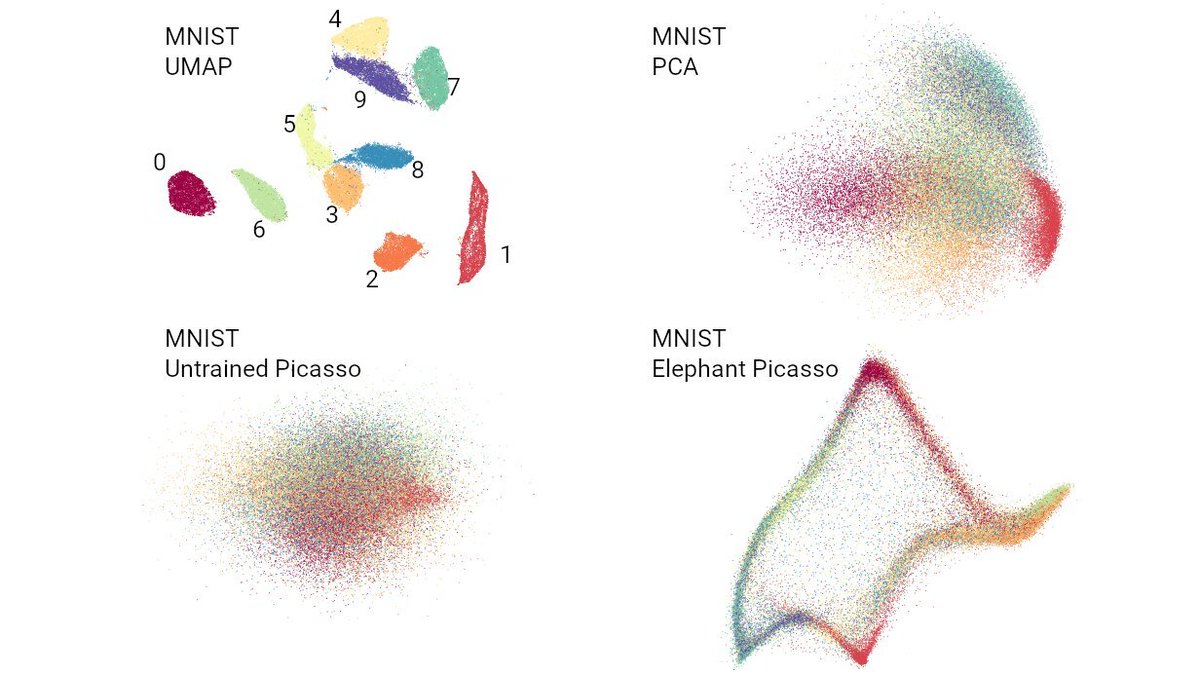
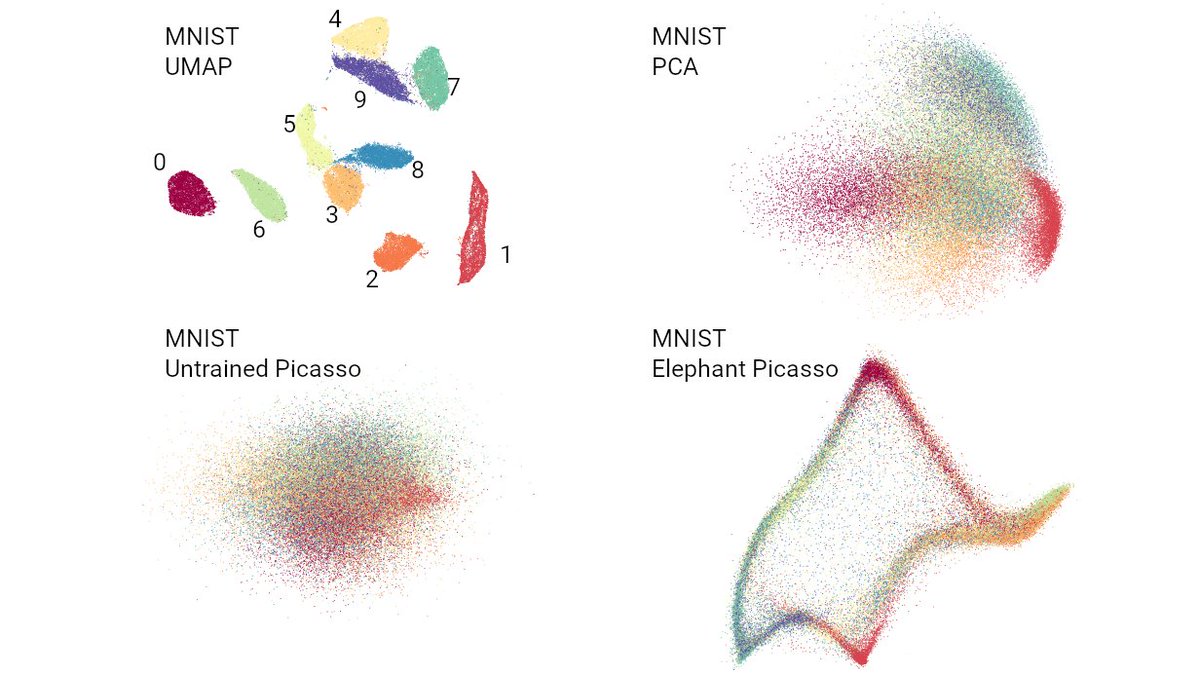
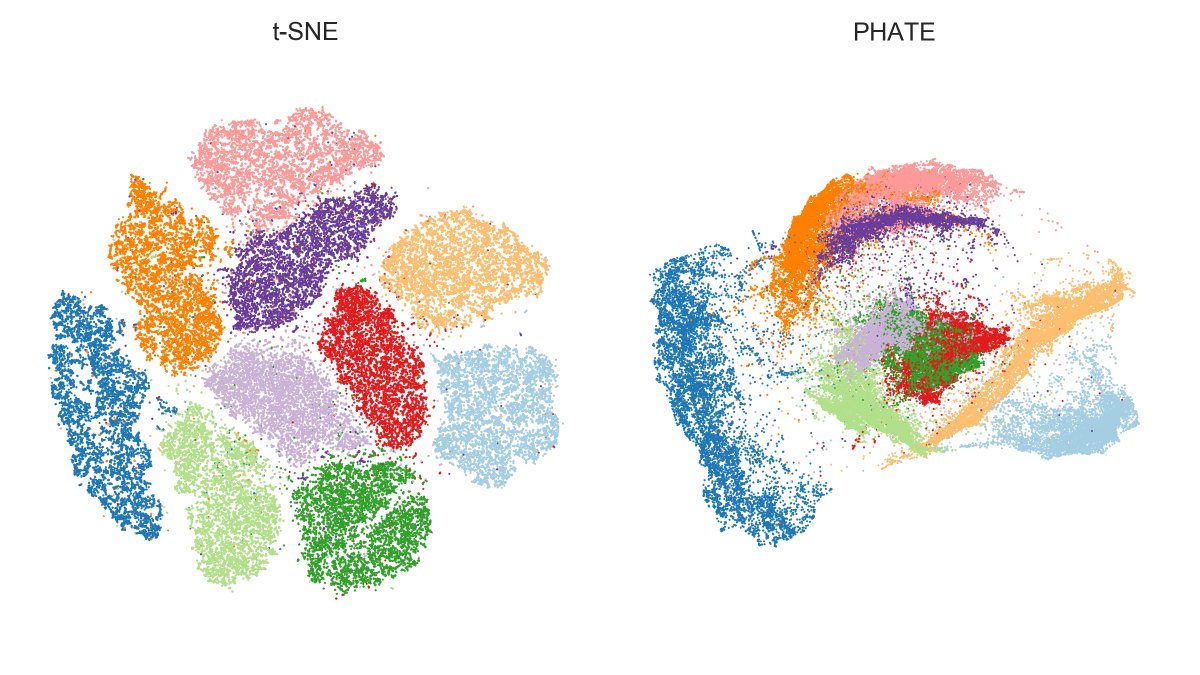
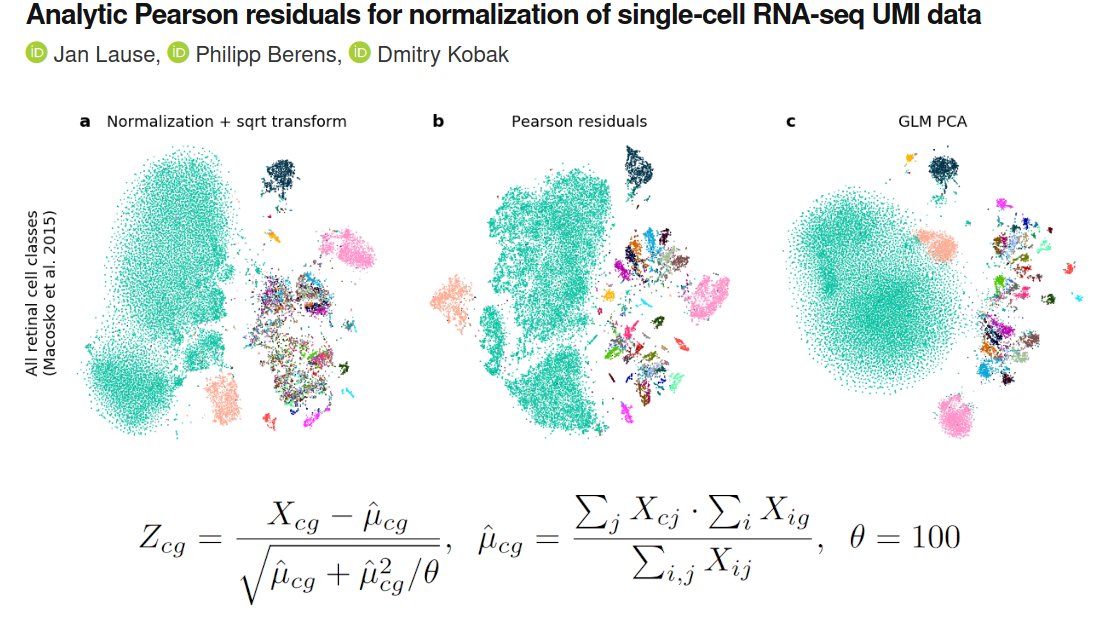
