I like mice…
So does some lab that derived Omicron.
Very good read.
You’ll see how Molnupiravir could not have done this alone.
Random C->U mutations are not elevated in omicron but GC-AT are different in early omicron Vs post outbreak.
Mice host!
sciencedirect.com/science/articl…
So does some lab that derived Omicron.
Very good read.
You’ll see how Molnupiravir could not have done this alone.
Random C->U mutations are not elevated in omicron but GC-AT are different in early omicron Vs post outbreak.
Mice host!
sciencedirect.com/science/articl…

GC-AT mutations are a signature of ROS damage that makes 8-OxoG.
ROS is elevated in febrile viral infections.
These authors looked at early omicron Vs post outbreak omicron and see a difference here suggesting omicron had a different host at some point.
Mice fit the bill.
ROS is elevated in febrile viral infections.
These authors looked at early omicron Vs post outbreak omicron and see a difference here suggesting omicron had a different host at some point.
Mice fit the bill.
Another kind cat sent me this paper.
Another important point is the Ka/Ks analysis they do (dN/dS).
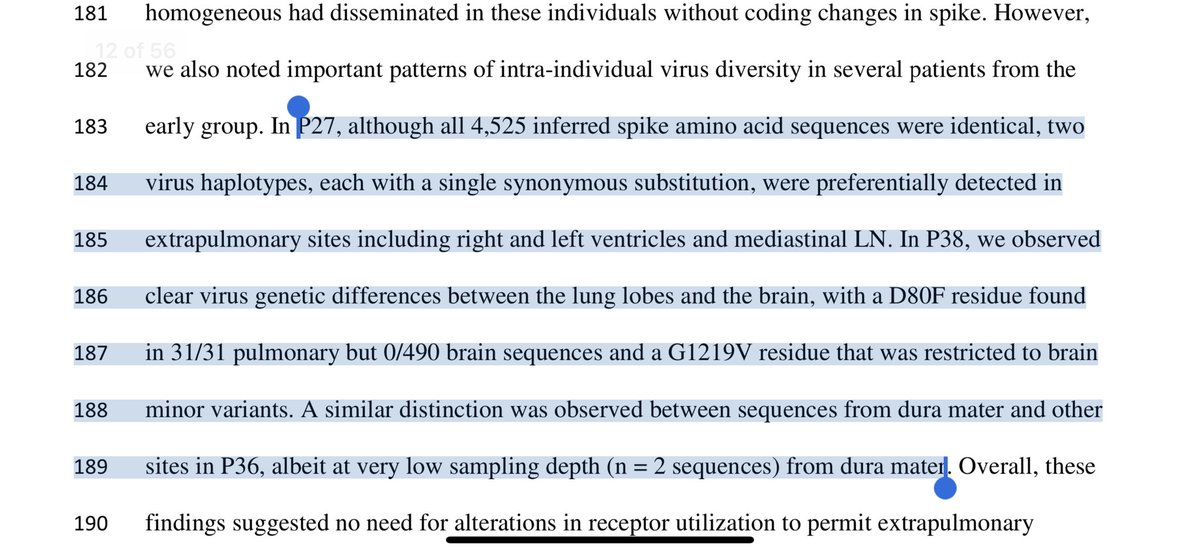
This shows 6.6X higher selective pressure on spike but not on the rest of the SARs genome.
Mice + selection against spike (vax of MAbs) could do this.
Lab derived.
Another important point is the Ka/Ks analysis they do (dN/dS).
This shows 6.6X higher selective pressure on spike but not on the rest of the SARs genome.
Mice + selection against spike (vax of MAbs) could do this.
Lab derived.
Just to clarify.
C-> U is the most common mutation in C19.
It’s also the mutation Molnupiravir makes.
So for Molnupiravir to be responsible for this, you would need more C->U than background C->U mutation rates….
And you need them focused on Spike.
And you need mis-sense SNPs.
C-> U is the most common mutation in C19.
It’s also the mutation Molnupiravir makes.
So for Molnupiravir to be responsible for this, you would need more C->U than background C->U mutation rates….
And you need them focused on Spike.
And you need mis-sense SNPs.
This isn’t an endorsement or Molnupiravir which is a trainwreck.
You do not approve a viral and human mutagen mid pandemic unless you want to fuel a leaky vaccine company with a perpetual booster gravy train.
You do not approve a viral and human mutagen mid pandemic unless you want to fuel a leaky vaccine company with a perpetual booster gravy train.
Some inquire about natural mice.
C19 needs Humanized AC2 mice aka lab mice.
C19 needs Humanized AC2 mice aka lab mice.
https://twitter.com/Kevin_McKernan/status/1477780480774782979
• • •
Missing some Tweet in this thread? You can try to
force a refresh