Great chat with @jjcouey
I have some errors to confess to as I spoke to fast.
1)I was enrolled in a PhD program at UW but dropped out to focus on my job when the HGP starting racing with Craig Venter. So No PhD.
twitch.tv/videos/1239989…
I have some errors to confess to as I spoke to fast.
1)I was enrolled in a PhD program at UW but dropped out to focus on my job when the HGP starting racing with Craig Venter. So No PhD.
twitch.tv/videos/1239989…
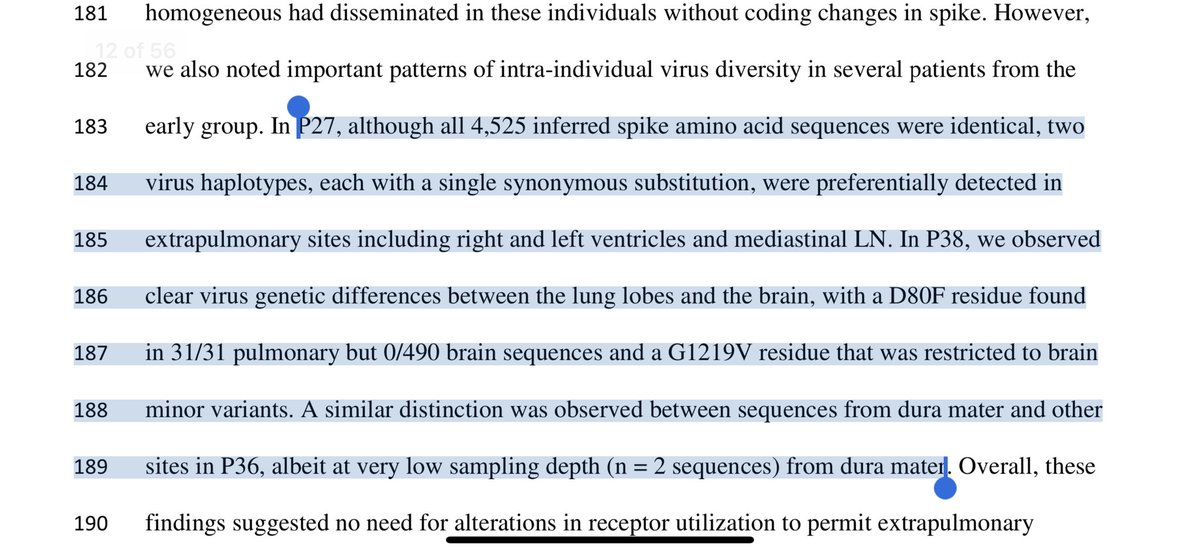
