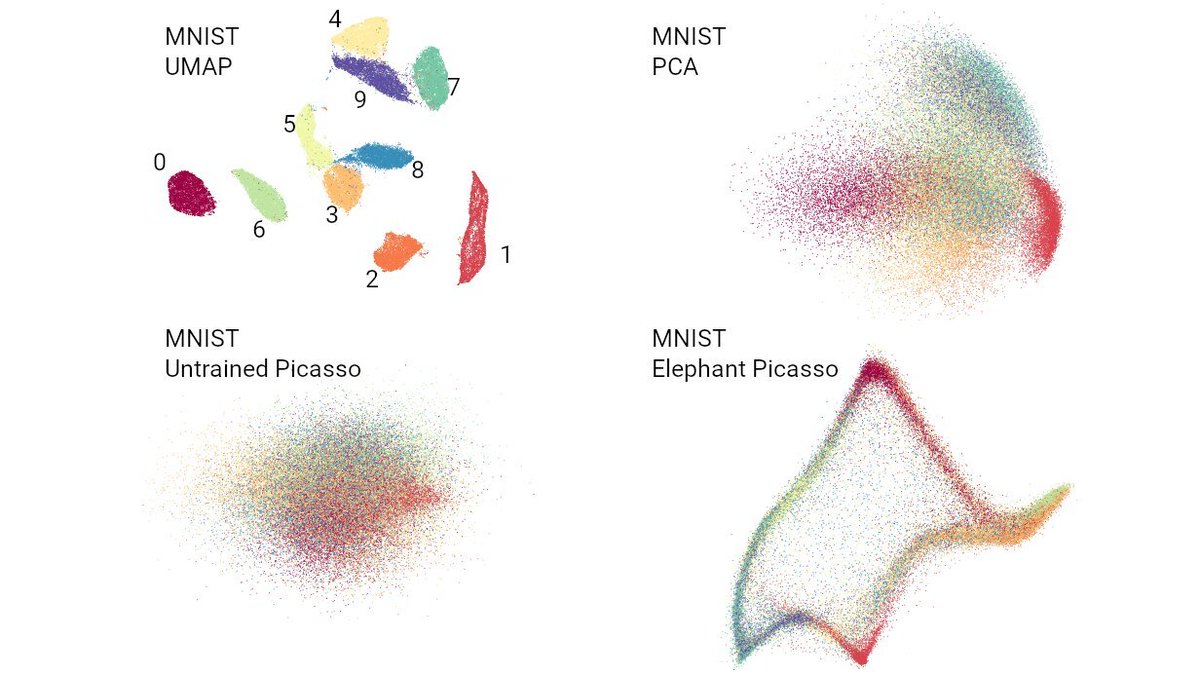
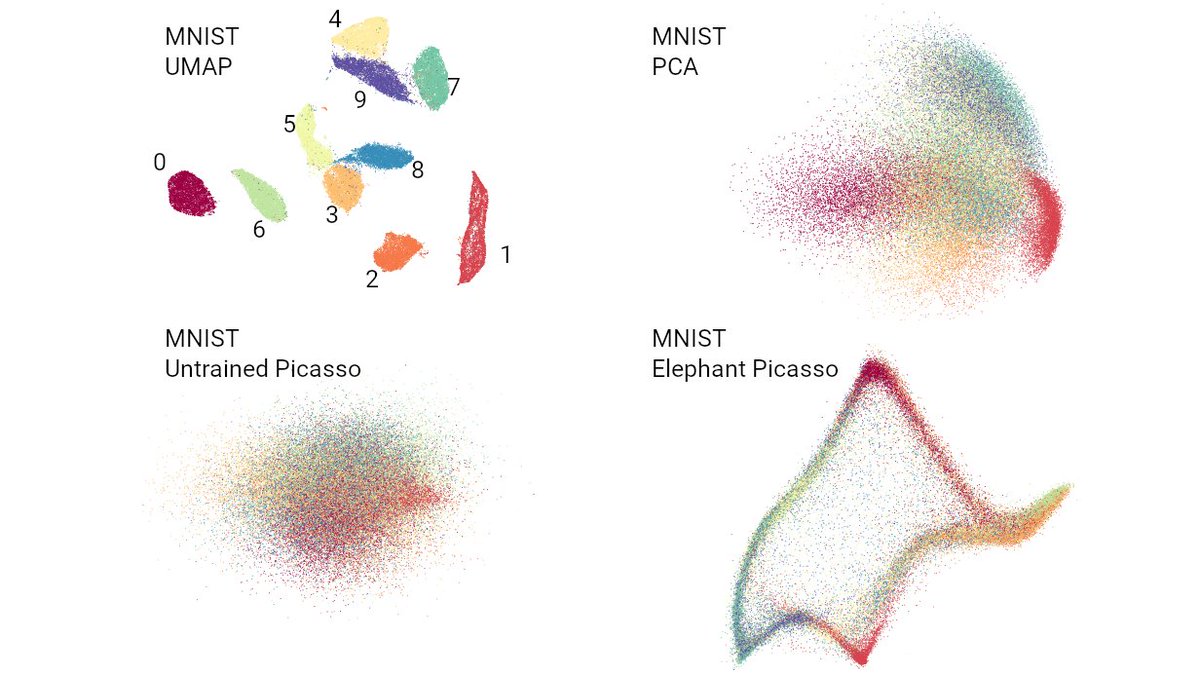
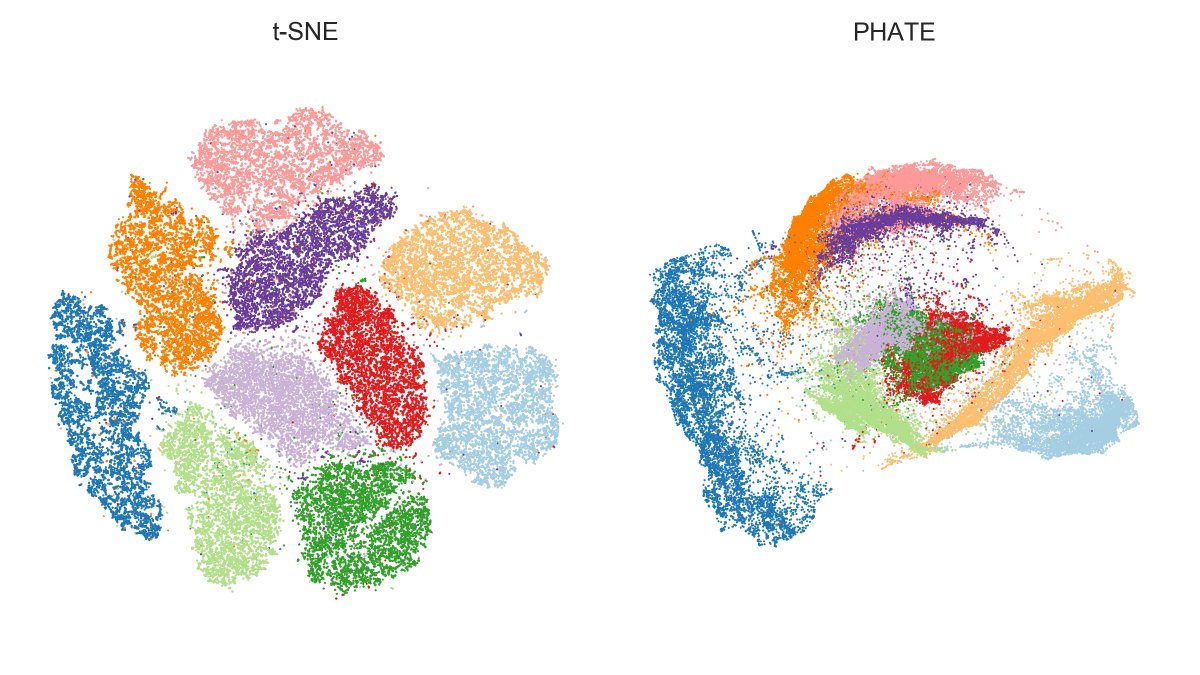
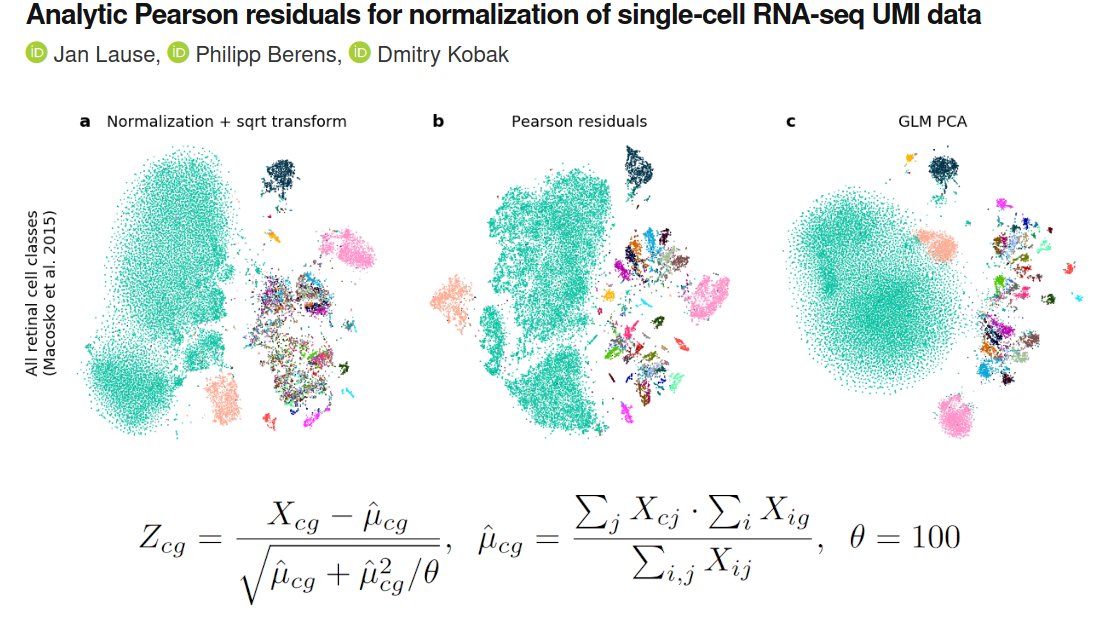
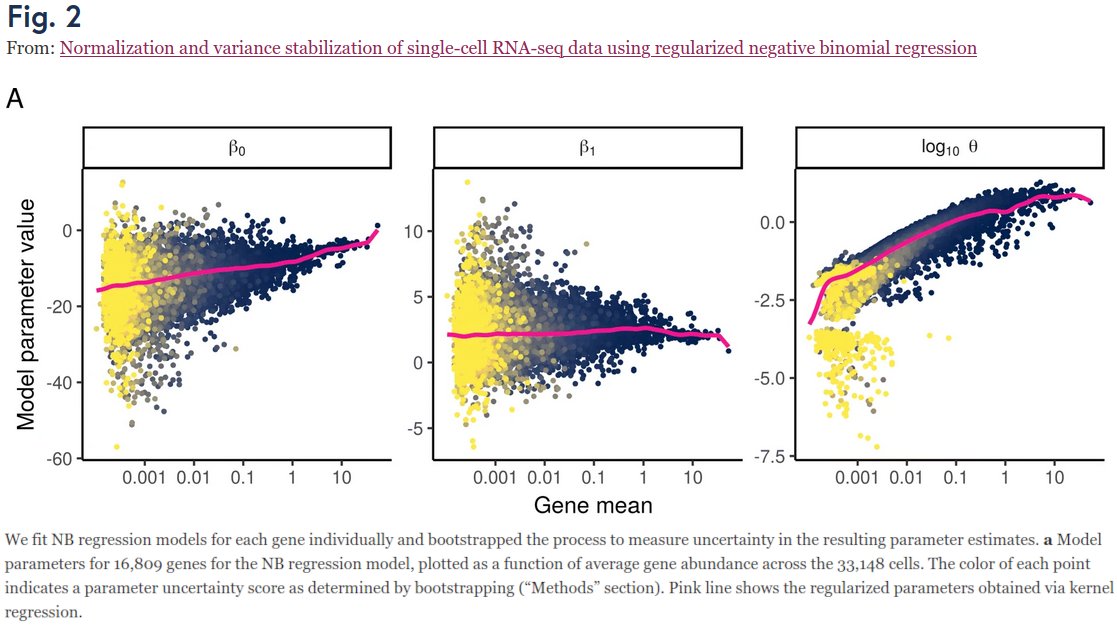
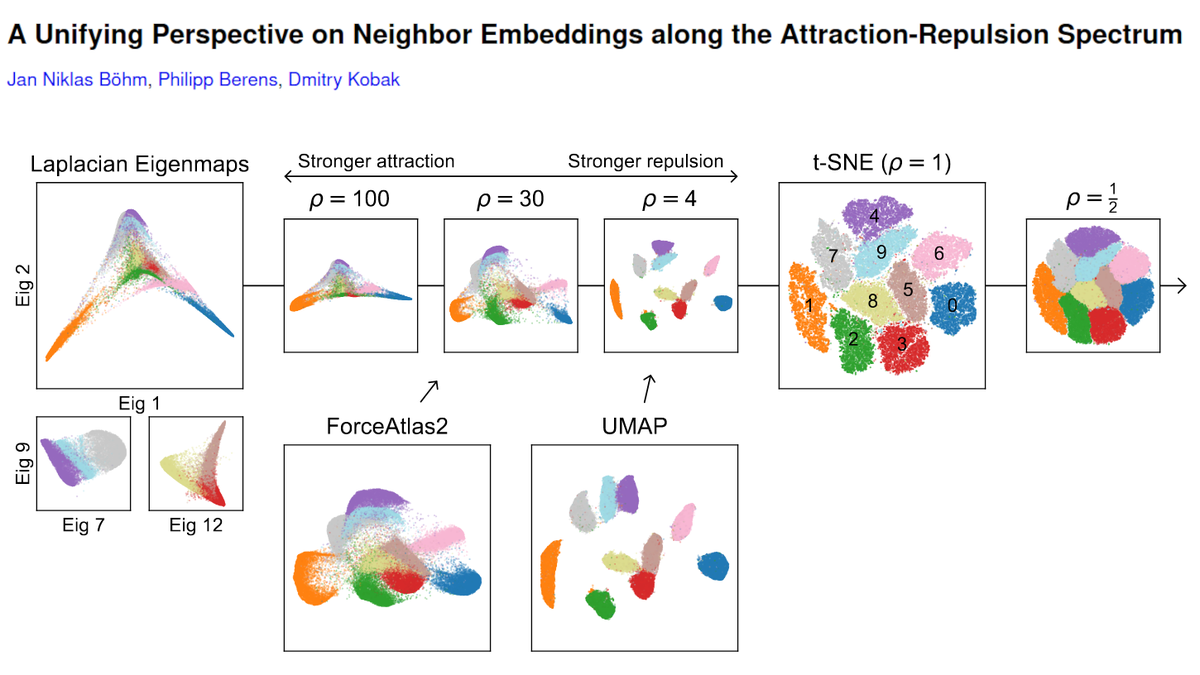
So what's up with the Russian election two weeks ago? Was there fraud?
Of course there was fraud. Widespread ballot stuffing was videotaped etc., but we can also prove fraud using statistics.
See these *integer peaks* in the histograms of the polling station results? 🕵️♂️ [1/n]
Of course there was fraud. Widespread ballot stuffing was videotaped etc., but we can also prove fraud using statistics.
See these *integer peaks* in the histograms of the polling station results? 🕵️♂️ [1/n]

