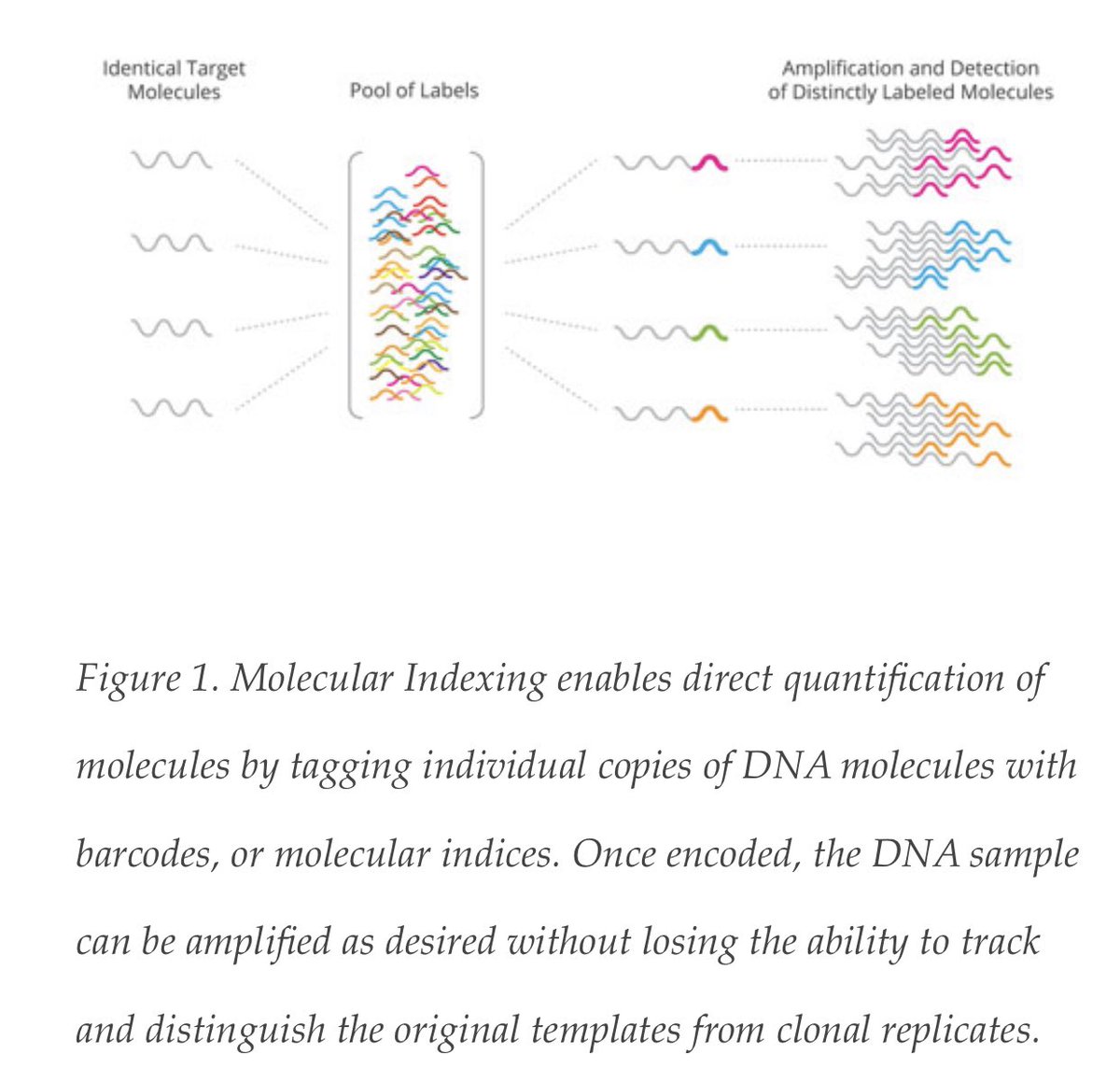
I’m in the process of updating this for submission (2,000 downloads before submission)
Several details people have brought up in support of this.
Will walk through them.
osf.io/bcsa6/
Several details people have brought up in support of this.
Will walk through them.
osf.io/bcsa6/
1)Some folks at NIH reached out pointed out the work on ribosomal pausing that we missed.
It’s very relevant.
