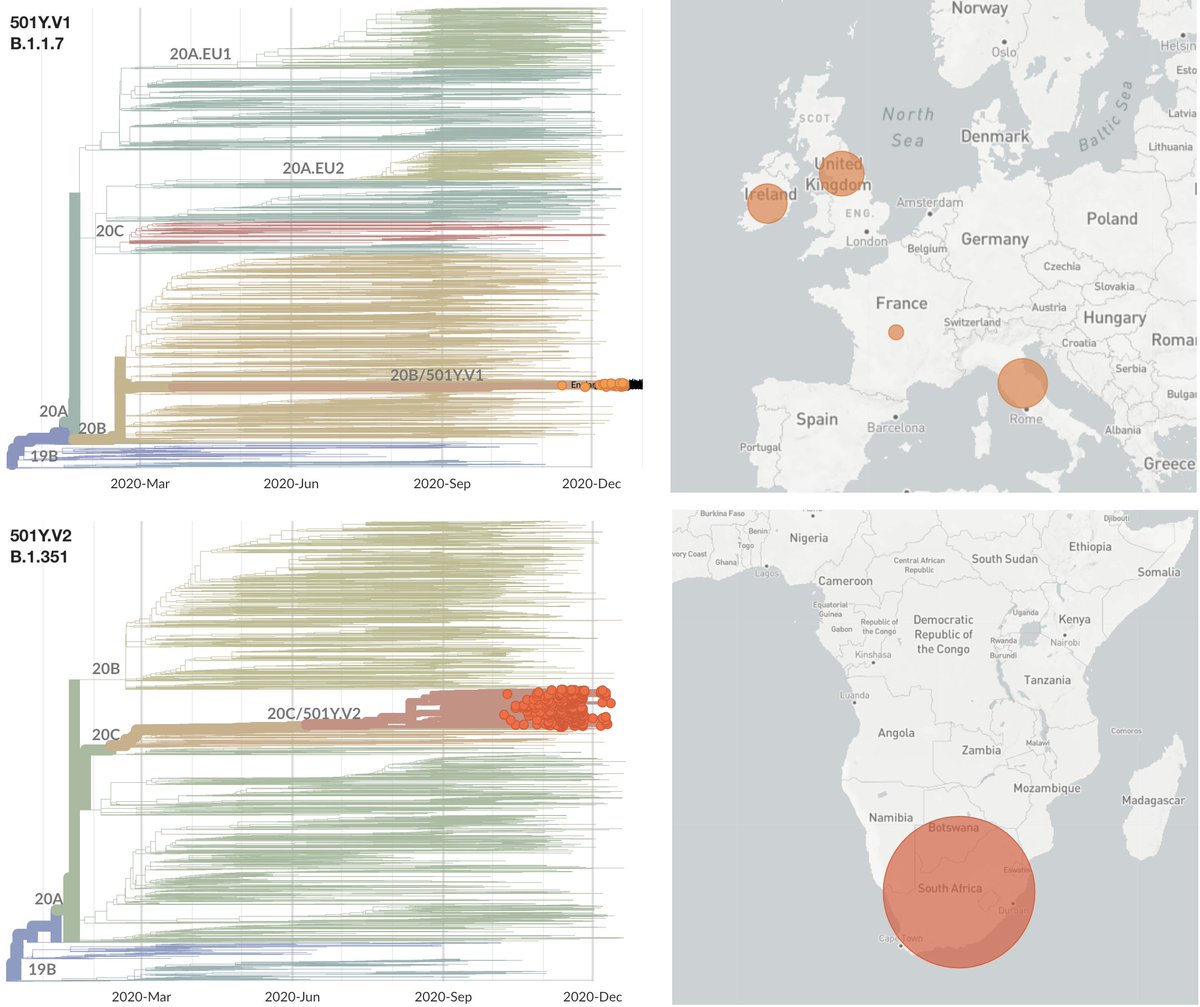
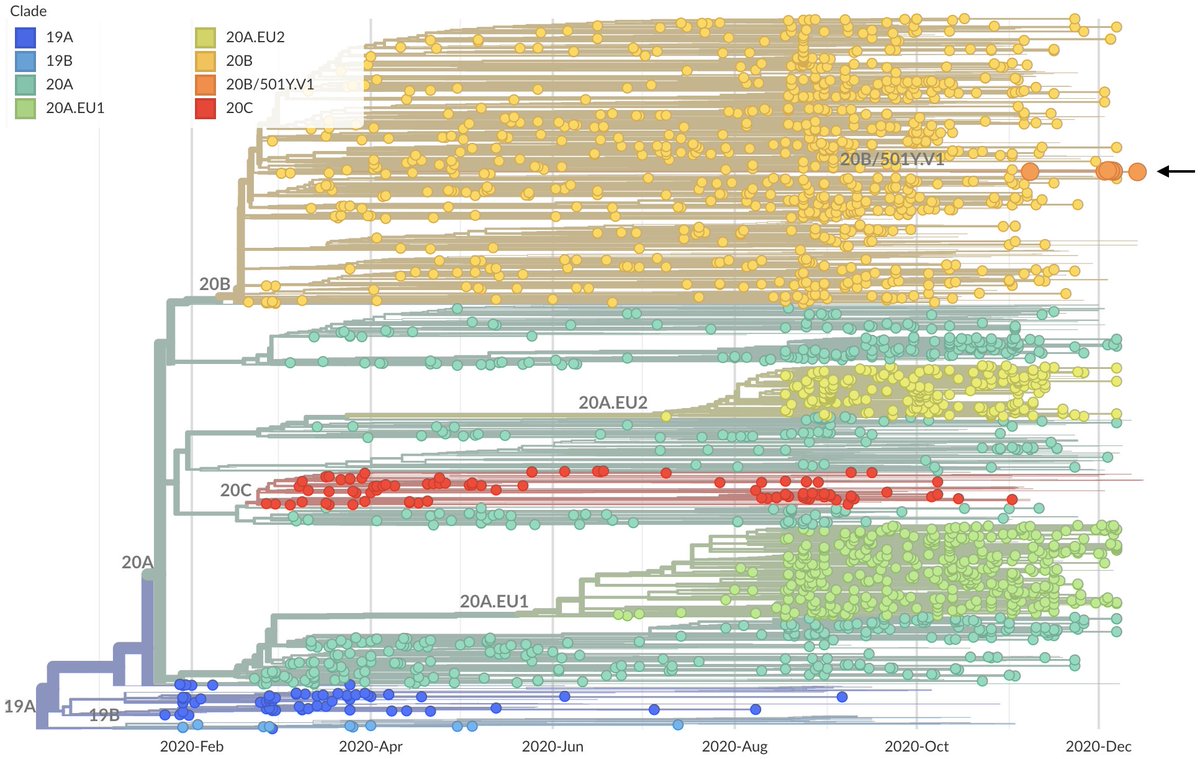
Important new study by Wibmer et al (biorxiv.org/content/10.110…) of neutralization by convalescent sera on wildtype vs 501Y.V2 variant viruses circulating in South Africa. It shows that mutations present in 501Y.V2 result in reduced neutralization capacity. 1/10
Here, I've replotted data from the preprint to make effect size a bit more clear. Each line is sera from one individual tested against wildtype virus on the left and 501Y.V2 variant virus on the right. Note the log y axis (as is common with this type of data). 2/10 
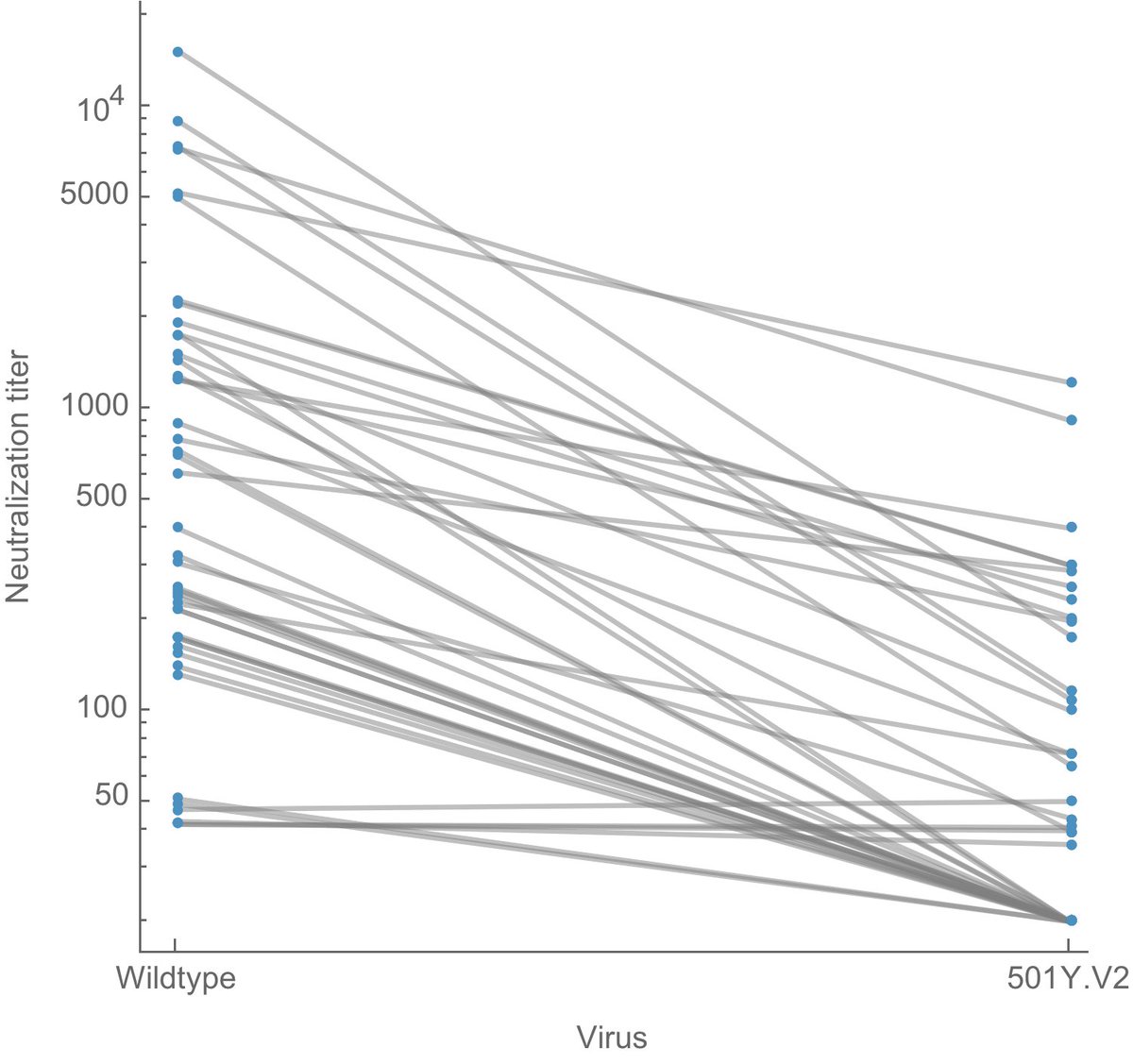