These peaks are formed by polling stations that report integer turnout percentage or United Russia percentage. E.g. 1492 ballots cast at a station with 1755 registered voters. 1492/1755 = 85.0%. Important: 1492 is not a suspicious number! It's 85.0% which is suspicious. [2/n]
We can use binomial Monte Carlo simulation to find how many polling stations with integer percentages there should be by chance. Then we can compute the number of EXCESS integer polling stations (roughly the summed heights of all INTEGER PEAKS).
Resulting excess is 1300. [3/n]
Resulting excess is 1300. [3/n]
1300 clearly fraudulent stations is a lot! But it's not as many as in the last years, especially in 2020 (constitutional referendum). [4/n]

https://twitter.com/hippopedoid/status/1278673265075118081

Does it mean that there was less fraud this time? Not at all! But it seems it was less stupidly done.
Here is a 2D scatter plot of turnout vs. United Russia result. This suggests the actual result was ~30%, possibly a few % more, instead of the official 49.8%. [5/n]
Here is a 2D scatter plot of turnout vs. United Russia result. This suggests the actual result was ~30%, possibly a few % more, instead of the official 49.8%. [5/n]

Here is how this "comet" compares to the previous federal elections over the Putin era.
In terms of how many % points were added to the leader's result during counting, this election may actually have been the worst ever (but it's a close call with 2011). [6/n]
In terms of how many % points were added to the leader's result during counting, this election may actually have been the worst ever (but it's a close call with 2011). [6/n]

See our series of papers (with Sergey Shpilkin and @MPchenitchnikov) regarding the methodology of integer peak calculations:
* projecteuclid.org/journals/annal…
* rss.onlinelibrary.wiley.com/doi/full/10.11…
* rss.onlinelibrary.wiley.com/doi/full/10.11…
* rss.onlinelibrary.wiley.com/doi/abs/10.111…
[7/n]



* projecteuclid.org/journals/annal…
* rss.onlinelibrary.wiley.com/doi/full/10.11…
* rss.onlinelibrary.wiley.com/doi/full/10.11…
* rss.onlinelibrary.wiley.com/doi/abs/10.111…
[7/n]




Just an example of how stupidly it _was_ sometimes done. This entire 2D integer peak with 75.0% turnout and 75.0% United Russia result (back in 2011) was due to one single city: Sterlitamak (in Bashkortostan). Obviously they did not even count the ballots. [8/n] 

You can find all the data (in CSV) and my analysis code (as a Python notebook) at github.com/dkobak/electio…. The data have been scraped by Sergey Shpilkin. [9/n]
Scraping the data was much more difficult this time, because it was deliberately obfuscated (see below). Of course eventually people wrote several de-obfuscators, e.g. see this very detailed write-up by Alexander Shpilkin: purl.org/cikrf/un/unfuc…. [10/10]
https://twitter.com/hippopedoid/status/1439897585783803914
Update: here is my new favourite plot on this topic. I pooled the data from all 11 federal elections from 2000 to 2021 and made a scatter plot of all 1+ million polling stations together. Just look at the periodic integer pattern in the top-right (i.e. fraudulent) corner! [11/10] 

• • •
Missing some Tweet in this thread? You can try to
force a refresh