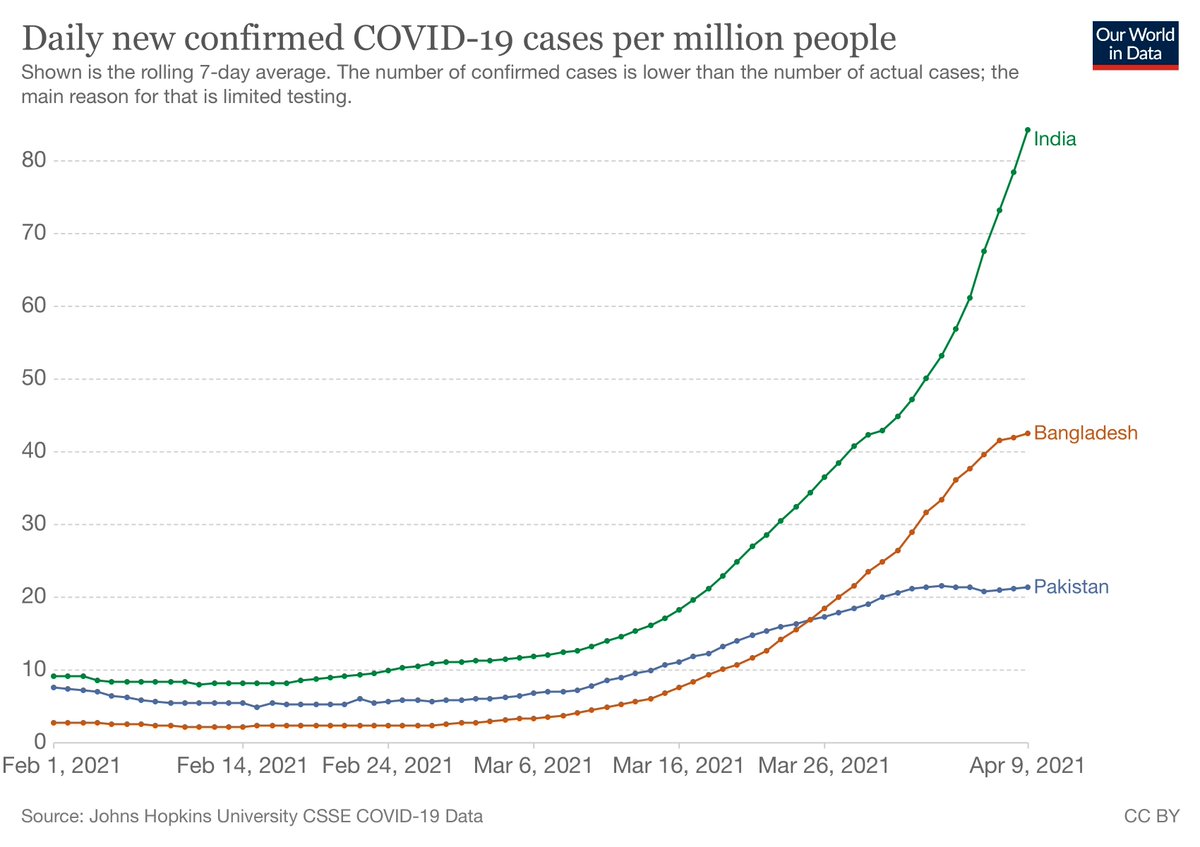
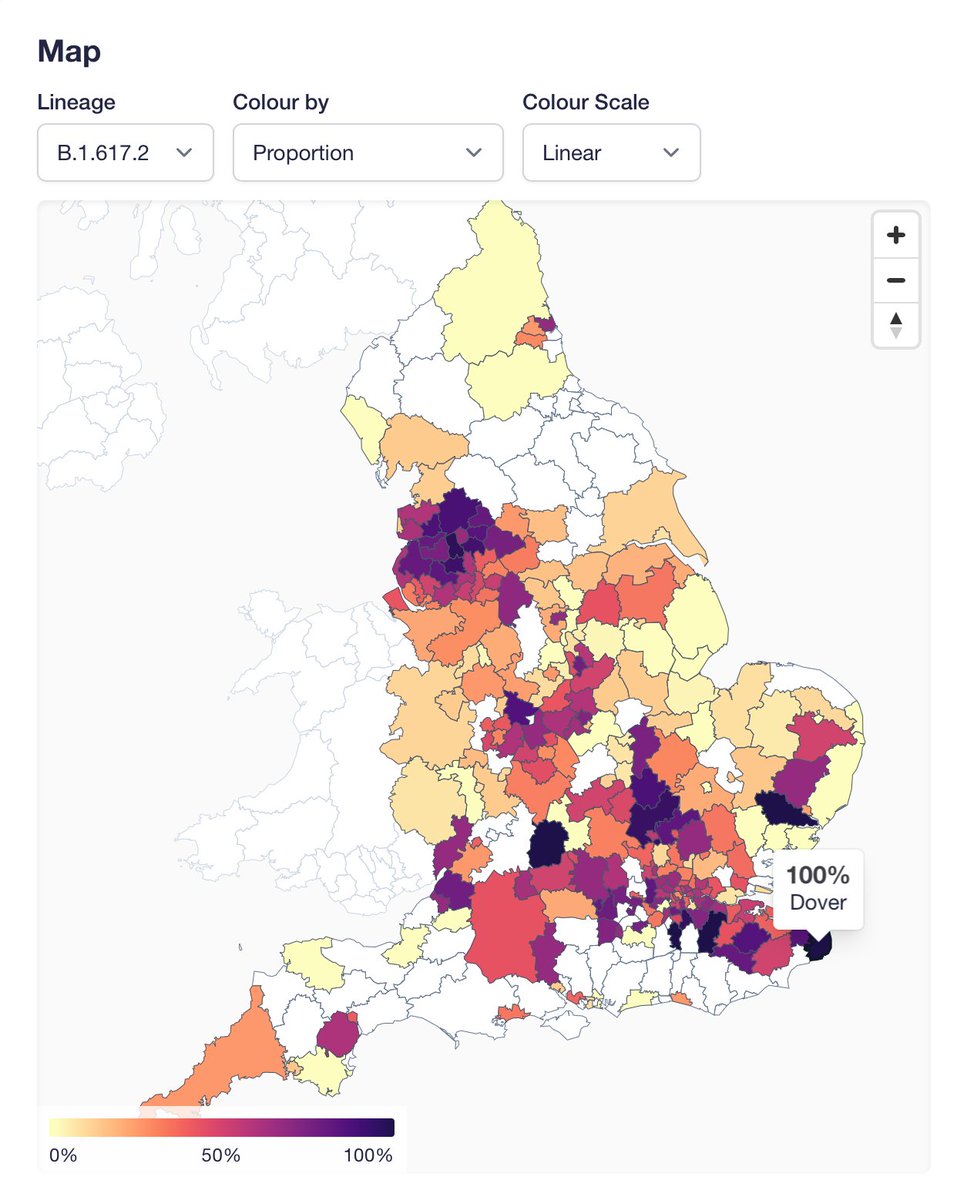
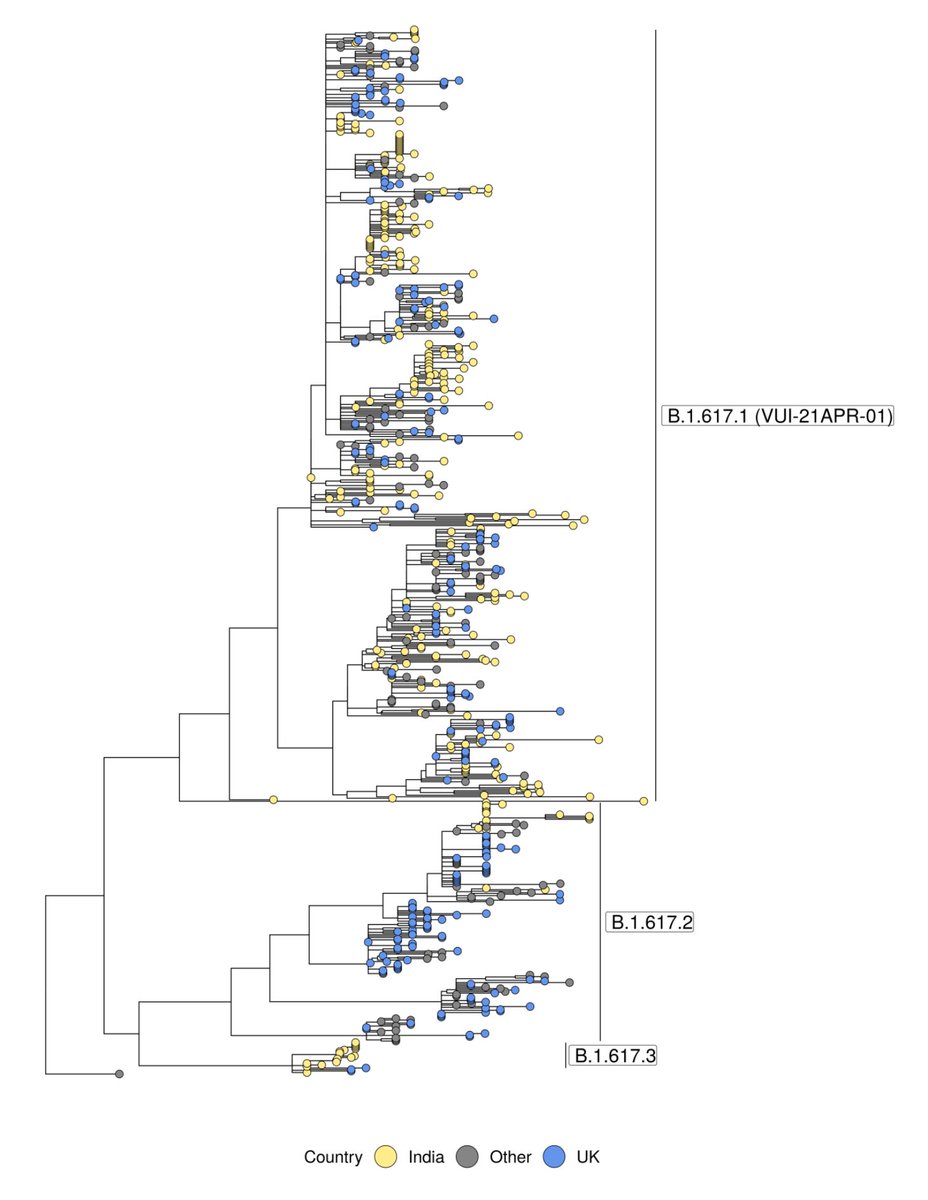
Monday always means new data at covid19.sanger.ac.uk, but this week also brings new features! Overall picture should come as no surprise: Delta variant growing in proportion and absolute numbers. In fact, next week