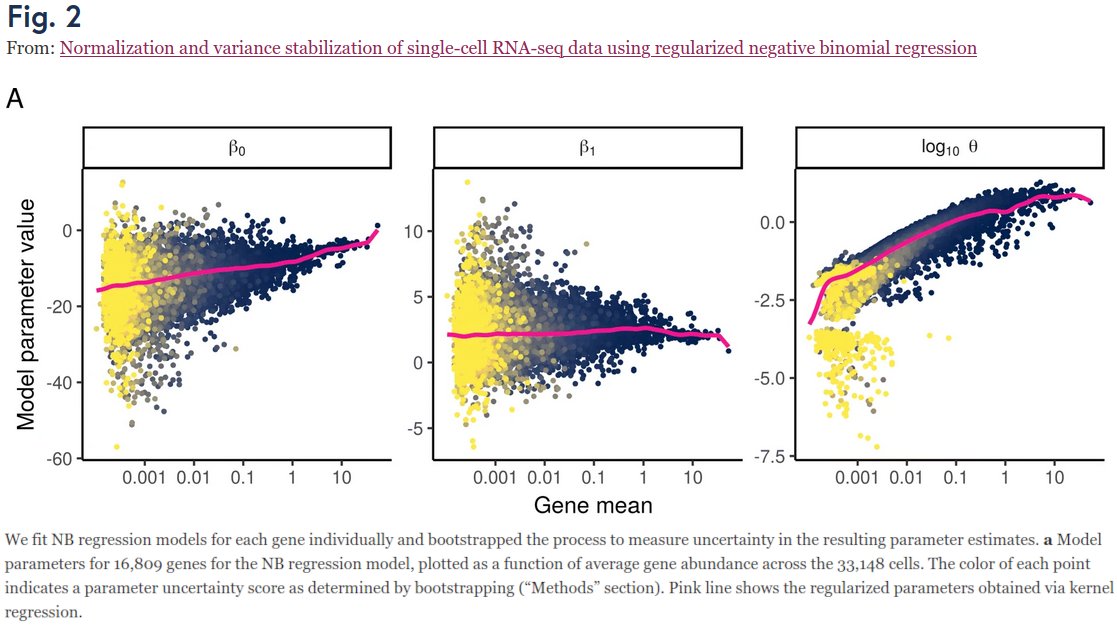
Want to know which countries fake their daily reported numbers of Covid19 cases and deaths? 🧐
I tested each country for underdispersion relative to the Poisson model: github.com/dkobak/excess-…. If the variance is lower than the Poisson variance, something is fishy (pun intended).
