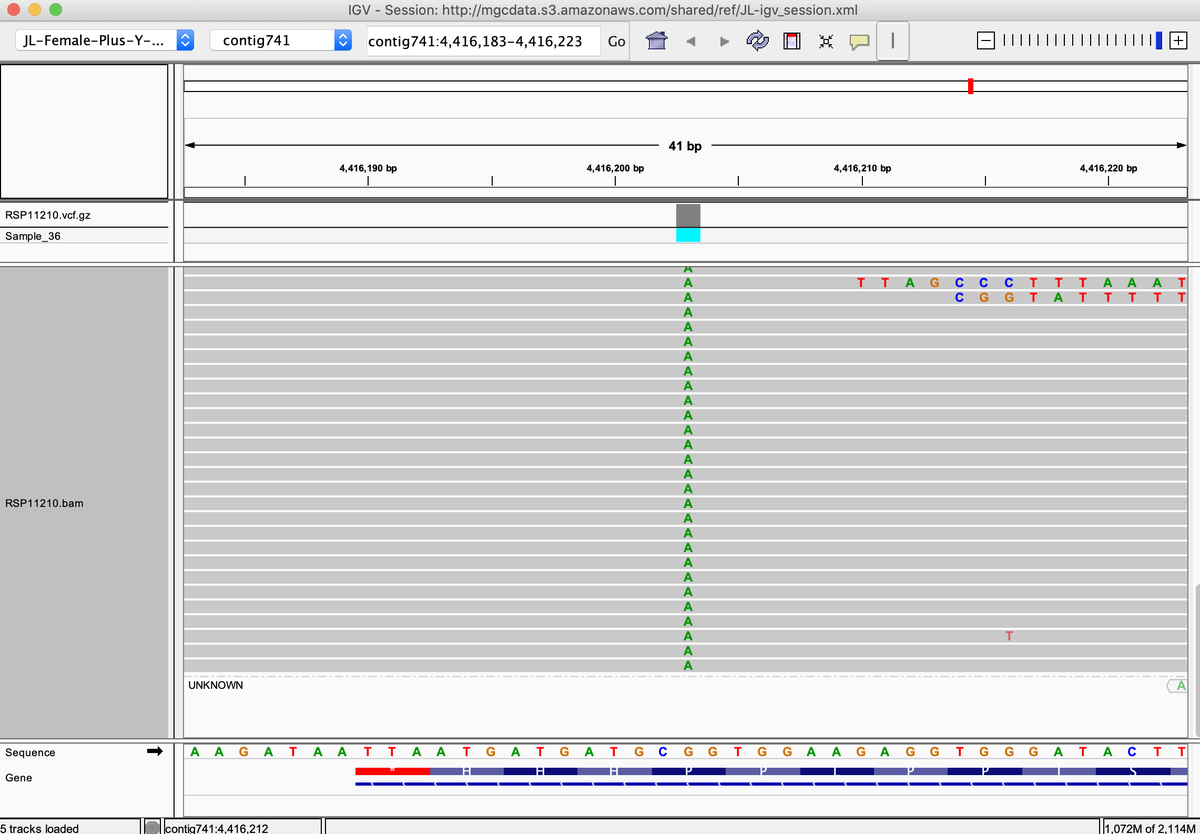
Sour Tsunami is another one. kannapedia.net/strains/rsp111…
This also happens with A250D. Most strains are heterozygous but some are homozygous for the variant. kannapedia.net/strains/rsp111… 

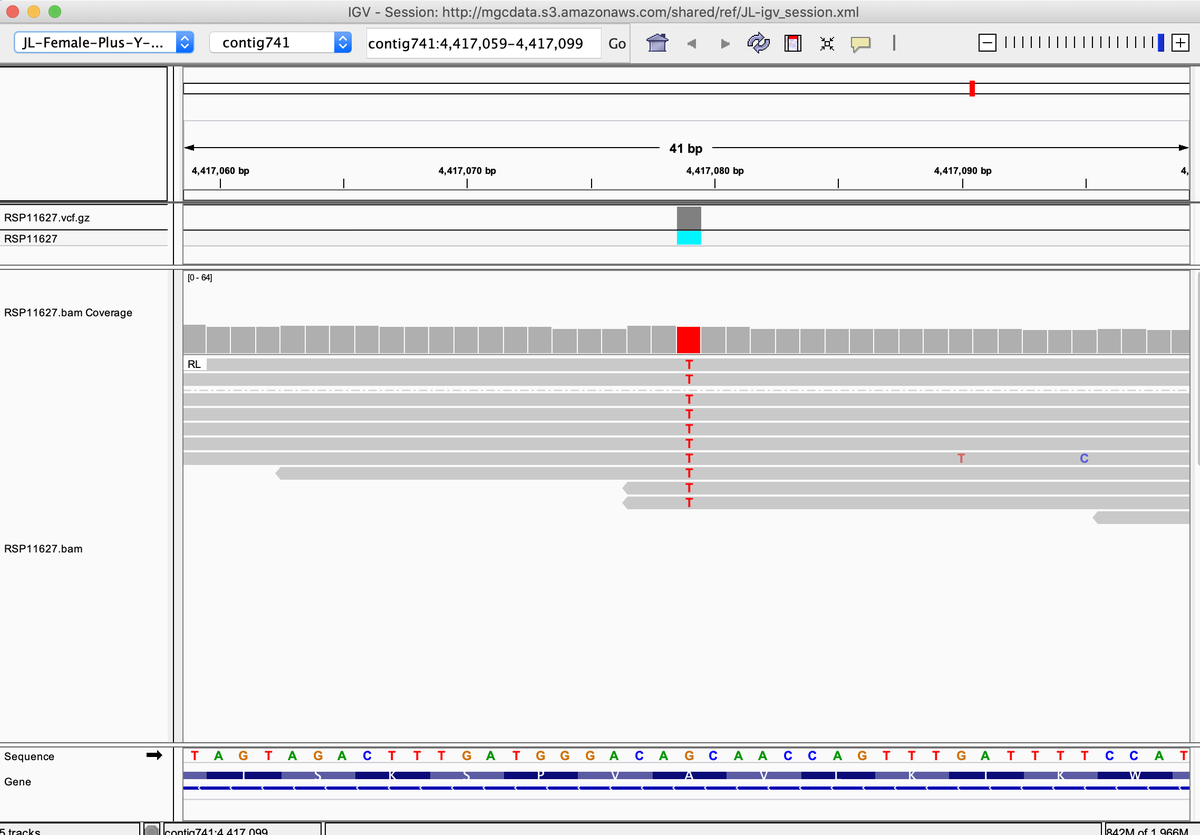