Just the codon change in the FCS is enough to pause the ribosomes and alter 3D structure of spike.
Just 2 codons!
mdpi.com/1422-0067/22/1…
It’s very relevant.
Just the codon change in the FCS is enough to pause the ribosomes and alter 3D structure of spike.
Just 2 codons!
mdpi.com/1422-0067/22/1…
They changed many codons.
2)pseudouridine and N1-methyl should be better spelled out. Xia et al conflates the two and we will spell this out more.
PseudoU wobbles more the methylpseudoU but both significantly alter Tm and thus are ‘stickier’ bases than disrupt translation.
2)pseudouridine and N1-methyl should be better spelled out. Xia et al conflates the two and we will spell this out more.
PseudoU wobbles more the methylpseudoU but both significantly alter Tm and thus are ‘stickier’ bases than disrupt translation.
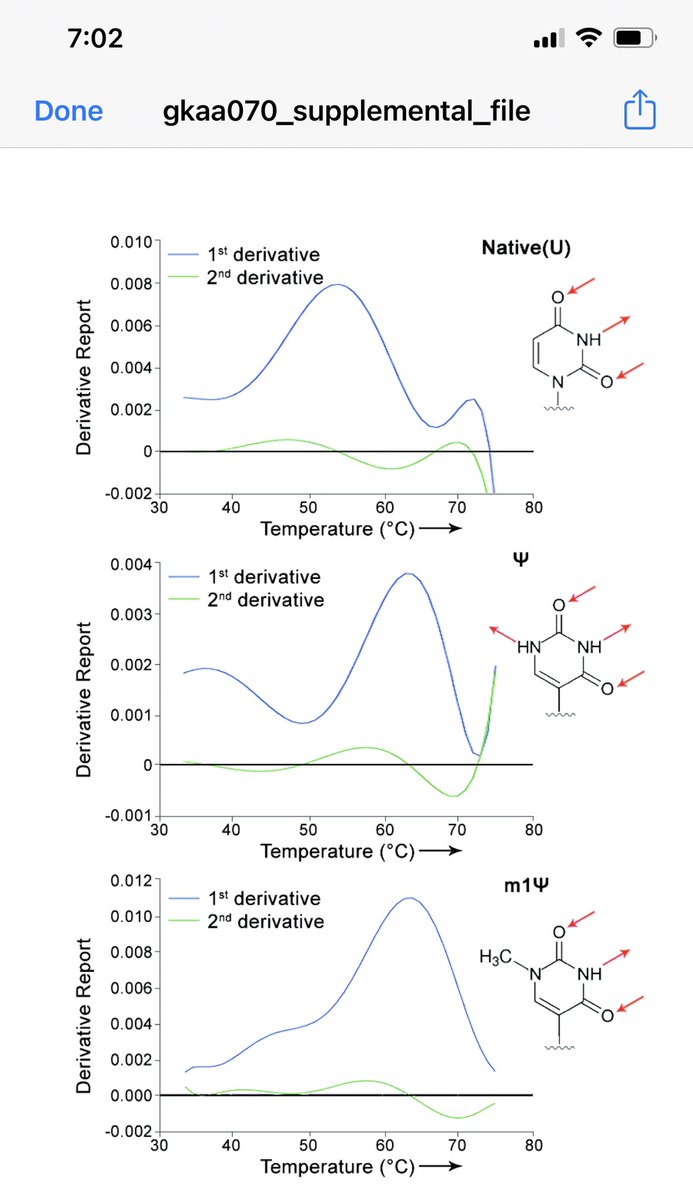
Parr et al.
Demonstrate 9C changes in Tm of a primer if you swap out just 4 Uridines with either base.
ncbi.nlm.nih.gov/labs/pmc/artic…
Demonstrate 9C changes in Tm of a primer if you swap out just 4 Uridines with either base.