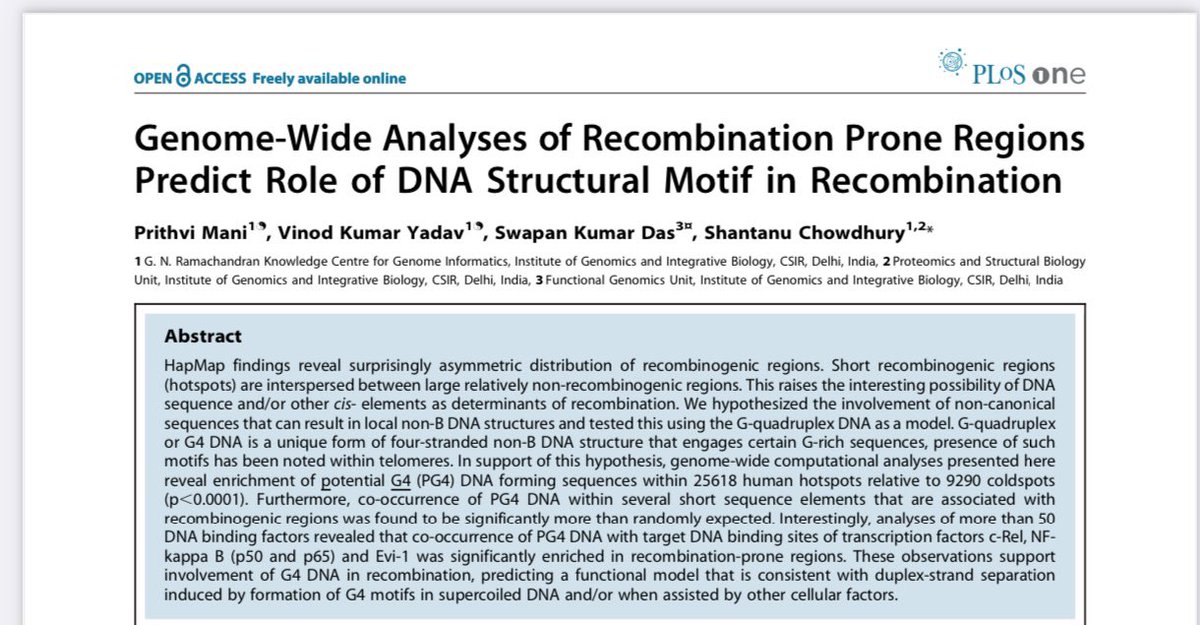
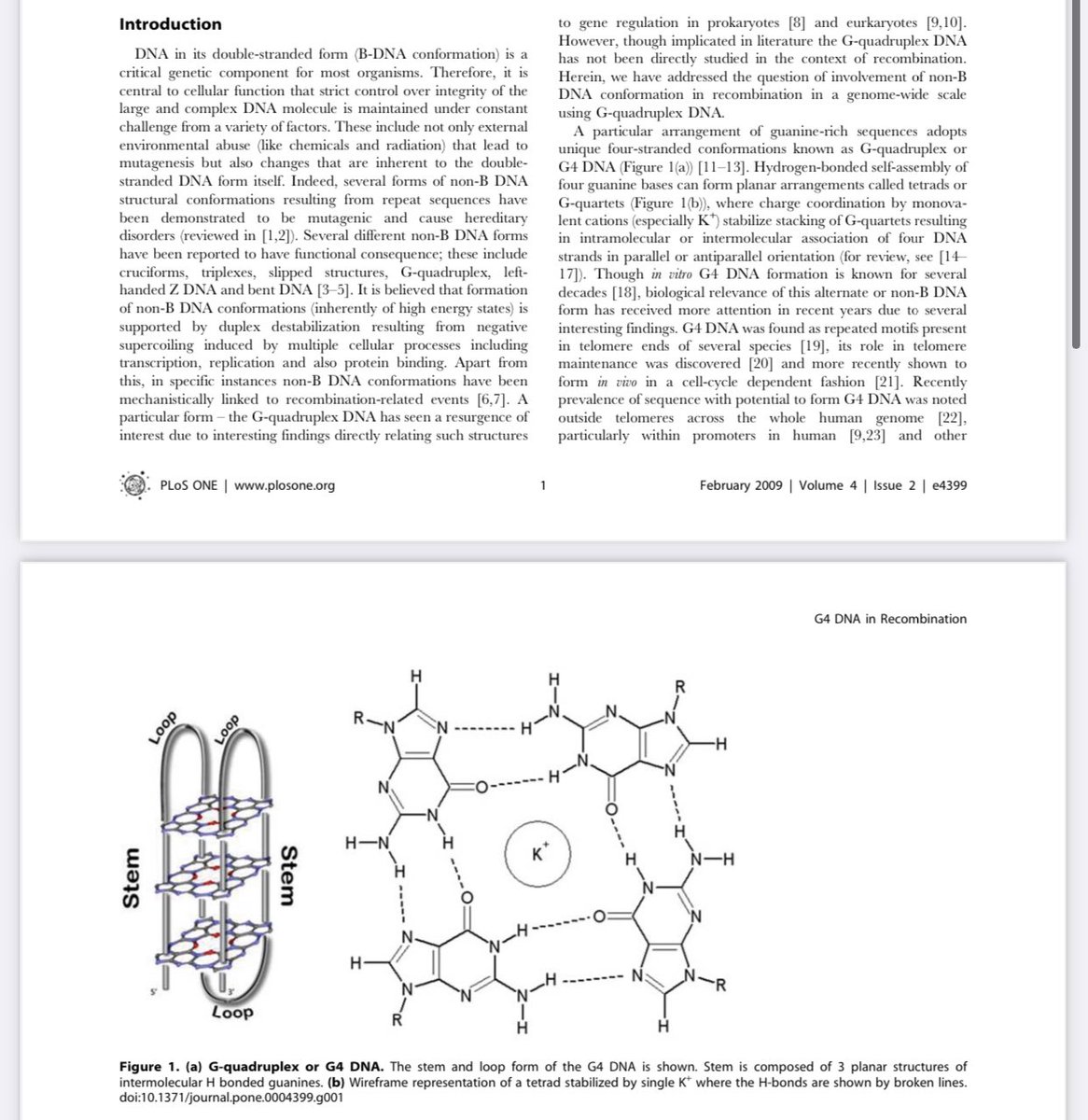
2)nsp13 should be nsp3. It interacts with quadruplex G
academic.oup.com/nar/article/49…
academic.oup.com/nar/article/49…
3)The vaccines have N1-Methyl Pseudouridine and I shouldn't shorthand this to Pseudouridine as the former has less wobble than the latter. N1-Methyl does alter the Tm and increase base stacking in RNA but its methyl group steals one potential H-bond.