It's clear that 501Y.V2 often results in reductions of neutralization titer, quantified as "fold-reduction" where, for example, a 2-fold reduction in titer would mean that you need twice as much sera to neutralize the same amount of virus in the assay. 3/10
Here, I'm plotting distribution of fold-reduction across the 44 individuals tested. You can see there is a median 8-fold reduction in titer when comparing wildtype to 501Y.V2 virus, though some individuals show no reduction and other individuals show a 64-fold reduction. 4/10 
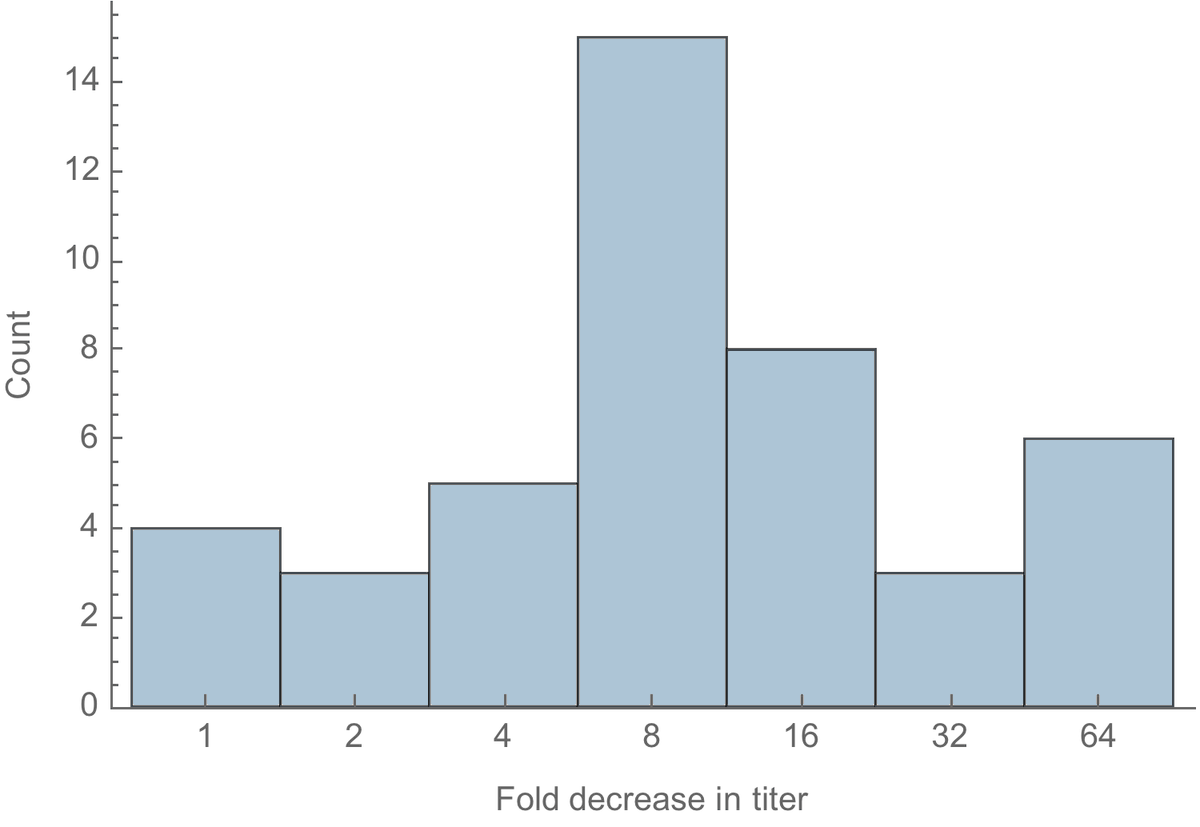
To put an 8-fold drop in context, the @WHO uses an 8-fold threshold when deciding to update the seasonal influenza vaccine (note this is a different virus and neutralization results may not be directly comparable, but it at least gives a ballpark comparison). 5/10
Also note that the mRNA vaccines in particular are really good vaccines and elicit strong immune responses. A reduction in neutralization from a high starting point will have less of an impact than a reduction from a lower starting point. 6/10
https://twitter.com/trvrb/status/1340410014255652865
We urgently need "immune correlates of protection" determined for COVID-19 vaccination. This would allow extrapolation from reductions in neutralization into expected effects on vaccine efficacy. At the moment, it's guesswork. 7/10
However, if these results are confirmed by further studies, my guess based on the seasonal influenza comparison is that we need to investigate the manufacturing timeline and regulatory steps required to update the "strain" used in the vaccine. 8/10
501Y.V2 is still largely restricted to South Africa, but it (or other antigenically drifted variants) may spread more widely in the coming months. I would be planning this potential "strain" update for fall 2021. 9/10
And all this said, I'll be getting the vaccine as soon as I'm able. We have an amazing vaccine now that works against currently circulating viruses. And if it becomes necessary, this emerging situation can be dealt with through a forthcoming vaccine update. 10/10
• • •
Missing some Tweet in this thread? You can try to
force a refresh