Vax longevity in the blood is buried in one of my posts.
It deserves its own thread
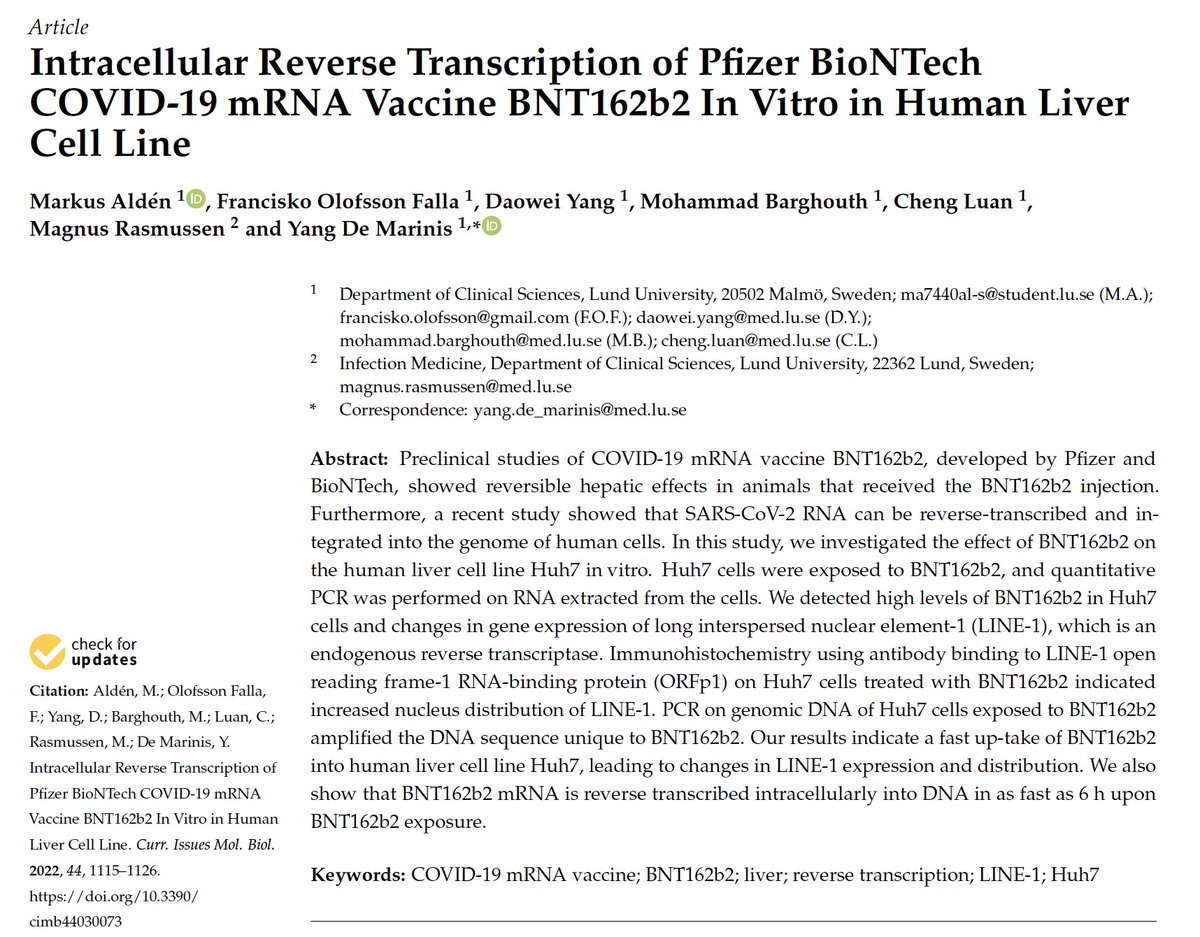
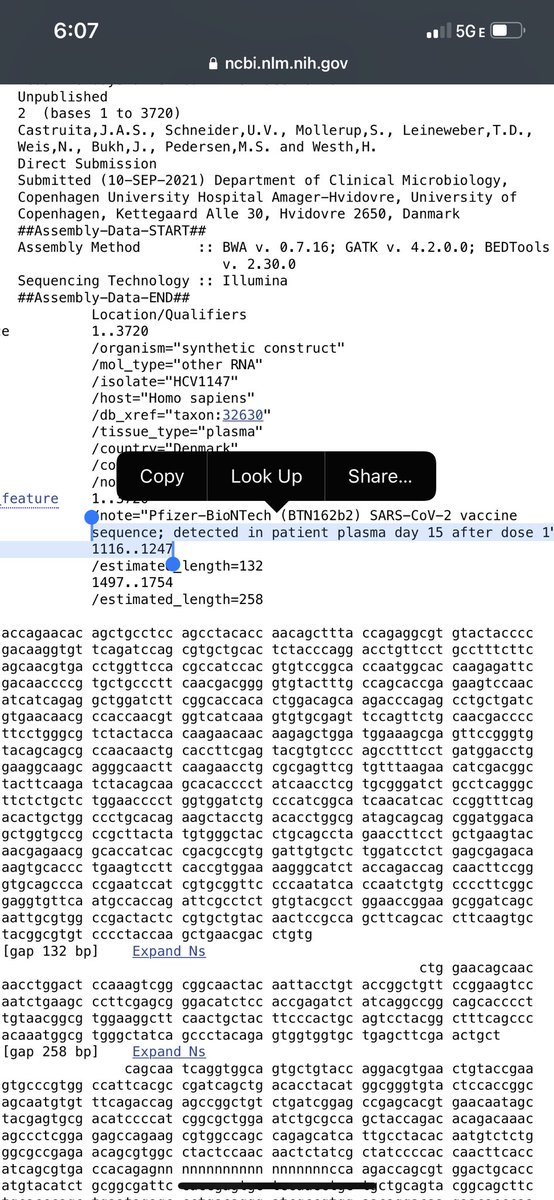
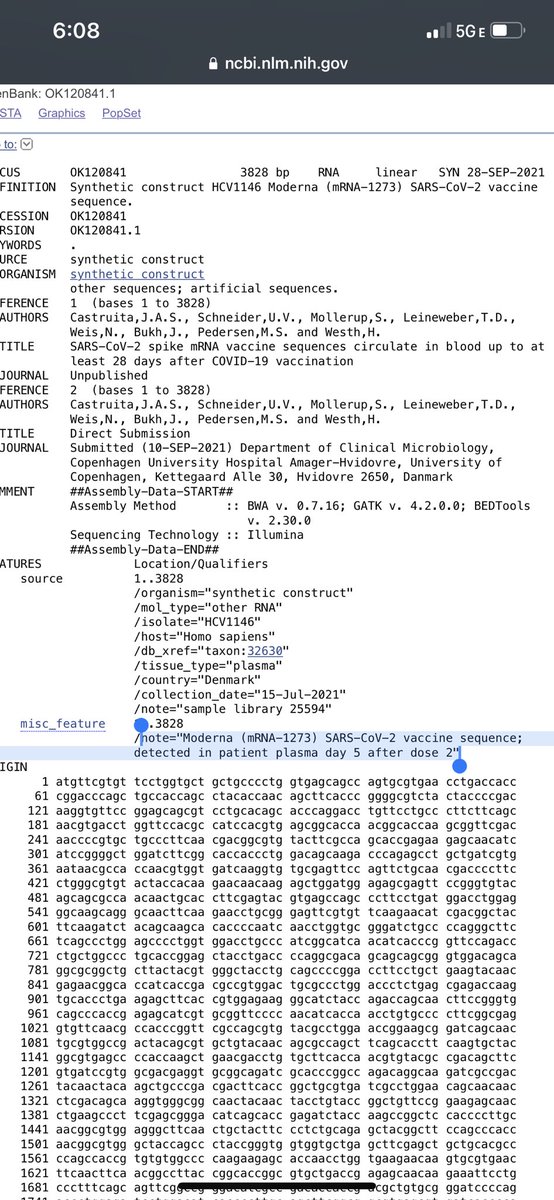
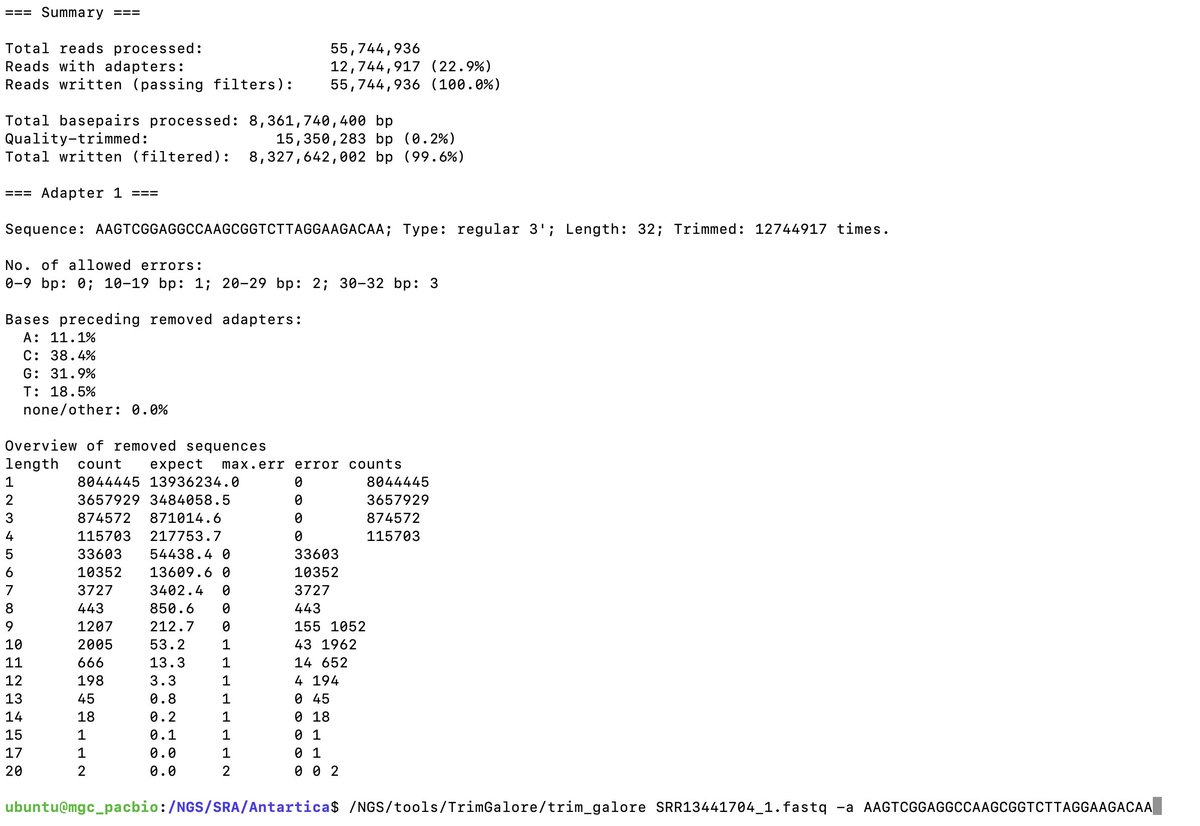
The title of this paper goes out to 28 days. Full seq is in NCBI.
It deserves its own thread
The title of this paper goes out to 28 days. Full seq is in NCBI.

Pfizer claimed it was gone in 24 hours.
Keep in mind normal C19 primers don’t amplify the vax due to the heavy codon optimization they did.
So no one is really looking for it.
Keep in mind normal C19 primers don’t amplify the vax due to the heavy codon optimization they did.
So no one is really looking for it.
When people do look… no one is listening.
To get full length sequencing of the virus, CDC asks for samples to be less than a CT of 28. So for this group to find it in plasma, there has to be a lot of it.
Keep in mind 80T of these 1um in length molecules can circle Earth.
Keep in mind 80T of these 1um in length molecules can circle Earth.
That’s about 13M molecules per UL if you have 6L of blood.
Your body is 87L is not much less if it distributes everywhere.
~1M copies/ul across your whole body.
Your body is 87L is not much less if it distributes everywhere.