ncbi.nlm.nih.gov/labs/pmc/artic…
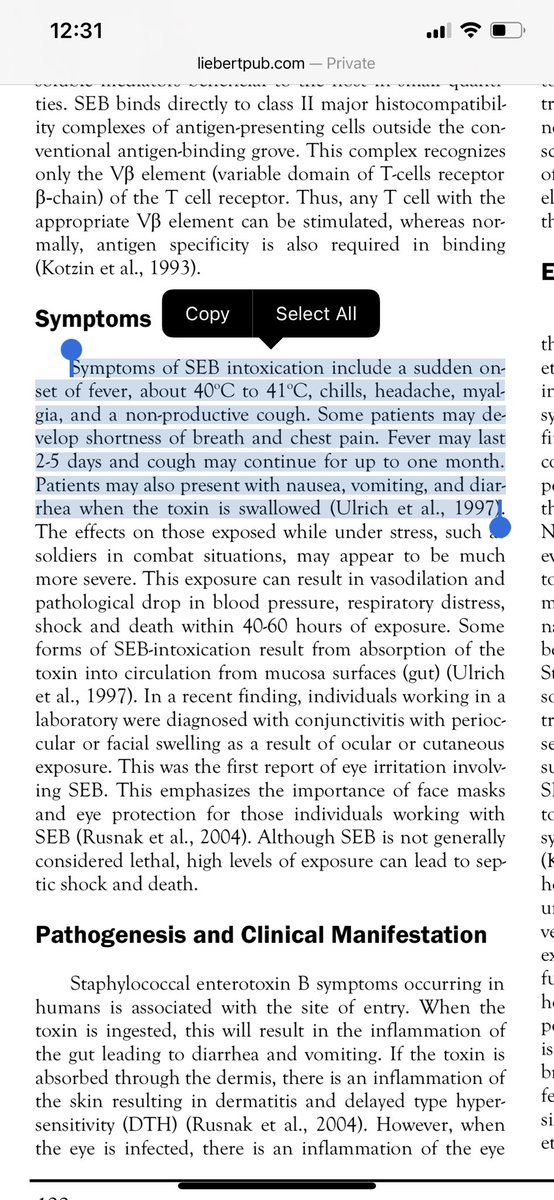
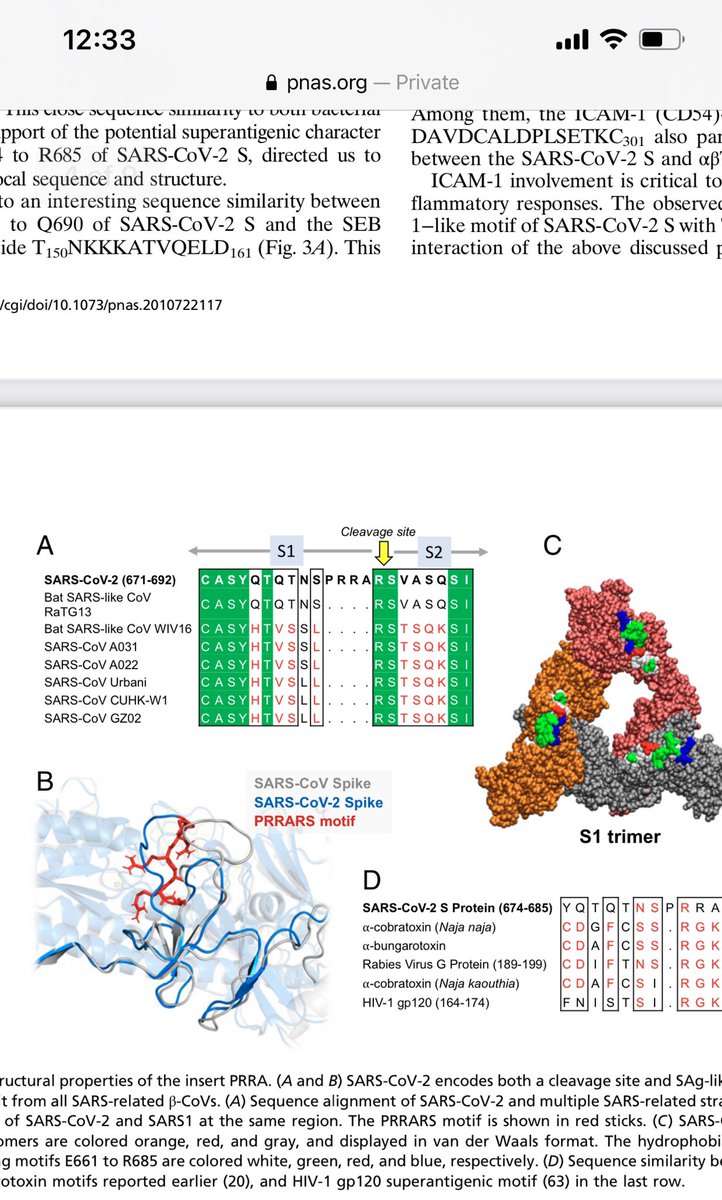
More data has emerged on SEB and its similarity to other venom toxins. The paper was already written to disclaimer SEB as being a superantigenic peptide that was a subset of the whole SEB toxin but Cheng et al raise an antibody to this and demonstrate its relevance.