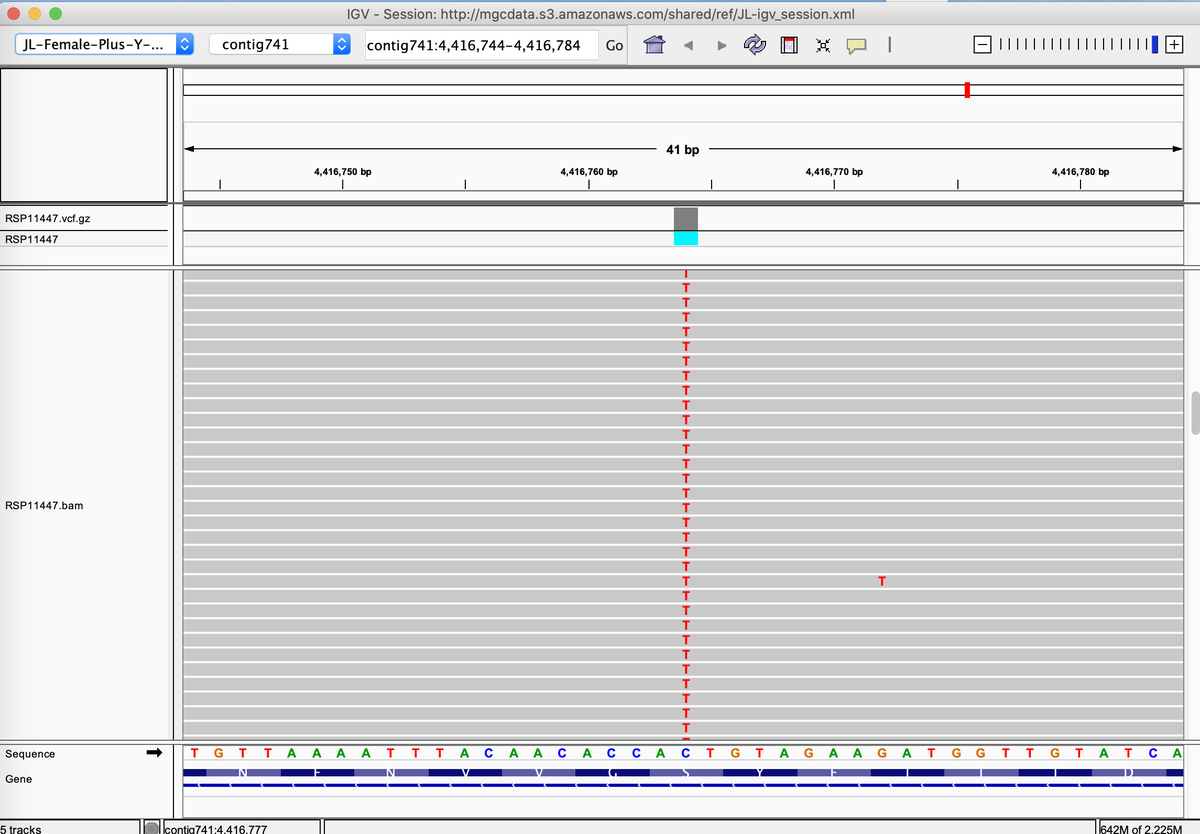
Why do these Homozygous variants matter?
When both copies of THCAS are impacted, their impact on expression is more obvious.
Have a look at CBG lines.
This is a common mutation we see in some CBG lines that