~1M copies/ul across your whole body.
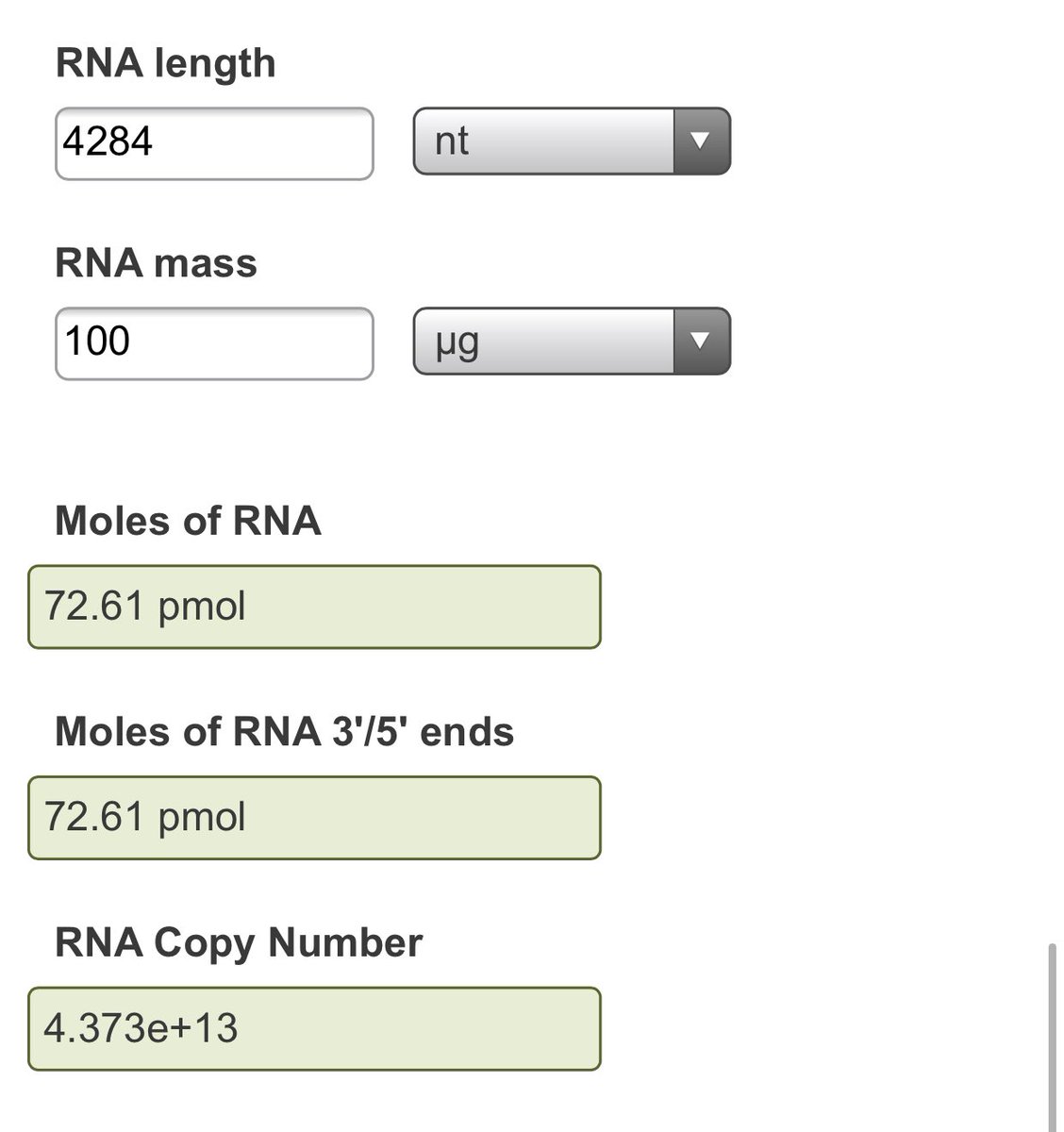
NEB calculator has 100ug at 43T molecules of a ~4.2kb ssRNA molecule.
