There is a new paper out discussion MSH3 homology to this sequence.
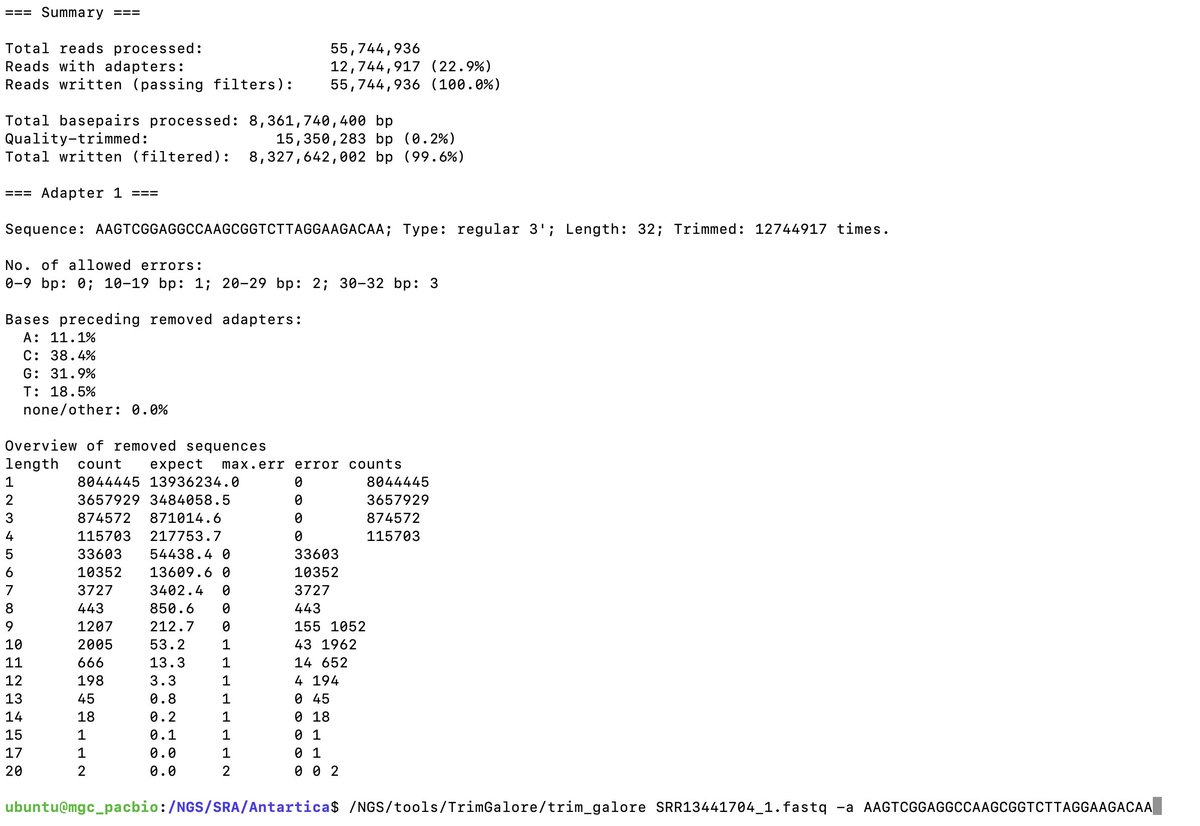
The tables in the supplement demonstrate the authors never blasted this sequence against 16S databases.
Had they done so, they probably wouldn’t have submitted that paper.
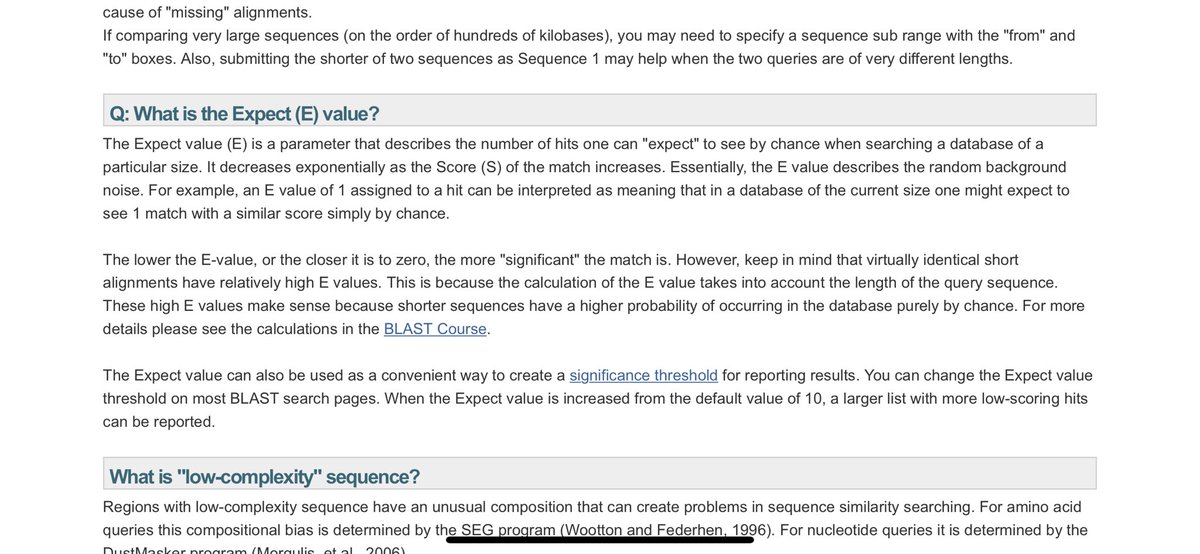
There is no E value to this sequence
The tables in the supplement demonstrate the authors never blasted this sequence against 16S databases.
Had they done so, they probably wouldn’t have submitted that paper.
There is no E value to this sequence
https://twitter.com/kevin_mckernan/status/1484987210508255233
Here is the MSH3 paper.
Note the supplement doesn’t look at 16S sequence. This is one of the most common sequences on Earth and this 19bp has a 10-11bp match to it. This destroys its E value.
It’s like finding water on Earth. Doesn’t 📌 you down.
frontiersin.org/articles/10.33…
Note the supplement doesn’t look at 16S sequence. This is one of the most common sequences on Earth and this 19bp has a 10-11bp match to it. This destroys its E value.
It’s like finding water on Earth. Doesn’t 📌 you down.
frontiersin.org/articles/10.33…
So Moderna doesn’t have any patent rights over this sequence.
• • •
Missing some Tweet in this thread? You can try to
force a refresh