knocks out THCAS activity. Often found with homozygous P333R & S355N

When both copies of THCAS are impacted, their impact on expression is more obvious.
Have a look at CBG lines.
This is a common mutation we see in some CBG lines that knocks out THCAS activity. Often found with homozygous P333R & S355N
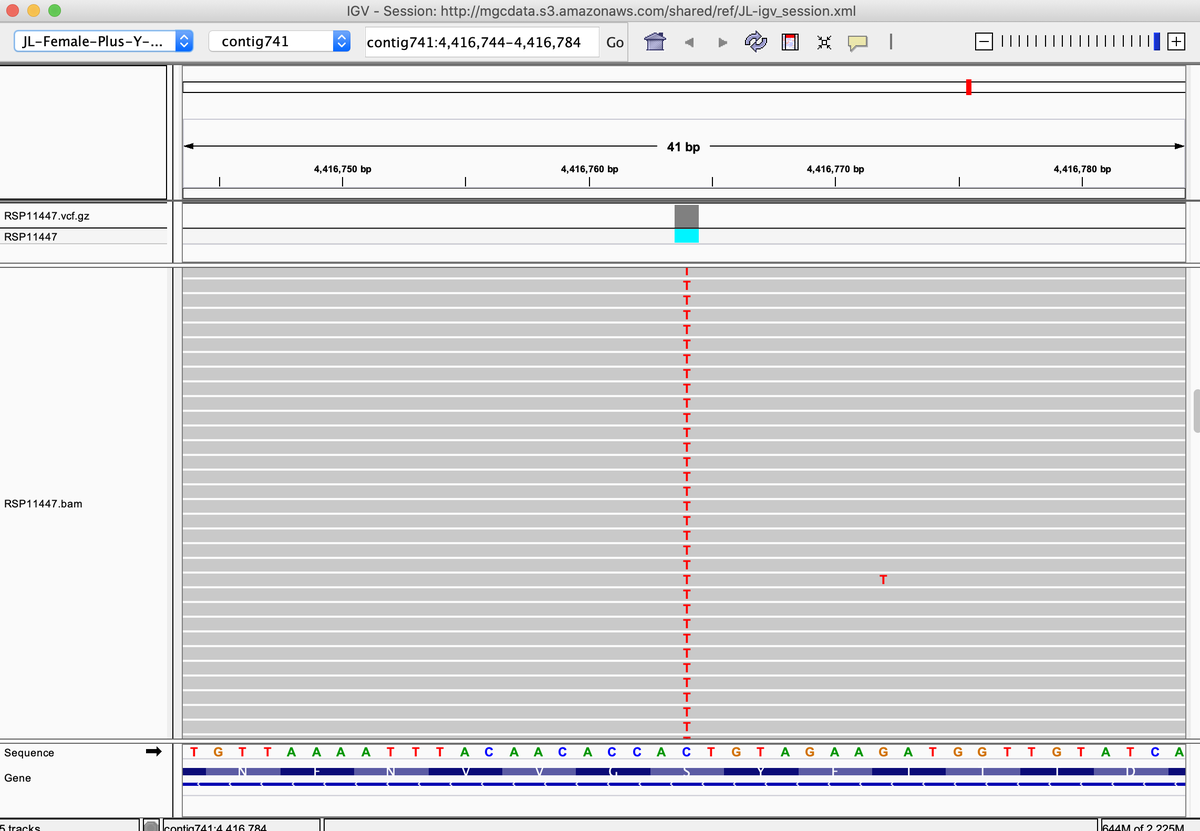
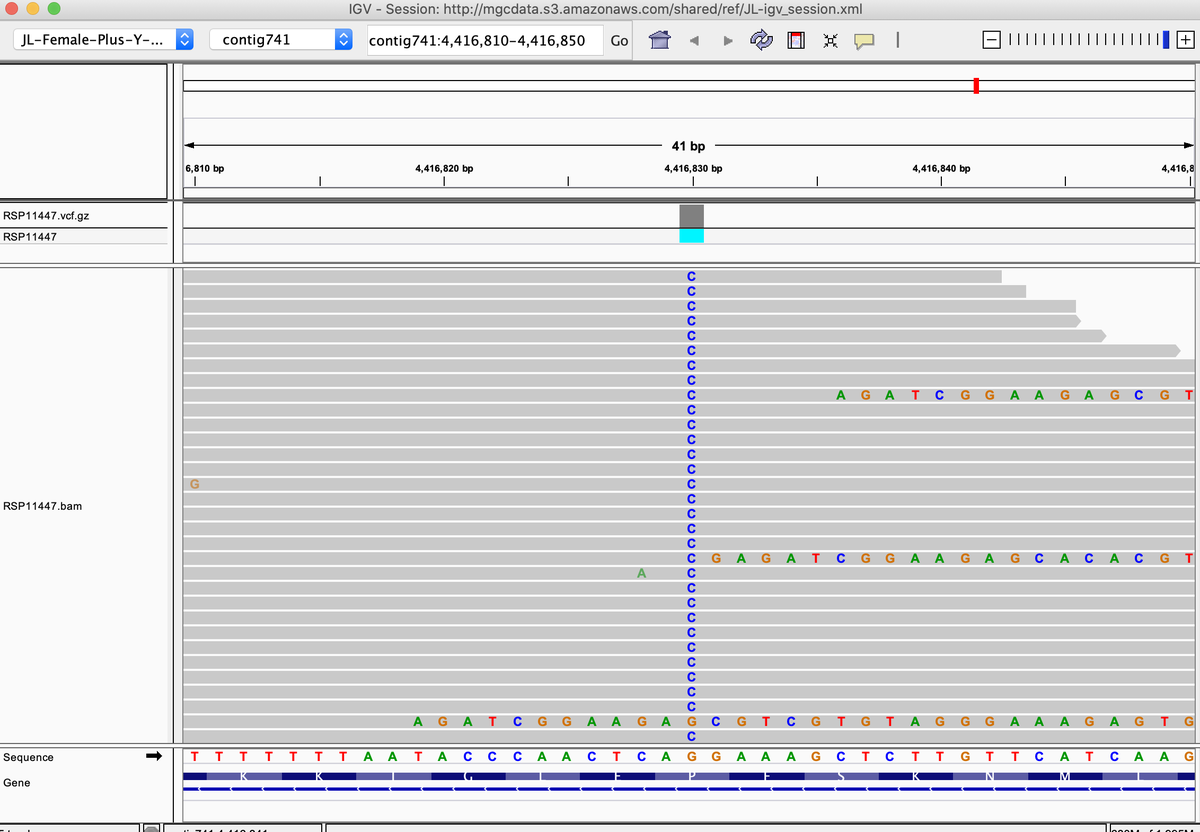
This can be seen in
kannapedia.net/strains/rsp114…
kannapedia.net/strains/rsp114…
There are 8 other CBG strains that have a THCAS gene with