ncbi.nlm.nih.gov/labs/pmc/artic…
ncbi.nlm.nih.gov/labs/pmc/artic…
There are Peudouridine synthases which convert U->PseudoU. There are N1-methyltransferase genes that convert Pseudo->N1-methylPseudo. This base methylation is a natural signaling system that likely has never seen a fully methylated mRNA before. Natural methylation is usually 1-2%
This is the regulatory document which identifies many smears. Just search this document for "frag" and you will find very interesting discussions. ema.europa.eu/en/documents/a… 
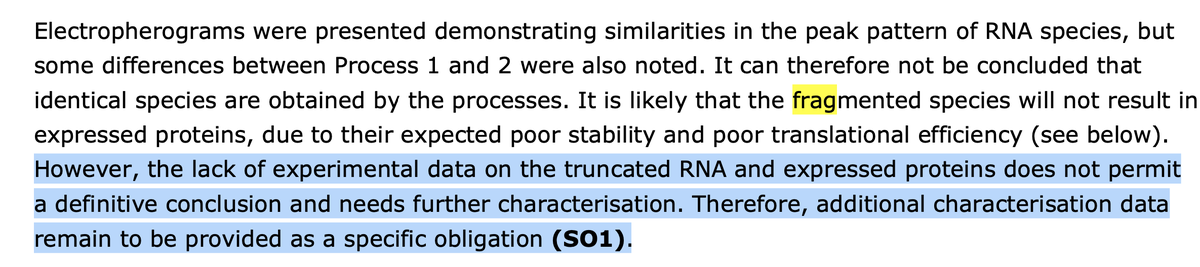

More on N1-methyl pseudouridine and Ribosomal Pausing. This has been shown to alter the protein folding of Spike proteins.
ncbi.nlm.nih.gov/labs/pmc/artic…
More on Ribosomal Pausing -
Postnikova et al.
mdpi.com/1422-0067/22/1…
ncbi.nlm.nih.gov/labs/pmc/artic…
More on Ribosomal Pausing -
Postnikova et al.