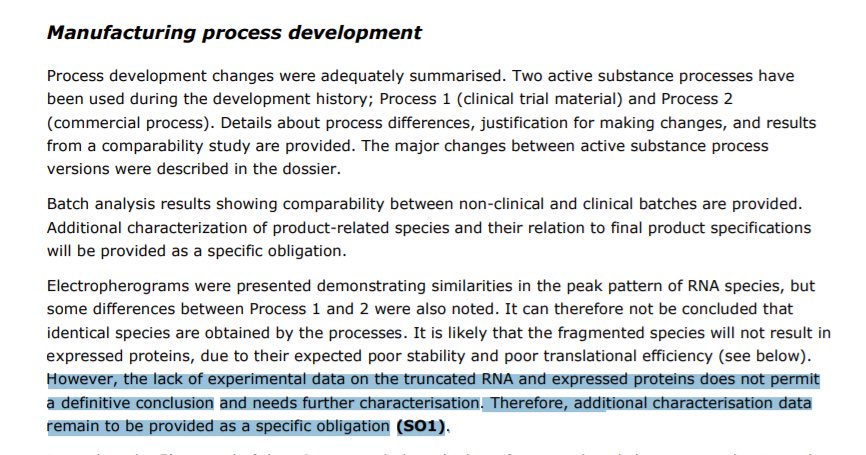
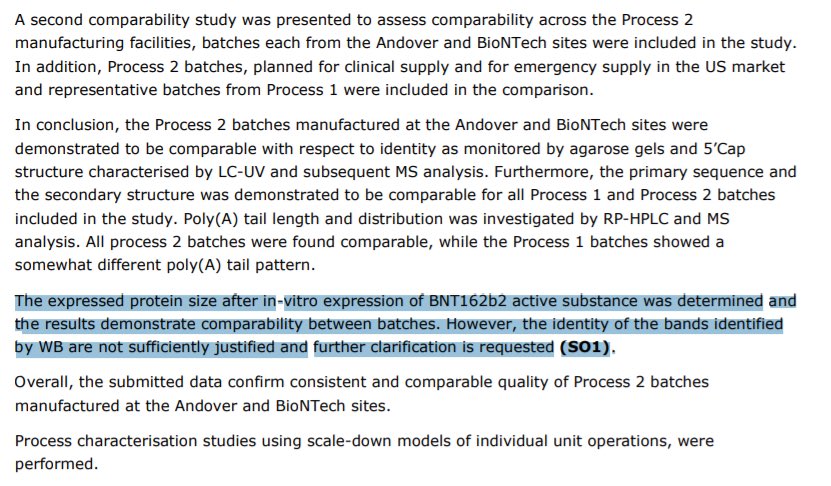
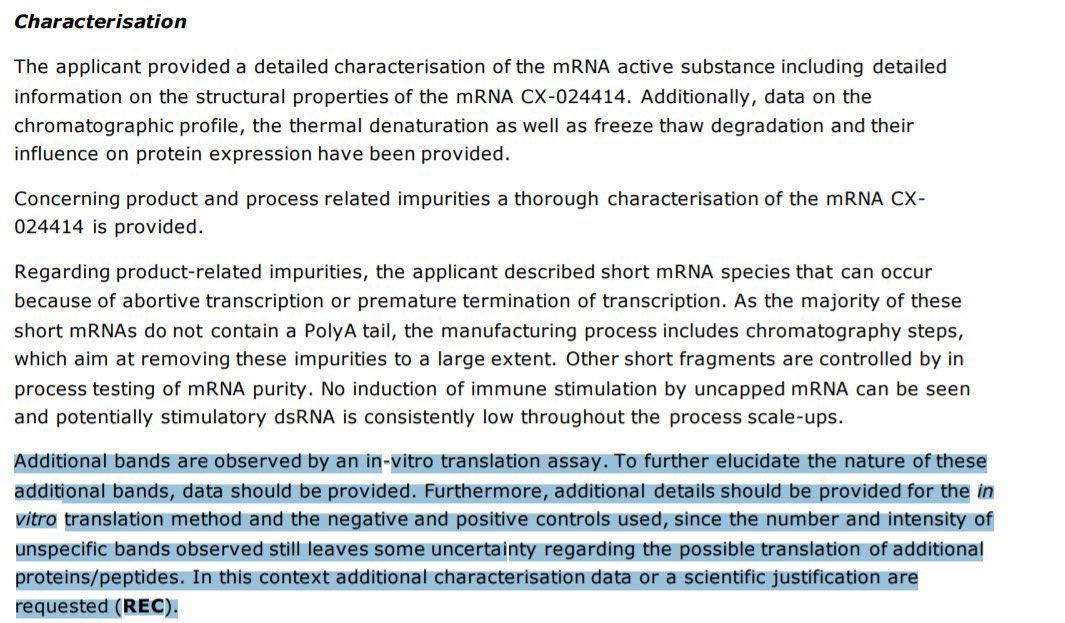
Perhaps the most damaging data to support our vax QC concerns comes out of the EMA.