2 shots= 86T
Booster even worse.
nebiocalculator.neb.com/#!/ssrnaamt
2 shots= 86T
Booster even worse.
nebiocalculator.neb.com/#!/ssrnaamt
Earth =~40,000 Km circumference
1million microns/meter
1billion microns/Km
1trillion microns/1,000 Km
40Trillion microns/circumference
Each Vax mRNA = 1um.
Persistence length of mRNA =~4200bp/micron.
1million microns/meter
1billion microns/Km
1trillion microns/1,000 Km
40Trillion microns/circumference
Each Vax mRNA = 1um.
Persistence length of mRNA =~4200bp/micron.
This study isn’t published yet so we don’t know if they stopped measuring at 28 days? Likewise, certain tissues may have higher concentrations than others.
Other studies show longer duration in GC/Lymphnodes.
Other studies show longer duration in GC/Lymphnodes.
https://twitter.com/BrendanEich/status/1494045047049768960
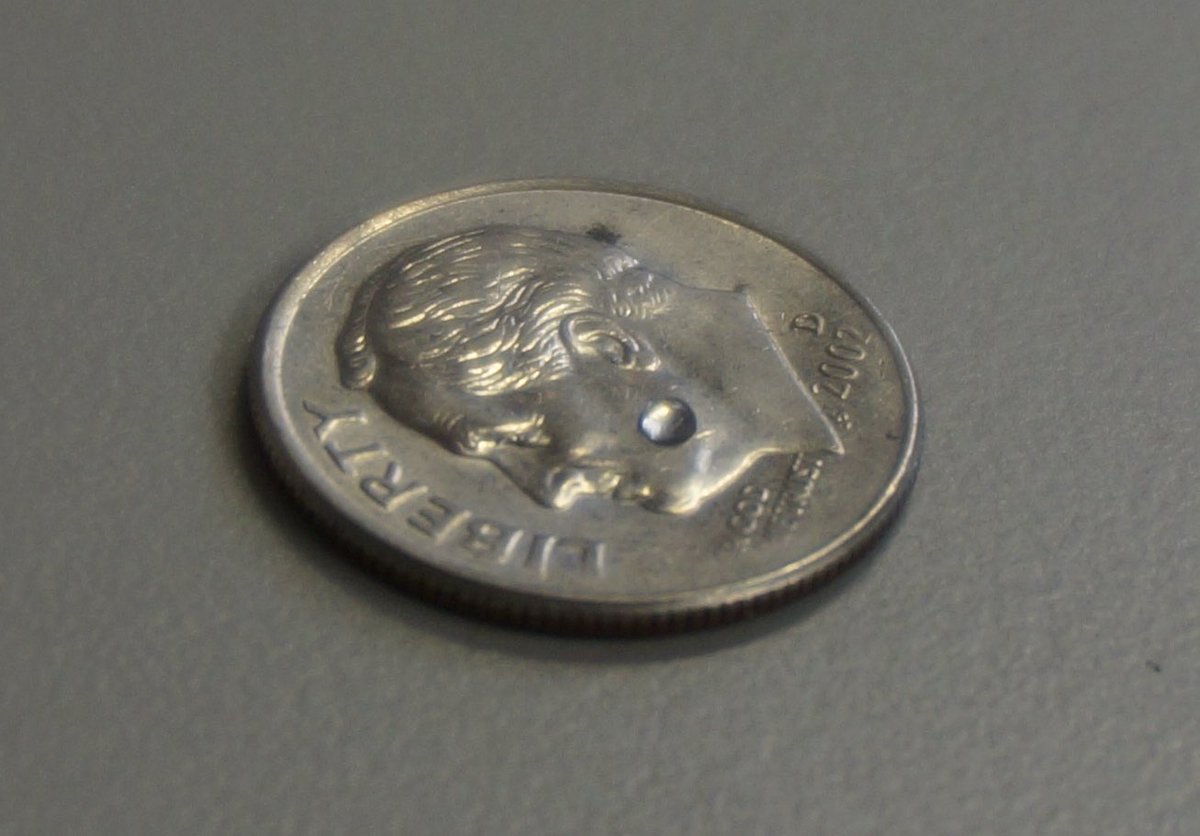
For those not used to working with microliters…
That spot on his cheek is a microliter.