mdpi.com/1422-0067/22/1…
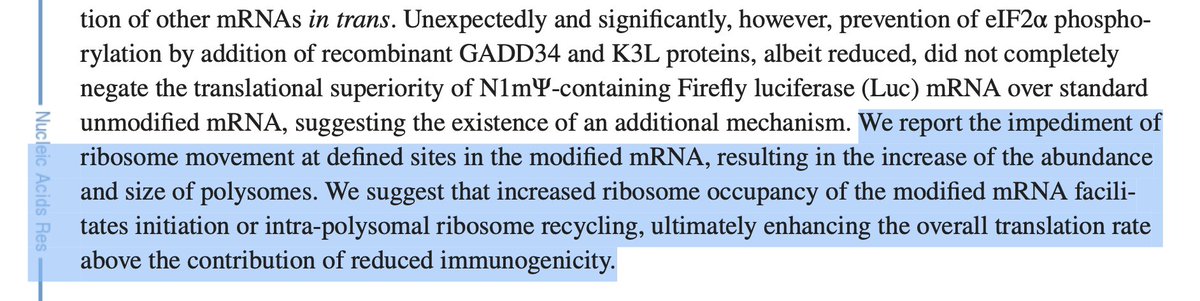

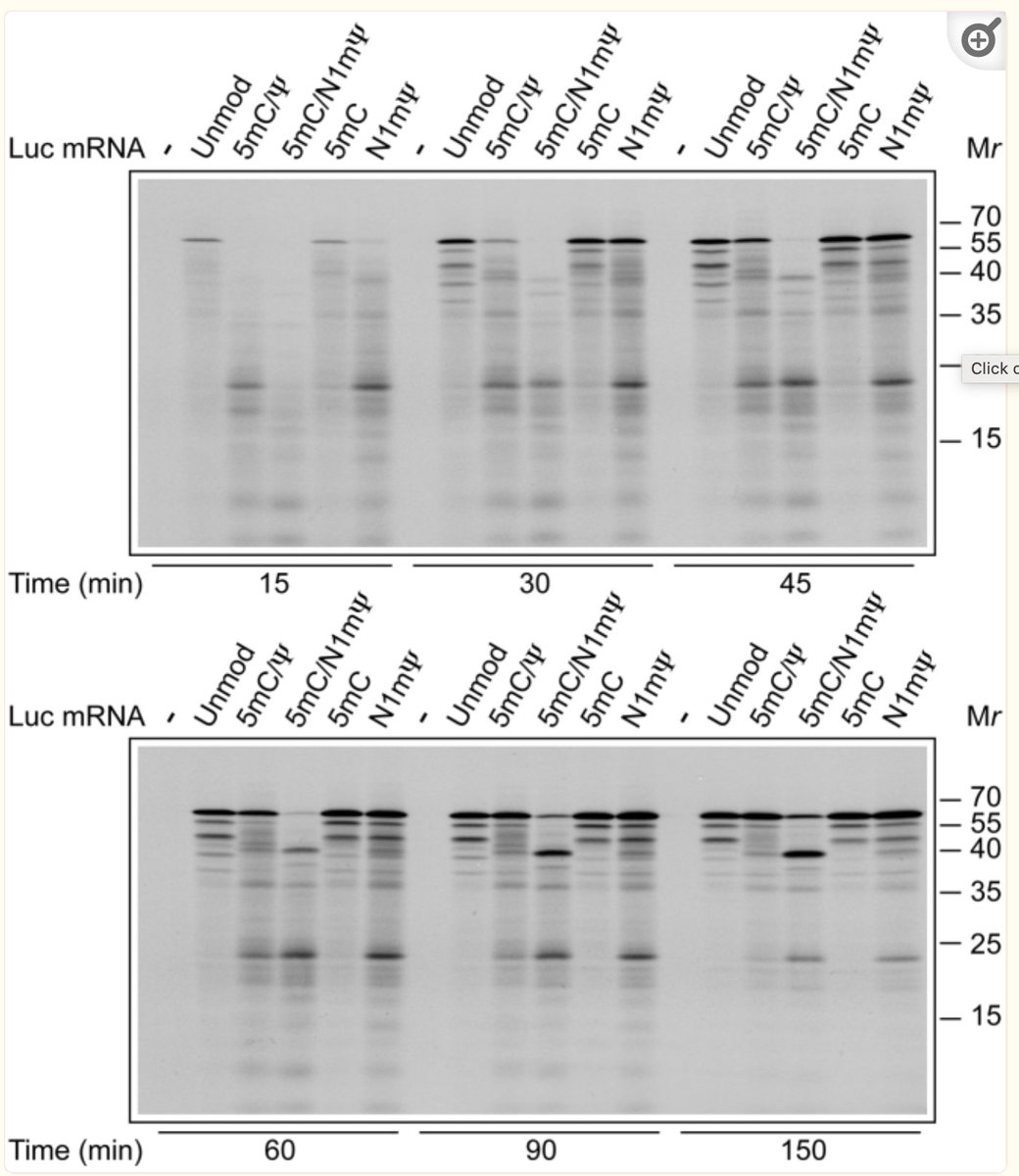
The unmodified mRNA provides mostly 1 band. Add in N1-methyl Pseudouridine and you get bright banding patterns and pause sites. There is no 6X His Tag to purify the full length proteins so all of these get injected. 