Regulatory officials highlighted their post Translational smears demanding an action item to resolve them.
ema.europa.eu/en/documents/v…


Regulatory officials highlighted their post Translational smears demanding an action item to resolve them.
ema.europa.eu/en/documents/v…



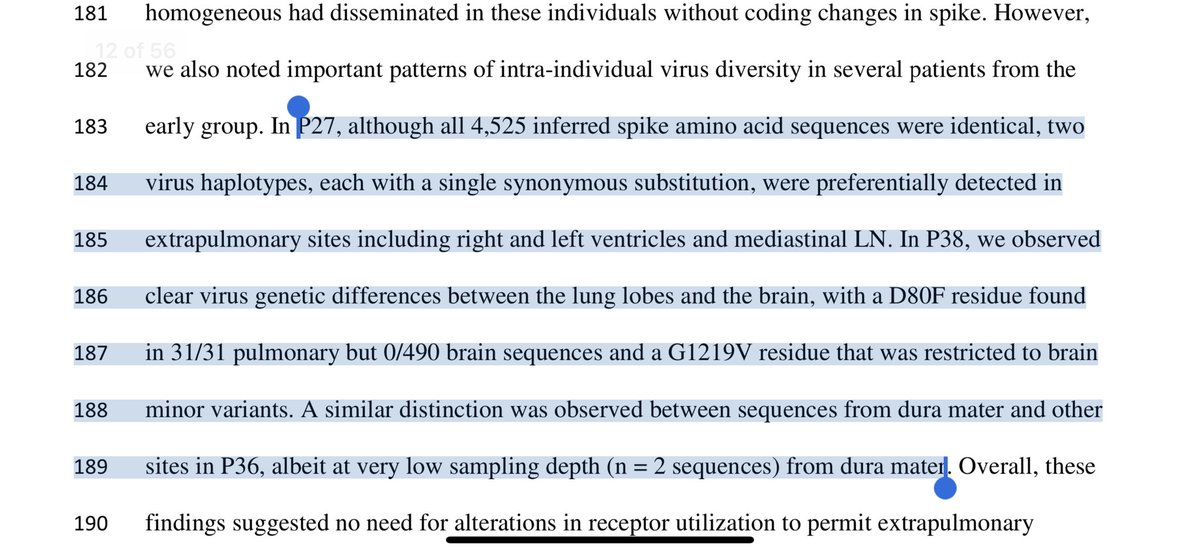
50 minutes into this video with @YoDoctorYo they notice S1 protein heterogeneity in the vaccinated.
Could this be differential glycosylation or translation error from methyl-pseudoU/codon pausing/stop codon ablation?
Could this be differential glycosylation or translation error from methyl-pseudoU/codon pausing/stop codon ablation?
This is pro-drug where the manufacturer has not provided evidence of the primary metabolite or active drug.
Lot to lot public QC is vacuous.
Liability waived
Unethical mandates
Lot to lot public QC is vacuous.
Liability waived
Unethical mandates
We have also added this reference regarding cancer risks.
Pfizer’s recent multi billion dollar acquisitions are for blood cancer companies and autoimmune disease.