The shot will give you ~1Million copies of the mRNA in every microliter of your body assuming a uniform distribution to your 87L body.
That spot on his cheek is a microliter.
The shot will give you ~1Million copies of the mRNA in every microliter of your body assuming a uniform distribution to your 87L body.

43Trillion mRNAs per 100ug shot.
87million microliters per 87L.
~500,000 copies/ul
2 shots = 1M/ul
87million microliters per 87L.
~500,000 copies/ul
2 shots = 1M/ul
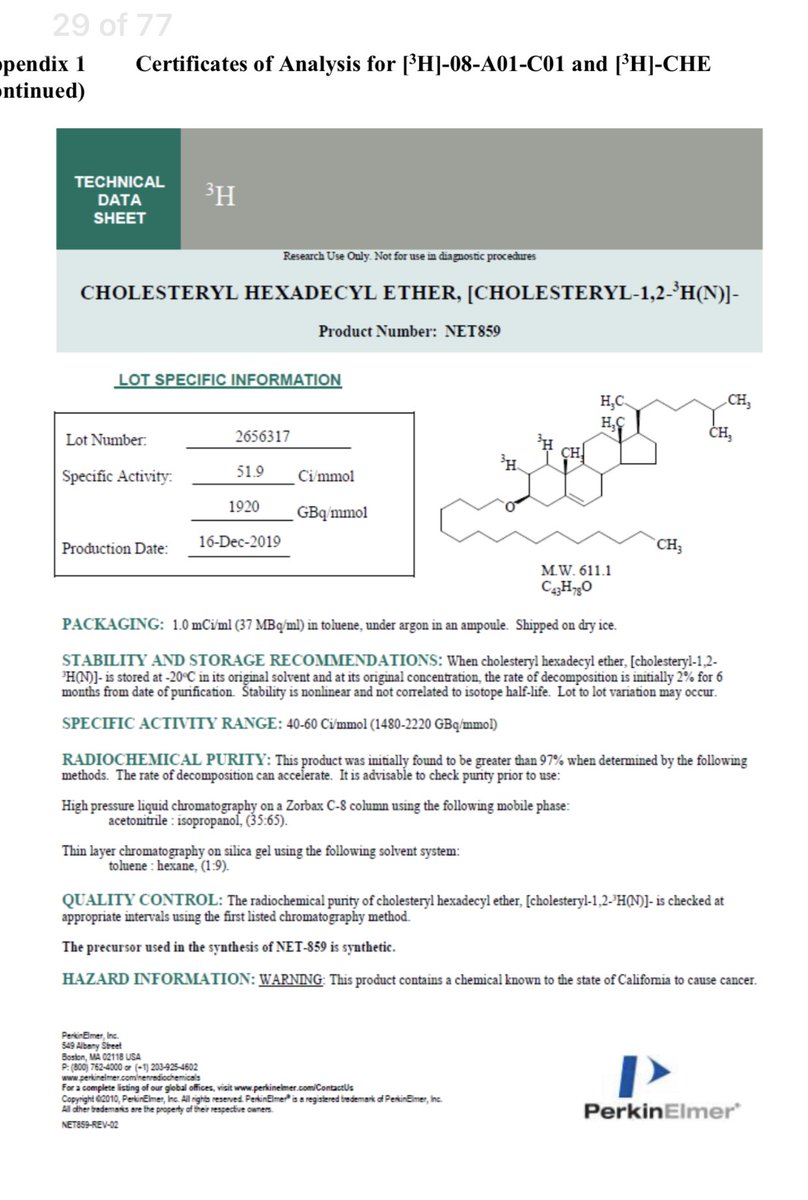
The Pfizer docs have a longevity study.
They didn’t radiolabel the mRNA.
They radiolabelled cholesterol as a proxy for the LNP bio distribution.
They didn’t radiolabel the mRNA.
They radiolabelled cholesterol as a proxy for the LNP bio distribution.

• • •
Missing some Tweet in this thread? You can try to
force a refresh