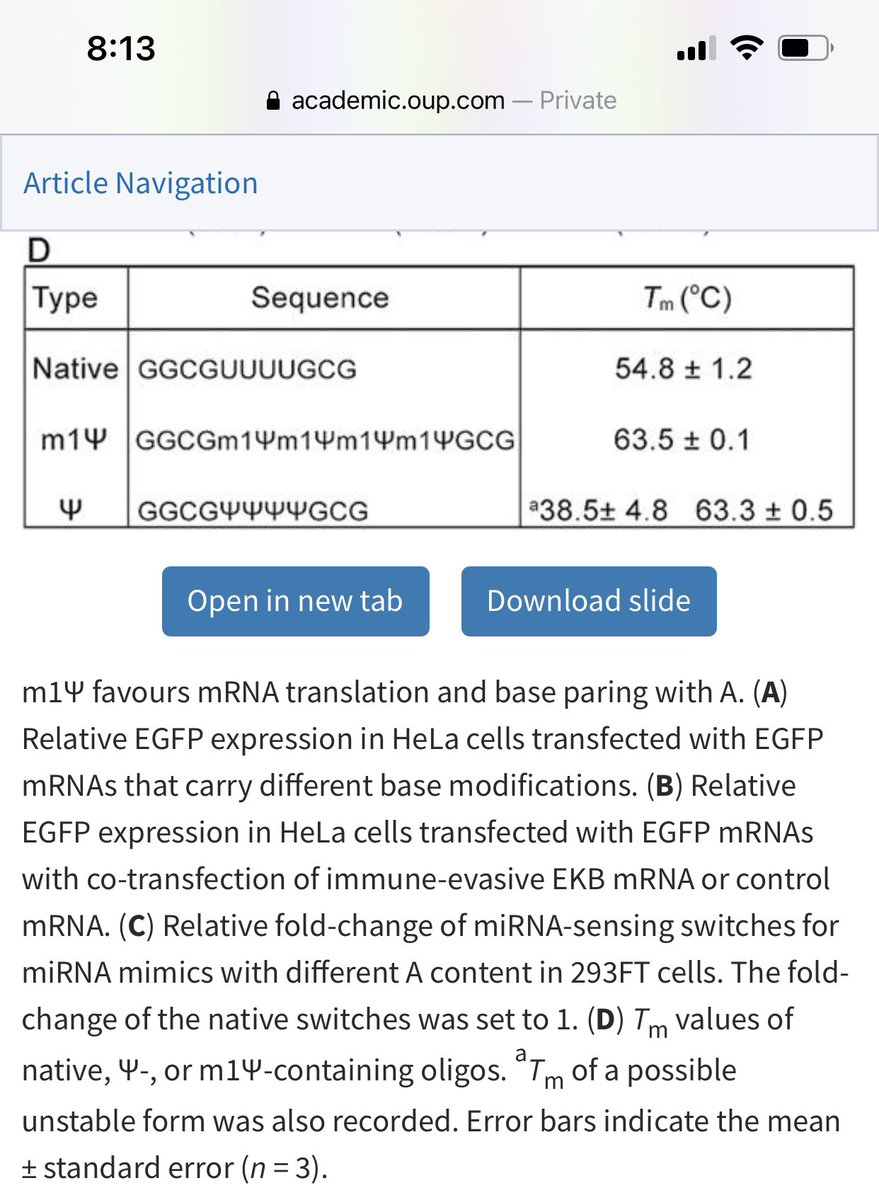
While pseudU has one more hydrogen donor than N1 methylpseudoU, the methyl group compensates with better base stacking and an equal shift in Tm.
Not clear what this does to translation fidelity in light of the bad wrap with pseudoU and the gel smears in regulatory docs.
Not clear what this does to translation fidelity in light of the bad wrap with pseudoU and the gel smears in regulatory docs.

Just the addition of 4 pseudo’s jumps the Tm 9C.
Imagine all of spike.
academic.oup.com/nar/article/48…
Supplement has the Tm figure.
Imagine all of spike.
academic.oup.com/nar/article/48…
Supplement has the Tm figure.

• • •
Missing some Tweet in this thread? You can try to
force a refresh