I tested each country for underdispersion relative to the Poisson model: github.com/dkobak/excess-…. If the variance is lower than the Poisson variance, something is fishy (pun intended).

Updated my analysis quantifying underdispersion (relative to Poisson 🐟) in daily reported Covid-19 cases and deaths. Countries listed below have likely been tampering with their data :-/
Manuscript upcoming.
Code moved here: github.com/dkobak/covid-u…
Manuscript upcoming.
Code moved here: github.com/dkobak/covid-u…
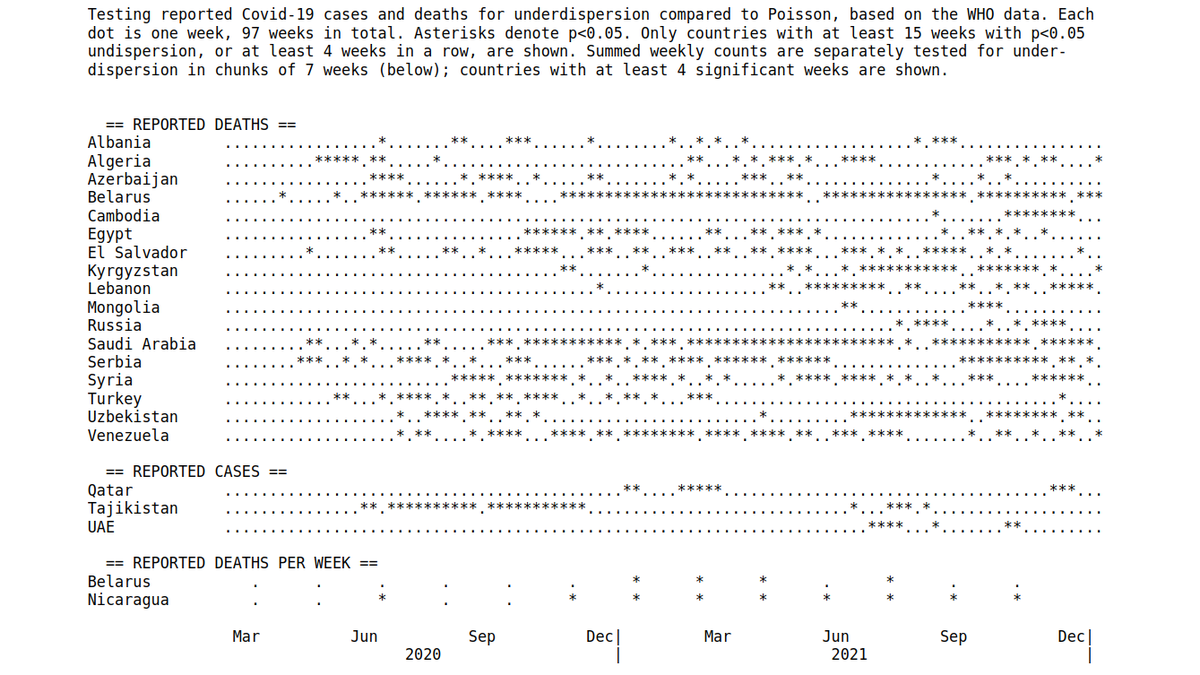
Update: preprint is up. The claim is that Poisson underdispersion in daily reported Covid-19 cases/deaths is a HUGE RED FLAG and suggests misreporting.
Underdispersion ratio (mean/variance) is correlated with undercount ratio based on excess mortality.
medrxiv.org/content/10.110…
Underdispersion ratio (mean/variance) is correlated with undercount ratio based on excess mortality.
medrxiv.org/content/10.110…
• • •
Missing some Tweet in this thread? You can try to
force a refresh