these 2 missense mutations. This is rare. Most CBG lines have both CBDAS and THCAS completely deleted. Convergent evolution. Many ways to skin a cat (please don't skin cats). 
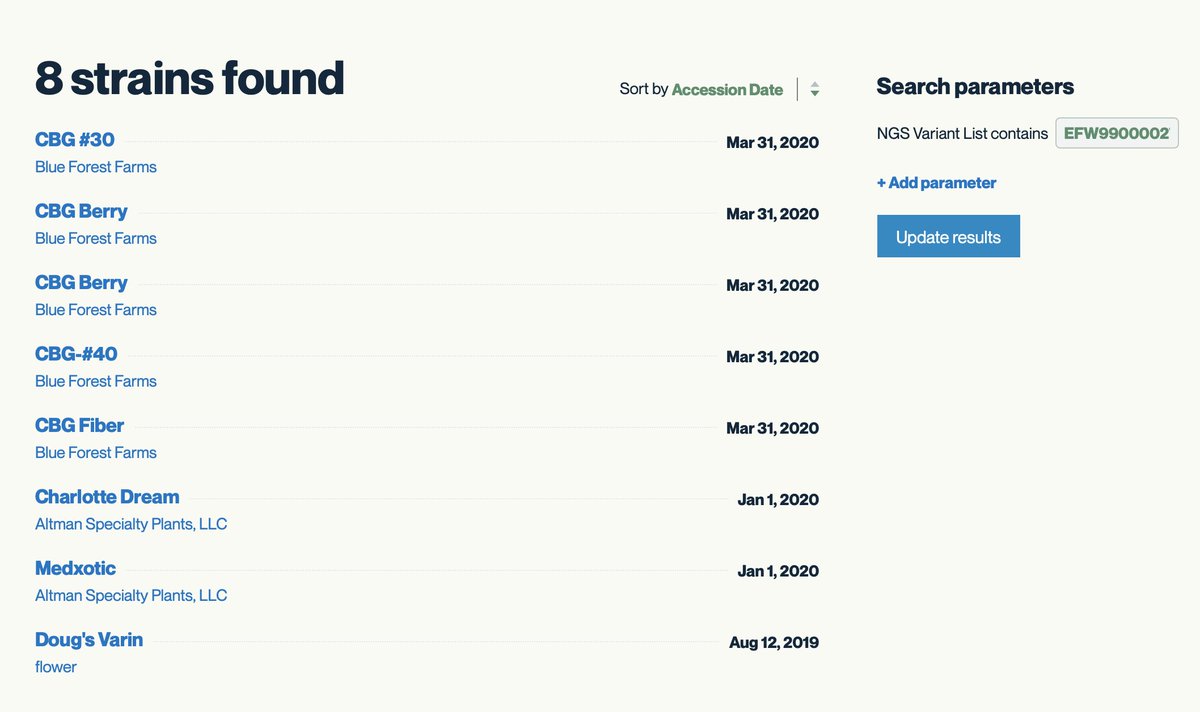
Of note is the THCV line known as Doug's Varin.