Holding back their data for 50 years will give them key insights on what to make next
frontiersin.org/articles/10.33…
Pfizer’s recent multi billion dollar acquisitions are for blood cancer companies and autoimmune disease.
Holding back their data for 50 years will give them key insights on what to make next
frontiersin.org/articles/10.33…
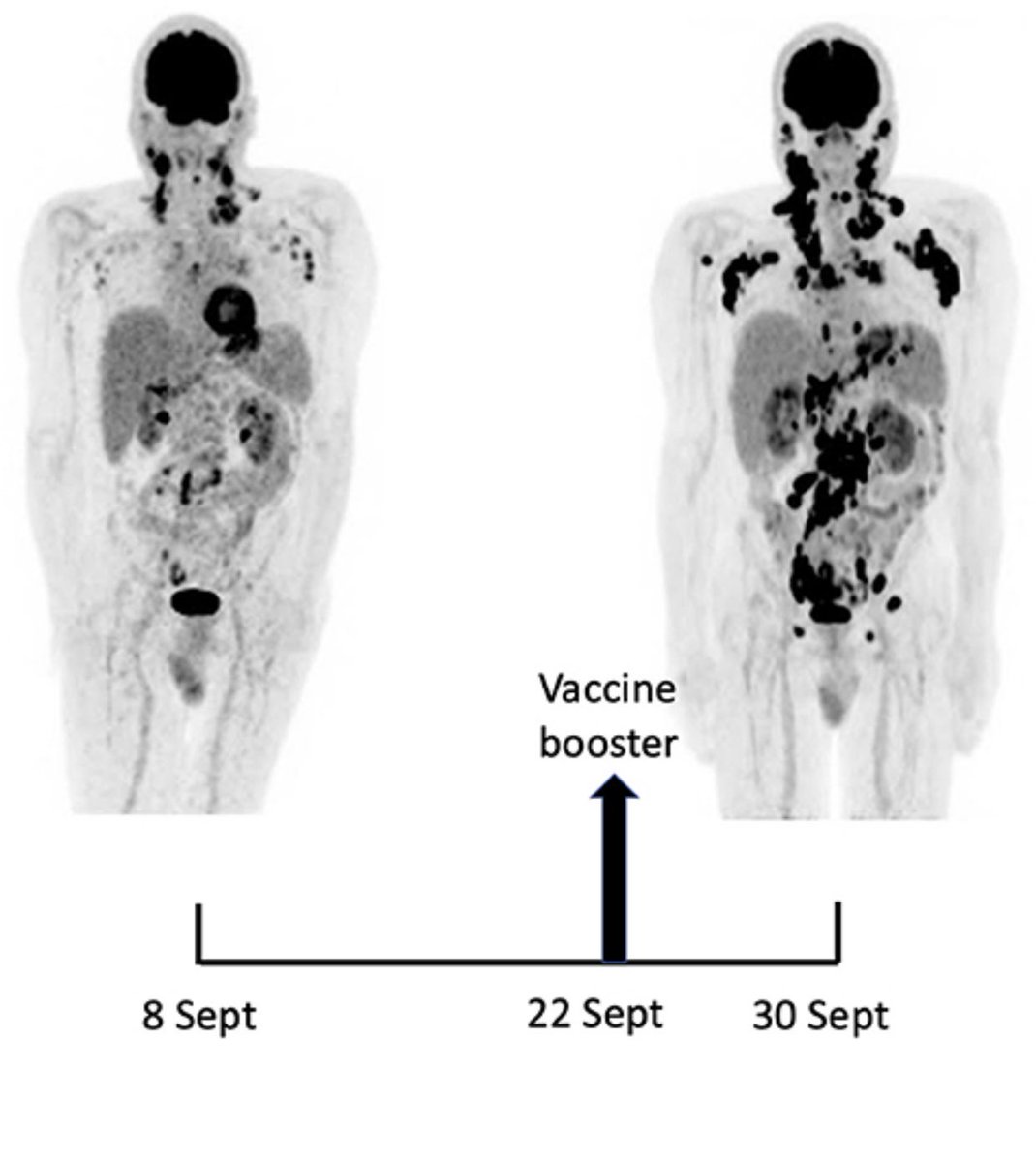
For more on codon optimizations…
• • •
Missing some Tweet in this thread? You can try to
force a refresh