is likely to be our highest ever count of one lineage🧵 

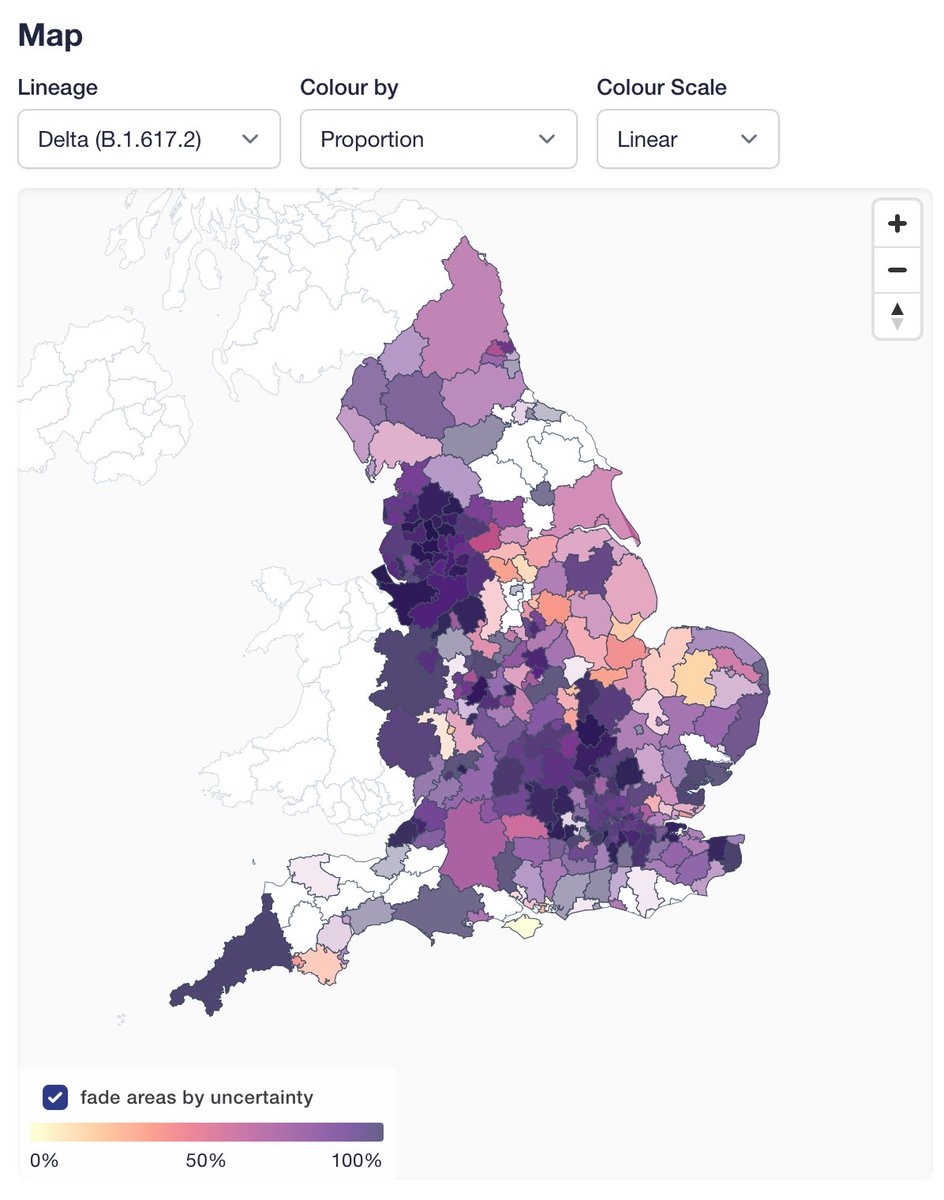
First new feature is "fade areas by uncertainty", or "nebulosity view", as I like to call it. We fade the colouring of local authorities on proportion view to give a visual hint that you shouldn't over-interpret an area with 100% Delta if it only has a handful of sequences. 
Related to this, we show confidence intervals on proportion tooltips and in the unstacked version of the proportion graph. Compare the following two local authorities with similar point estimates, but different amounts of data. 



Finally a couple of technical points: we fixed the creeping mystery "B" lineages, that are actually Delta, and added support for Pango aliasing so everything goes up to "A" or "B" at the root, and there's no more "other".
Thanks as usual to the whole team, especially @theosanderson, @richgoater, and @msinnott123 for this week's updates!
• • •
Missing some Tweet in this thread? You can try to
force a refresh