That's odd, as THCV production needs a functional THCAS. But alas, its heterozygous for P333R and S355N. It has a functional THCAS but also a bunk copy of THCAS from CBG lines.
kannapedia.net/strains/rsp112…

That's odd, as THCV production needs a functional THCAS. But alas, its heterozygous for P333R and S355N. It has a functional THCAS but also a bunk copy of THCAS from CBG lines.
kannapedia.net/strains/rsp112…
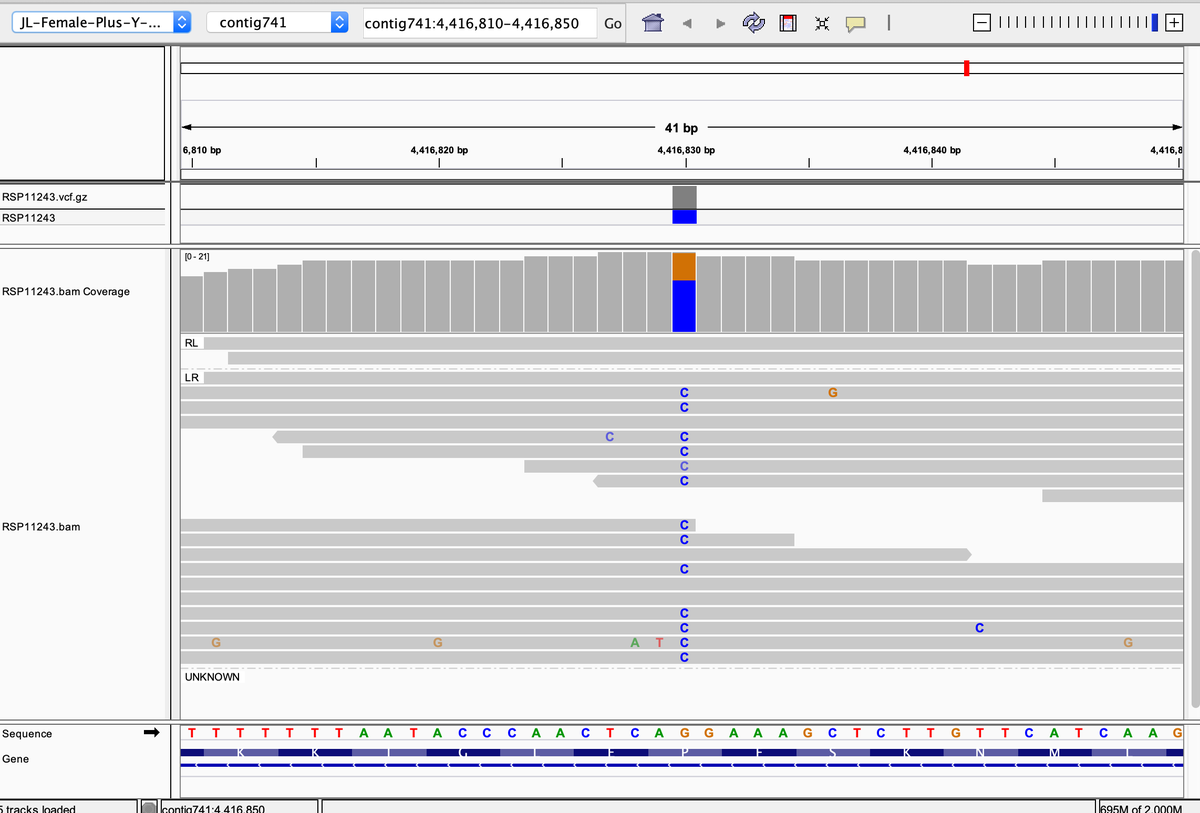
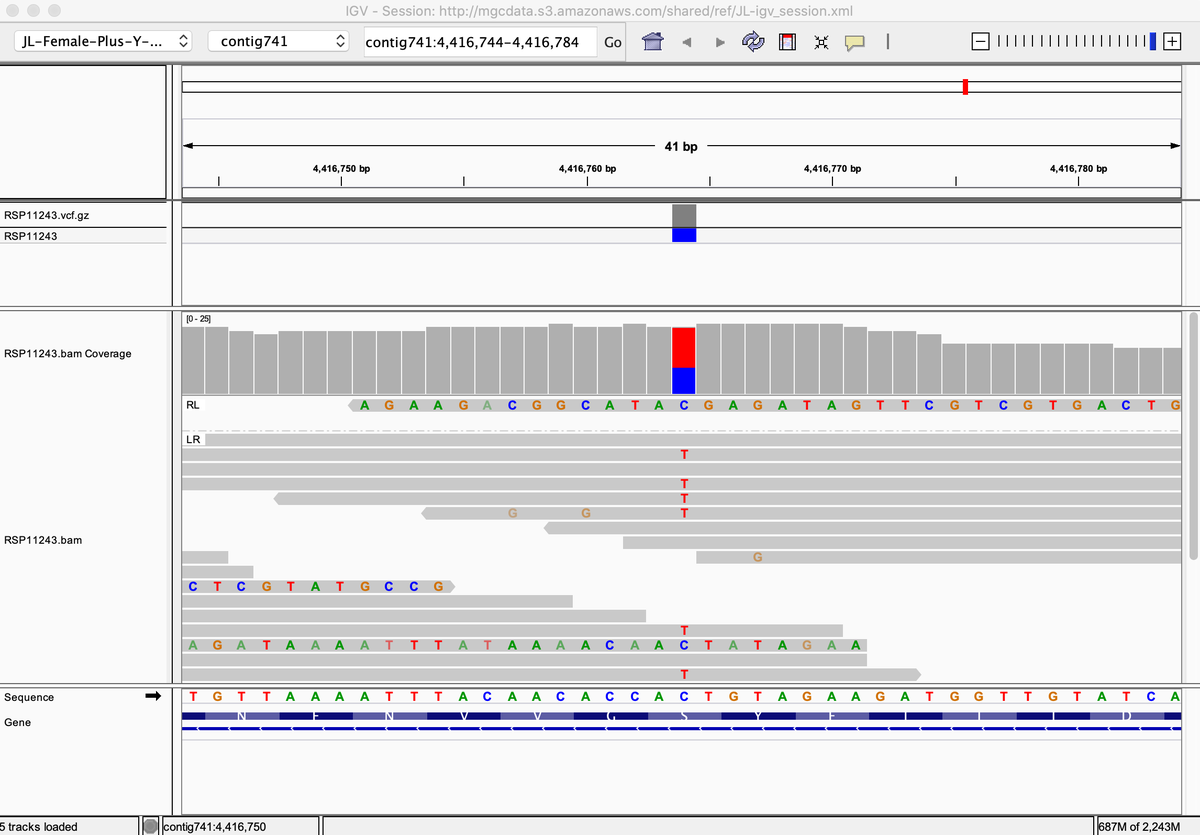
In summary, the latest Skunk to enter Kannapedia.net is unique compared to the others but the database needs more Skunks to really draw any conclusions on ancestry. Tracking the THCAS variants alone can be very helpful.
• • •
Missing some Tweet in this thread? You can try to
force a refresh